Abstract
Objectives
We evaluate MR radiomics and develop machine learning–based classifiers to predict MYCN amplification in neuroblastomas.
Methods
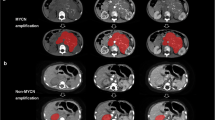
A total of 120 patients with neuroblastomas and baseline MR imaging examination available were identified of whom 74 (mean age ± standard deviation [SD] of 6 years and 2 months ± 4 years and 9 months; 43 females and 31 males, 14 MYCN amplified) underwent imaging at our institution. This was therefore used to develop radiomics models. The model was tested in a cohort of children with the same diagnosis but imaged elsewhere (n = 46, mean age ± SD: 5 years 11 months ± 3 years 9 months, 26 females and 14 MYCN amplified). Whole tumour volumes of interest were adopted to extract first-order histogram and second-order radiomics features. Interclass correlation coefficient and maximum relevance and minimum redundancy algorithm were applied for feature selection. Logistic regression, support vector machine, and random forest were employed as the classifiers. Receiver operating characteristic (ROC) analysis was performed to evaluate the diagnostic accuracy of the classifiers on the external test set.
Results
The logistