Abstract
Objectives
Extraprostatic extension (EPE) of prostate cancer (PCa) is predicted using clinical nomograms. Incorporating MRI could represent a leap forward, although poor sensitivity and standardization represent unsolved issues. MRI radiomics has been proposed for EPE prediction. The aim of the study was to systematically review the literature and perform a meta-analysis of MRI-based radiomics approaches for EPE prediction.
Materials and methods
Multiple databases were systematically searched for radiomics studies on EPE detection up to June 2022. Methodological quality was appraised according to Quality Assessment of Diagnostic Accuracy Studies 2 (QUADAS-2) tool and radiomics quality score (RQS). The area under the receiver operating characteristic curves (AUC) was pooled to estimate predictive accuracy. A random-effects model estimated overall effect size. Statistical heterogeneity was assessed with I2 value. Publication bias was evaluated with a funnel plot. Subgroup analyses were performed to explore heterogeneity.
Results
Thirteen studies were included, showing limitations in study design and methodological quality (median RQS 10/36), with high statistical heterogeneity. Pooled AUC for EPE identification was 0.80. In subgroup analysis, test-set and cross-validation-based studies had pooled AUC of 0.85 and 0.89 respectively. Pooled AUC was 0.72 for deep learning (DL)–based and 0.82 for handcrafted radiomics studies and 0.79 and 0.83 for studies with multiple and single scanner data, respectively. Finally, models with the best predictive performance obtained using radiomics features showed pooled AUC of 0.82, while those including clinical data of 0.76.
Conclusion
MRI radiomics–powered models to identify EPE in PCa showed a promising predictive performance overall. However, methodologically robust, clinically driven research evaluating their diagnostic and therapeutic impact is still needed.
Clinical relevance statement
Radiomics might improve the management of prostate cancer patients increasing the value of MRI in the assessment of extraprostatic extension. However, it is imperative that forthcoming research prioritizes confirmation studies and a stronger clinical orientation to solidify these advancements.
Key Points
• MRI radiomics deserves attention as a tool to overcome the limitations of MRI in prostate cancer local staging.
• Pooled AUC was 0.80 for the 13 included studies, with high heterogeneity (84.7%, p < .001), methodological issues, and poor clinical orientation.
• Methodologically robust radiomics research needs to focus on increasing MRI sensitivity and bringing added value to clinical nomograms at patient level.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Prostate MRI has reshaped prostate cancer (PCa) diagnostic pathway, as the “MRI-first” approach is increasingly gaining recognition as standard-of-care, with lesion detection to be performed following acknowledged MRI guidelines (Prostate Imaging-Reporting and Data Systems version 2.1 (PI-RADS v2.1)) [1, 2]. As for local staging, MRI-based grading systems for extraprostatic extension (EPE) detection appear promising, with suspicious imaging features mainly represented by tumor capsular contact length, capsular bulge/irregularity, and frank capsular breach [3,4,5,6,7]. Nevertheless, present guidelines still base pre-treatment local staging (cT) assessment exclusively on digital rectal examination findings, with only a weak recommendation to the use of MRI in this setting [8, 9]. This might be partly due to the lack of standardization affecting the ability of radiologists to detect EPE on MRI, as well as to the reported poor sensitivity [3, 10]. The adoption of different clinical nomograms differently incorporating cT, patient’s demographics, and laboratory and biopsy findings is widespread as a tool for EPE prediction and overall risk stratification, but these do not include imaging features [11]. However, in external validation cohorts, multivariate risk calculators accounting for MRI features have shown significantly higher performance compared to clinical nomograms [12, 13].
Novel MRI biomarkers and advanced quantitative techniques have been recently investigated to further enhance the value of MRI for EPE detection and possibly overcome current limits [14,15,16,17,18,19,20,21,22,23,24,25,26]. In particular, radiomics is a novel approach that can translate images into valuable quantitative datasets by the analysis of many mathematical parameters describing different MR image properties [27]. Furthermore, the application of artificial intelligence (AI) and machine learning (ML) may improve the discovery of task-specific features such as anatomic localization, tumor contacting, neurovascular bundles, or direct evidence of abnormalities in periprostatic soft tissue. However, AI in medicine is facing a reproducibility crisis which is hindering the translation of radiomics research into clinical practice, with the scientific community advocating for more robust methodology in the field [28,29,30,31,32]. Additionally, radiomics studies should be specifically designed to address unsolved clinical needs. In the case of EPE prediction, this would mean contextualizing the possible added value of radiomics compared to current standard (i.e., cT and/or clinical nomograms) as well as possibly viable alternatives (i.e., conventional MRI approaches).
In this light, we performed a systematic review and meta-analysis aiming to provide insights into MRI-based radiomics approaches for EPE prediction, by estimating their performance, exploring their heterogeneity, summarizing the main factors impacting the diagnostic accuracy, and focusing on those methodological and study design shortcomings that must be addressed to increase their clinical value.
Materials and methods
This meta-analysis followed the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) statement (supplementary material for PRISMA Checklist) [33]. The review protocol is registered on PROSPERO (CRD42023392319) [34].
Literature search and study selection
An English literature search was performed using the PubMed, Embase, Scopus, and Web of Science databases to identify articles published until June 30th, 2022. The study search was restricted to data obtained in humans and conducted using the following key words with their variations: “Magnetic Resonance Imaging,” “MRI,” “Prostate cancer,” “Machine Learning,” “Radiomics,” and “Extraprostatic.” The full search strategy is presented in the supplementary materials. The title and abstract of potentially relevant studies were screened for appropriateness before retrieval of the full article by two reviewers (M.G. and A.S.) and disagreements were discussed with a third reviewer (A.P.) to reach a consensus. The full-published reports of the abstracts selected by the reviewers were retrieved and the same reviewers independently performed a second-step selection based on the inclusion criteria; disagreements were resolved by consensus. Furthermore, in accordance with PRISMA guidelines, the bibliographies of retrieved articles were manually reviewed to identify additional items meeting inclusion criteria.
Data extraction and eligibility
Data from the included studies was collected in a database. Each study was initially identified considering author, journal, and year of publication. Total patient population, number of positive and negative cases, study type, and MRI field strength were recorded. Furthermore, information regarding the radiomics and ML pipeline, MRI sequences included, lesion segmentation details, image and data preprocessing steps, feature extraction, feature selection, algorithm, prediction model, and validation strategy were collected. Additionally, the following details regarding study design were also registered: (1) whether the radiomics predictive model incorporated non-radiomics characteristics and if so their specifics; (2) whether a comparison with clinical nomograms and/or conventional imaging assessment was performed to investigate the added value of radiomics. Finally, relevant accuracy metrics were extracted for the subsequent pooled analysis. No minimum sample size was chosen for inclusion. A study was included if all the following criteria were met: (1) an analysis focused on EPE prediction was presented; (2) information on area under the receiver operating curve (AUC) and total number of positive and negative cases, respectively defined as patients with or without EPE included in the analysis, were reported; (3) clear definition of the dataset used in the study and input data source. For the purposes of data pooling, in case of studies reporting data for either internal or external test-set, we considered them separately, while if different models were built, the one with the best predictive performance was selected. Reviews, editorials, abstracts, animal studies, conference presentations, and studies not focused on the topic of interest, published in languages other than English, or presenting insufficient data for pooling were excluded.
Data quality assessment
The methodological assessment of the quality of eligible studies was performed by two reviewers independently (A.S. and A.P.), according to the Quality Assessment of Diagnostic Accuracy Studies 2 (QUADAS-2) tool [35] and radiomics quality score (RQS) [36]. The QUADAS-2 tool offers obvious grades of bias and applicability of primary diagnostic accuracy studies. It comprises four significant domains, namely (1) patient selection; (2) index test; (3) reference standard; and (4) the flow and timing. Each domain contains several signal questions used to help judge the risk of bias (low, high, or unclear). The two reviewers completed the screening process independently. Disagreement in the process of answering questions was discussed until consensus was reached. The RQS represents a system of metrics for the overall evaluation of the methodological validity and thoroughness of radiomics-based studies, and has been adopted in different topics, mainly focused on oncology [37, 38]. It consists of 16 items regarding different steps in the workflow of radiomics. The summed total score ranges between − 8 and 36, while the percentage is calculated on a 0–36 scale.
Statistical analysis
The predictive accuracy (predicting presence of EPE) was quantified using the AUC for the receiver operating characteristic curve analysis. For each of the included studies, the AUC was extracted with corresponding 95% confidence intervals. The AUC standard error was calculated from the total number of positive and negative EPE patients. The I2 value was used to assess statistical heterogeneity, providing an estimate of the percentage of variability among included studies. I2 values of 0–25%, 25–50%, 50–75%, and > 75% represent very low, low, medium, and high heterogeneity, respectively. I2 was calculated as follows: I2 = 100% × (Q − df) / Q, where Q is Cochran’s heterogeneity statistic and df the degrees of freedom. The weight of each study was calculated with the inverse variance method, in which the weight given to each study is chosen to be the inverse of the variance of the effect estimate, minimizing the uncertainty of the pooled effect estimate. Pooling of studies was conducted, and effect size assessed using a random-effects model, which allows to estimate the distribution of true effects between studies accounting for heterogeneity. Publication bias was explored using the effective sample size funnel plot described by Egger et al Two-sided p values ≤ 0.05 were considered statistically significant [39]. Subgroup analyses were also performed in relation to the use of dedicated test-set or not, deep learning (DL) or not, single or multiple scanners, and if the best predictive models only included radiomics features or if they combined radiomics features with clinical data.
Results
Study selection and data extraction
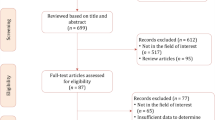
The complete literature search process is presented in Fig. 1. In brief, the initial search identified 260 potentially eligible citations. After removing duplicates, 206 records were screened by the reviewers. After the titles and abstracts evaluation, 192 citations were discharged because they were judged to be non-relevant or non-pertinent. Thus, 14 full-text articles were blindly assessed by each investigator for eligibility. After revision, 2 articles were excluded leaving 12 articles. Furthermore, after screening the reference lists of the eligible studies, we identified one additional article that had not been initially captured in our initial search, despite the presence of the selected keywords. Finally, 13 items were the basis of the present meta-analysis [14,15,16,17,18,19,20,21,22,23,24,25,26].
The baseline characteristics of the included studies are shown in Tables 1 and 2. Briefly, 3 studies involved more than one institution [14, 15, 19], while 6 investigations adopted multiple scanners [14, 15, 17, 19, 21, 22]. Regarding MRI field strength, a 3-T scanner was mostly adopted (10/13) [14, 17,18,19, 21,22,23,24,25,26], while one study was conducted on both 1.5-T and 3-T scanners [15]. Only 1 study used a semi-automatic lesion segmentation approach [16], while the remaining investigations performed either 3D-manual segmentation [14, 15, 17, 18, 21, 22, 24,25,26] or a combination of manual and automatic approach [20]. Details regarding the study design are presented in Supplementary Tables 1 and 2. Finally, 9 studies had a dedicated test-set [14, 15, 17,18,19, 21,22,23, 26].
Data quality assessment
The methodological quality assessment of risk of bias within eligible studies according to QUADAS-2 is shown in Fig. 2. In particular, risk of bias due to patient selection was unclear in six studies because a statement on consecutive or random selection was not present [14, 16, 17, 23, 24, 26]. Regarding index test domain, the risk of bias was unclear in seven studies due to the lack of preprocessing details [14, 18,19,20,21,22, 26], while it was high in three studies who did not test feature robustness [17, 24, 25]. Only one study had an unclear risk of bias for providing very few details regarding reference standard [24]. In five cases, the authors did not clearly report the time passed between MRI and radical prostatectomy, receiving an unclear risk of bias [14, 16, 17, 23, 24]. As for applicability concerns, one study scored at unclear risk of bias for the lack of sufficient details regarding patient selection and reference standard [24].
Methodological quality of the included studies assessed according to the Quality Assessment of Diagnostic Accuracy Studies 2 tool for risk of bias and applicability concerns. The green circle represents the low risk of bias, the yellow circle the unclear risk of bias, and the red circle the high risk of bias
The methodological quality assessment of the included studies according to RQS is shown in Supplementary Table 3. The total RQS ranged from 0 to 33% of the maximum rating, with a median score of 10/36 (interquartile range 11). The RQS was low especially due to the lack of prospective design (all studies were retrospective) and of comparison with gold standard. All investigations performed discriminations statistics while none of them made their code or data publicly available.
Statistical analysis
The radiomics models for EPE characterization showed an overall pooled AUC = 0.80 (95% CI = 0.74–0.86) (Fig. 3). Study heterogeneity was 84.7% (p < 0.001).
Forest plot of single studies for the pooled area under the curve (AUC) and 95% CI of extraprostatic extension (EPE) characterization. Horizontal lines represent 95% confidence interval of the point estimates. The diamond means the pooled AUC estimate. The red dotted vertical line represents the overall pooled estimate. ^ internal test-set, * external test-set 1, ° external test-set 2
Subgroup analysis was performed to compare studies employing a dedicated test-set within their pipeline and those who did not. Four studies belonged to the latter group with a pooled AUC of 0.89 (95% CI = 0.78–0.99) and heterogeneity of 89.6% (p < 0.001). The pooled AUC for the remaining studies was 0.78 (95% CI = 0.73–0.82) and 46.4% heterogeneity (p = 0.038). The corresponding plots are presented in Supplementary Fig. 1.
Three studies employed DL within their pipeline for EPE assessment, with a pooled AUC of 0.72 (95% CI = 0.60–0.84) and heterogeneity of 68.4% (p = 0.042). The pooled AUC for the studies not adopting DL was 0.82 (95% CI = 0.76–0.89) with heterogeneity of 83.7% (p < 0.001). The corresponding plots are presented in Supplementary Fig. 2.
Nine studies employed multiple scanners with a pooled AUC of 0.79 (95% CI = 0.74–0.84) and heterogeneity of 32.6% (p = 0.16). The pooled AUC for the investigations adopting single scanners was 0.83 (95% CI = 0.73–0.92) with heterogeneity of 89.8% (p < 0.001). The corresponding plots are presented in Supplementary Fig. 3.
Finally, the five studies in which the best predictive performance was achieved by combined models showed a pooled AUC of 0.76 (95% CI = 0.71–0.82) and heterogeneity of 14.8% (p = 0.320). The pooled AUC for the remaining studies wherein the best predictive performance was obtained with only-radiomics models was 0.82 (95% CI = 0.75–0.89) with heterogeneity of 84.3% (p < 0.001). The corresponding plots are presented in Supplementary Fig. 4.
Discussion
The present systematic review allowed to identify 13 studies focused on MRI radiomics applications for EPE detection, published between 2019 and 2022, which is in line with the growing overall interest in the field [40, 41]. Most involved more than 100 patients and explored the feasibility of a broad spectrum of different algorithms and models, with none reporting negative results. Nevertheless, limitations in study design and methodological quality were present. In order to be clinically relevant, these studies should seek proofs of an added value of radiomics for EPE prediction over or in addition to MRI and/or currently embraced clinical nomograms. Unfortunately, less than half of the included studies proposed a comparison with conventional MRI assessment as performed by a radiologist while only a single study comparing the radiomics model with a recognized clinical nomogram was found. Similarly, a minority of the studies included clinical data in a holistic radiomics model, and only one presented a fully integrated prediction model obtained combining an established clinical nomogram, the radiologist assessment, and MRI radiomics features [20]. Finally, none of the studies was specifically designed to investigate the possible role of radiomics in filling the previously reported sensitivity gap of conventional MRI assessment for EPE detection [10]. Regarding methodological concerns emerged with QUADAS-2, the shortage of preprocessing details and lack of test feature robustness represent crucial points, as also highlighted by “how to” guides recently published aiming to standardize practice in radiomics [42, 43]. Furthermore, dataset quality should be prioritized to avoid the “garbage-in, garbage-out” phenomenon, for example, by defining the maximum time elapsed between MRI and radical prostatectomy to ensure that EPE cases are reliably classified [32]. Using a dedicated tool (RQS), the overall methodological quality of included studies was found to be heterogeneous, with no prospectively designed studies and sometimes inadequate validation strategies. Neglected items also include investigating biological correlates and publicly sharing data, which might allow researchers to increase the understanding of how radiomics features can play a role for EPE prediction. Indeed, poor explainability is a recognized problem with radiomics, especially compared to more understandable MRI quantitative parameters, like the apparent diffusion coefficient, that have also shown a potential value for EPE prediction [44]. However, low RQS scores should not be intended as a synonym of poor quality but rather as a guide to identify room for improvement; it was also pointed out that DL studies might be penalized by this tool specifically tailored for handcrafted radiomics [45]. In addition to the RQS, there are other checklists currently accessible [46, 47]. For instance, the CLAIM one proves valuable in reporting the modeling components of radiomics research [46]. Furthermore, CLEAR presents a viable alternative, encompassing even more aspects of the studies comprehensively through a single checklist, with a public repository being available to allow the radiomics community to comment on the items and adapt the tool for future versions [47]. All those checklists should be considered to enhance the quality and dependability of radiomics research, and consequently fostering its reproducibility.
Overall, our findings are in line with those of a recent systematic review which included 11 radiomics studies on the topic and similarly underlined the need for greater standardization and rigorousness in methodology [48]. To their merit, the authors of this previous work also included a qualitative synthesis of non-radiomics nomograms for EPE prediction, suggesting a possible added value of MRI to clinical data. Approaching this matter from an alternative perspective, we further expand the knowledge in the field providing a quantitative synthesis of the literature that has shown promising results, with a pooled AUC of 0.80 for EPE prediction MRI-based radiomics models. Different clinical nomograms have been proposed for EPE prediction, including the Memorial Sloan Kettering Cancer Center nomogram and the Partin tables [49, 50]. Nonetheless, these risk stratification tools displayed low accuracy and are strictly correlated with final histopathologic results, with reported AUCs ranging from 0.61 to 0.81 [51]. In a previous meta-analysis, de Rooij et al showed that the pooled sensitivity and specificity were 0.57 and 0.91 for detection of EPE with prostate MRI [10]. However, the included studies assessed EPE with different modalities, including dichotomization, Likert scales, or a standardized lexicon. PI-RADS also addresses EPE, reporting the suspicious features such as tumor contact length, capsule bulging, irregularity, and gross extension, as well as loss of rectoprostatic angle [1]. In a recent meta-analysis of ten studies based on the ESUR PI-RADS scoring system, Li et al reported a pooled AUC of 0.80, with pooled sensitivity and specificity respectively of 0.71 and 0.76 [52]. Furthermore, a 3-point EPE grading system has been recently proposed by Mehralivand and colleagues [53]. In a meta-analysis of four studies using the EPE grading system, Li et al [51] reported a pooled area under the hierarchical summary receiver operating characteristic curve of 0.82. Therefore, our meta-analysis shows that radiomics models align in terms of performance to that of conventional MRI assessment and possibly exceed that of clinical nomograms for the prediction of EPE in PCa, although conflicting data of sensitivity and specificity have been reported.
The subgroup analysis showed lower AUCs for studies with a dedicated (either internal or external) validation set compared to those without that (0.78 vs 0.89). This is unsurprising and widely explained by overfitting, a statistical modeling error that occurs when a function is too closely aligned to a limited set of data [54]. Conversely, a reasonable performance drop is expected when testing a model on an independent external dataset, but this allows to determine its reproducibility and generalizability which are pivotal for clinical translation [55, 56]. Furthermore, the subgroup analysis showed better predictive values for the models using handcrafted radiomics and non-deep ML algorithms compared to those that employed DL (AUC, 0.82 vs 0.72). These results could be at least in part explained by the intrinsic nature of the post-processing ML pipelines, with handcrafted radiomics approaches performing comparably or better than DL algorithm on relatively small-sized datasets [57]. Although new DL algorithms receive much interest from the scientific community, greater attention should be paid to the quality and size of datasets to choose the algorithm that favors the best generalizability of the predictive model. However, it should also be considered that all three studies exploring the predictive performance of DL algorithms had a dedicated test-set. Similar predictive values were found for studies employing single and multiple scanners to assess EPE (AUC of 0.83 and 0.79 respectively). To allow radiomics crossing the translational line between an exploratory investigation method and a standardized added value to precision medicine workflow, more efforts should be done to overcome issues related to multi-scanners and non-uniform scanning parameters from different centers. Finally, the counterintuitive finding of the radiomics-clinical vs radiomics-only models (AUC respectively 0.76 vs 0.82) may be at least partly explained looking at the characteristics of the studies. Indeed, the highest performances of radiomics-only studies were almost exclusively reported in settings with no dedicated test-set [16, 21, 24, 25], with one reaching an AUC as high as 0.98 [16]. Without a proper independent validation, the risk of overly optimistic performance estimates is not negligible. On the other hand, all but one of the radiomics-clinical models [17] were evaluated on a test-set, thus possibly showing lower but more reliable assessments.
Based on the qualitative and quantitative synthesis conducted, we must acknowledge that the primary objective of all the included investigations was to assess new algorithms rather than confirming the predictive performance of previously tested radiomics models. As a result, this has unavoidably led to increased heterogeneity. Given the plethora of available radiomics approaches, we strongly advocate for the necessity of replicative and confirmatory studies to enhance the quality and reliability of radiomics research. Moreover, it is crucial to prioritize the clinical setting wherein radiomics could potentially provide added value. Therefore, radiomics studies should not be limited to technical modeling exercises; instead, they should strive to build compelling evidence and instill confidence in the clinical significance of radiomics.
Our meta-analysis has some limitations to acknowledge. First of all, since accuracy metrics reporting was inconsistent, we were only able to focus on AUC values to perform our meta-analysis. Although previous studies adopted this approach and offer an insight on discriminatory ability [57], pooling additional accuracy metrics would have provided valuable information. Due to the relatively small number of studies included in the quantitative synthesis as well as the high (although expected) heterogeneity, the pooled data should be cautiously interpreted. However, this is a common occurrence in diagnostic accuracy meta-analysis and we explored sources of heterogeneity with the subgroup analyses [58]. Finally, the gray literature was not searched; while some relevant articles might have been missed, gray literature searches are not standardized and source reliability is difficult to prove [59].
In conclusion, radiomics introduces an added layer of complexity to prostate MRI and while it might open an exciting path toward more personalized and precise EPE assessment, its possible role must be brought into context with established tools and more practical alternatives. Technical and diagnostic efficacy studies indicate that radiomics could contribute to redefine how EPE is predicted, alongside radiologist’s evaluation. Methodologically robust research evaluating its diagnostic and therapeutic impact are advocated.
Abbreviations
- AI:
-
Artificial intelligence
- AUC:
-
Area under the curve
- CI:
-
Confidence interval
- DL:
-
Deep learning
- EPE:
-
Extraprostatic extension
- ML:
-
Machine learning
- PCa:
-
Prostate cancer
- QUADAS-2:
-
Quality Assessment of Diagnostic Accuracy Studies 2
- RQS:
-
Radiomics quality score
References
Turkbey B, Rosenkrantz AB, Haider MA et al (2019) Prostate imaging reporting and data system version 2.1: 2019 update of prostate imaging reporting and data system version 2. Eur Urol 76:340–351. https://doi.org/10.1016/j.eururo.2019.02.033
Morlacco A, Sharma V, Viers BR et al (2017) The incremental role of magnetic resonance imaging for prostate cancer staging before radical prostatectomy. Eur Urol 71:701–704. https://doi.org/10.1016/j.eururo.2016.08.015
Falagario UG, Jambor I, Ratnani P et al (2020) Performance of prostate multiparametric MRI for prediction of prostate cancer extra-prostatic extension according to NCCN risk categories: implication for surgical planning. Minerva Urol Nefrol 72:746–754. https://doi.org/10.23736/S0393-2249.20.03688-7
Gatti M, Faletti R, Gentile F et al (2022) mEPE-score: a comprehensive grading system for predicting pathologic extraprostatic extension of prostate cancer at multiparametric magnetic resonance imaging. Eur Radiol 32:4942–4953. https://doi.org/10.1007/s00330-022-08595-9
Asfuroğlu U, Asfuroğlu BB, Özer H et al (2022) Which one is better for predicting extraprostatic extension on multiparametric MRI: ESUR score, Likert scale, tumor contact length, or EPE grade? Eur J Radiol 149:110228. https://doi.org/10.1016/j.ejrad.2022.110228
Huebner NA, Shariat SF (2021) Clinical impact and statistical significance of multiparametric magnetic resonance imaging for local staging of prostate cancer. Eur Urol 79:186–187. https://doi.org/10.1016/j.eururo.2020.11.002
Krishna S, Lim CS, McInnes MDF et al (2018) Evaluation of MRI for diagnosis of extraprostatic extension in prostate cancer. J Magn Reson Imaging 47:176–185. https://doi.org/10.1002/jmri.25729
EAU Guidelines. Edn. Presented at the EAU Annual Congress Amsterdam 2022. ISBN 978-94-92671-16-5.
Parker C, Castro E, Fizazi K et al (2020) Prostate cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 31:1119–1134. https://doi.org/10.1016/j.annonc.2020.06.011
de Rooij M, Hamoen EHJ, Witjes JA et al (2016) Accuracy of magnetic resonance imaging for local staging of prostate cancer: a diagnostic meta-analysis. Eur Urol 70:233–245. https://doi.org/10.1016/j.eururo.2015.07.029
Zelic R, Garmo H, Zugna D et al (2020) Predicting prostate cancer death with different pretreatment risk stratification tools: a head-to-head comparison in a nationwide cohort study. Eur Urol 77:180–188. https://doi.org/10.1016/j.eururo.2019.09.027
Alves JR, Muglia VF, Lucchesi FR et al (2020) Independent external validation of nomogram to predict extracapsular extension in patients with prostate cancer. Eur Radiol 30:5004–5010. https://doi.org/10.1007/s00330-020-06839-0
Diamand R, Ploussard G, Roumiguié M et al (2021) External validation of a multiparametric magnetic resonance imaging–based nomogram for the prediction of extracapsular extension and seminal vesicle invasion in prostate cancer patients undergoing radical prostatectomy. Eur Urol 79:180–185. https://doi.org/10.1016/j.eururo.2020.09.037
Bai H, Xia W, Ji X et al (2021) Multiparametric magnetic resonance imaging-based peritumoral radiomics for preoperative prediction of the presence of extracapsular extension with prostate cancer. J Magn Reson Imaging 54:1222–1230. https://doi.org/10.1002/jmri.27678
Cuocolo R, Stanzione A, Faletti R et al (2021) MRI index lesion radiomics and machine learning for detection of extraprostatic extension of disease: a multicenter study. Eur Radiol 31:7575–7583. https://doi.org/10.1007/s00330-021-07856-3
Damascelli A, Gallivanone F, Cristel G et al (2021) Advanced imaging analysis in prostate MRI: building a radiomic signature to predict tumor aggressiveness. Diagnostics 11:594. https://doi.org/10.3390/diagnostics11040594
Fan X, Xie N, Chen J et al (2022) Multiparametric MRI and machine learning based radiomic models for preoperative prediction of multiple biological characteristics in prostate cancer. Front Oncol 12:839621. https://doi.org/10.3389/fonc.2022.839621
He D, Wang X, Fu C et al (2021) MRI-based radiomics models to assess prostate cancer, extracapsular extension and positive surgical margins. Cancer Imaging 21:46. https://doi.org/10.1186/s40644-021-00414-6
Hou Y, Zhang Y-H, Bao J et al (2021) Artificial intelligence is a promising prospect for the detection of prostate cancer extracapsular extension with mpMRI: a two-center comparative study. Eur J Nucl Med Mol Imaging 48:3805–3816. https://doi.org/10.1007/s00259-021-05381-5
Losnegård A, Reisæter LAR, Halvorsen OJ et al (2020) Magnetic resonance radiomics for prediction of extraprostatic extension in non-favorable intermediate- and high-risk prostate cancer patients. Acta Radiol 61:1570–1579. https://doi.org/10.1177/0284185120905066
Ma S, Xie H, Wang H et al (2019) MRI-based radiomics signature for the preoperative prediction of extracapsular extension of prostate cancer. J Magn Reson Imaging 50:1914–1925. https://doi.org/10.1002/jmri.26777
Ma S, Xie H, Wang H et al (2020) Preoperative prediction of extracapsular extension: radiomics signature based on magnetic resonance imaging to stage prostate cancer. Mol Imaging Biol 22:711–721. https://doi.org/10.1007/s11307-019-01405-7
Moroianu ŞL, Bhattacharya I, Seetharaman A et al (2022) Computational detection of extraprostatic extension of prostate cancer on multiparametric MRI using deep learning. Cancers (Basel) 14:2821. https://doi.org/10.3390/cancers14122821
Shiradkar R, Zuo R, Mahran A et al (2020) Radiomic features derived from periprostatic fat on pre-surgical T2w MRI predict extraprostatic extension of prostate cancer identified on post-surgical pathology: preliminary results. In: Hahn HK, Mazurowski MA (eds) Medical imaging 2020: computer-aided diagnosis. SPIE, p 121
Stanzione A, Cuocolo R, Cocozza S et al (2019) Detection of extraprostatic extension of cancer on biparametric MRI combining texture analysis and machine learning: preliminary results. Acad Radiol 26:1338–1344. https://doi.org/10.1016/j.acra.2018.12.025
Xu L, Zhang G, Zhao L et al (2020) Radiomics based on multiparametric magnetic resonance imaging to predict extraprostatic extension of prostate cancer. Front Oncol 10:40. https://doi.org/10.3389/fonc.2020.00940
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Kapoor S, Narayanan A (2022) Leakage and the reproducibility crisis in ML-based science. https://doi.org/10.48550/arXiv.2207.07048
Pinto dos Santos D, Dietzel M, Baessler B (2021) A decade of radiomics research: are images really data or just patterns in the noise? Eur Radiol 31:1–4. https://doi.org/10.1007/s00330-020-07108-w
Marcadent S, Hofmeister J, Preti MG et al (2020) Generative adversarial networks improve the reproducibility and discriminative power of radiomic features. Radiol Artif Intell 2:e190035. https://doi.org/10.1148/ryai.2020190035
Alderson PO (2020) The quest for generalizability in radiomics. Radiol Artif Intell 2:e200068. https://doi.org/10.1148/ryai.2020200068
Koçak B, Cuocolo R, dos Santos DP et al (2023) Must-have qualities of clinical research on artificial intelligence and machine learning. Balkan Med J 40:3–12. https://doi.org/10.4274/balkanmedj.galenos.2022.2022-11-51
Moher D, Liberati A, Tetzlaff J et al (2009) Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. BMJ 339:b2535. https://doi.org/10.1136/bmj.b2535
PROSPERO: International Prospective Register of Systematic Reviews. https://www.crd.york.ac.uk/prospero/
Whiting PF (2011) QUADAS-2: a revised tool for the Quality Assessment of Diagnostic Accuracy Studies. Ann Intern Med 155:529. https://doi.org/10.7326/0003-4819-155-8-201110180-00009
Lambin P, Leijenaar RTH, Deist TM et al (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14:749–762. https://doi.org/10.1038/nrclinonc.2017.141
Ponsiglione A, Stanzione A, Spadarella G et al (2022) Ovarian imaging radiomics quality score assessment: an EuSoMII radiomics auditing group initiative. Eur Radiol 33:2239–2247. https://doi.org/10.1007/s00330-022-09180-w
Zhong J, Hu Y, Si L et al (2021) A systematic review of radiomics in osteosarcoma: utilizing radiomics quality score as a tool promoting clinical translation. Eur Radiol 31:1526–1535. https://doi.org/10.1007/s00330-020-07221-w
Egger M, Smith GD, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. BMJ 315:629–634. https://doi.org/10.1136/bmj.315.7109.629
Song J, Yin Y, Wang H et al (2020) A review of original articles published in the emerging field of radiomics. Eur J Radiol 127:108991. https://doi.org/10.1016/j.ejrad.2020.108991
Stanzione A, Ponsiglione A, Alessandrino F et al (2023) Beyond diagnosis: is there a role for radiomics in prostate cancer management? Eur Radiol Exp 7:13. https://doi.org/10.1186/s41747-023-00321-4
van Timmeren JE, Cester D, Tanadini-Lang S et al (2020) Radiomics in medical imaging—“how-to” guide and critical reflection. Insights Imaging 11:91. https://doi.org/10.1186/s13244-020-00887-2
Zwanenburg A, Vallières M, Abdalah MA et al (2020) The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping. Radiology 295:328–338. https://doi.org/10.1148/radiol.2020191145
Bai K, Sun Y, Li W, Zhang L (2019) Apparent diffusion coefficient in extraprostatic extension of prostate cancer: a systematic review and diagnostic meta-analysis. Cancer Manag Res 11:3125–3137. https://doi.org/10.2147/CMAR.S191738
Guiot J, Vaidyanathan A, Deprez L et al (2022) A review in radiomics: making personalized medicine a reality via routine imaging. Med Res Rev 42:426–440. https://doi.org/10.1002/med.21846
Mongan J, Moy L, Kahn CE (2020) Checklist for Artificial Intelligence in Medical Imaging (CLAIM): a guide for authors and reviewers. Radiol Artif Intell 2:e200029. https://doi.org/10.1148/ryai.2020200029
Kocak B, Baessler B, Bakas S et al (2023) CheckList for EvaluAtion of Radiomics research (CLEAR): a step-by-step reporting guideline for authors and reviewers endorsed by ESR and EuSoMII. Insights Imaging 14:75. https://doi.org/10.1186/s13244-023-01415-8
Calimano-Ramirez LF, Virarkar MK, Hernandez M et al (2023) MRI-based nomograms and radiomics in presurgical prediction of extraprostatic extension in prostate cancer: a systematic review. Abdominal Radiol (NY) 48:2379–2400. https://doi.org/10.1007/s00261-023-03924-y
Eifler JB, Feng Z, Lin BM et al (2013) An updated prostate cancer staging nomogram (Partin tables) based on cases from 2006 to 2011. BJU Int 111:22–29. https://doi.org/10.1111/j.1464-410X.2012.11324.x
Ohori M, Kattan MW, Koh H et al (2004) Predicting the presence and side of extracapsular extension: a nomogram for staging prostate cancer. J Urol 171:1844–9; discussion 1849. https://doi.org/10.1097/01.ju.0000121693.05077.3d
Li W, Shang W, Lu F et al (2022) Diagnostic performance of extraprostatic extension grading system for detection of extraprostatic extension in prostate cancer: a diagnostic systematic review and meta-analysis. Front Oncol 11:792120. https://doi.org/10.3389/fonc.2021.792120
Li W, Dong A, Hong G et al (2021) Diagnostic performance of ESUR scoring system for extraprostatic prostate cancer extension: a meta-analysis. Eur J Radiol 143:109896. https://doi.org/10.1016/j.ejrad.2021.109896
Mehralivand S, Shih JH, Harmon S et al (2019) A grading system for the assessment of risk of extraprostatic extension of prostate cancer at multiparametric MRI. Radiology 290:709–719. https://doi.org/10.1148/radiol.2018181278
Steyerberg EW, Vickers AJ, Cook NR et al (2010) Assessing the performance of prediction models. Epidemiology 21:128–138. https://doi.org/10.1097/EDE.0b013e3181c30fb2
Stanzione A, Cuocolo R, Ugga L et al (2022) Oncologic imaging and radiomics: a walkthrough review of methodological challenges. Cancers (Basel) 14:4871. https://doi.org/10.3390/cancers14194871
Moons KGM, Altman DG, Reitsma JB et al (2015) Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis (TRIPOD): explanation and elaboration. Ann Intern Med 162:W1–W73. https://doi.org/10.7326/M14-0698
Cuocolo R, Cipullo MB, Stanzione A et al (2020) Machine learning for the identification of clinically significant prostate cancer on MRI: a meta-analysis. Eur Radiol 30:6877–6887. https://doi.org/10.1007/s00330-020-07027-w
Cronin P, Kelly AM, Altaee D et al (2018) How to perform a systematic review and meta-analysis of diagnostic imaging studies. Acad Radiol 25:573–593. https://doi.org/10.1016/j.acra.2017.12.007
Adams J, Hillier-Brown FC, Moore HJ et al (2016) Searching and synthesising ‘grey literature’ and ‘grey information’ in public health: critical reflections on three case studies. Syst Rev 5:164. https://doi.org/10.1186/s13643-016-0337-y
Funding
Open access funding provided by Università degli Studi di Napoli Federico II within the CRUI-CARE Agreement. The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Professor Massimo Imbriaco, MD.
Conflict of interest
Arnaldo Stanzione and Renato Cuocolo serve as Editorial Board Members of European Radiology. They have not taken part in the review or selection process of this article.
Andrea Ponsiglione serves as Editorial Board Member of Insights into Imaging. Andrea Ponsiglione is junior deputy editor in European Radiology. He has not taken part in the review or selection process of this article.
Renato Cuocolo serves as Editorial Board Member of European Radiology Experimental. The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was not required for this study because it is a systematic literature review and meta-analysis.
Ethical approval
Institutional Review Board approval was not required because it is a systematic literature review and meta-analysis.
Study subjects or cohorts overlap
Not applicable
Methodology
• Systematic review and meta-analysis
• Performed at one institution
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ponsiglione, A., Gambardella, M., Stanzione, A. et al. Radiomics for the identification of extraprostatic extension with prostate MRI: a systematic review and meta-analysis. Eur Radiol 34, 3981–3991 (2024). https://doi.org/10.1007/s00330-023-10427-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-10427-3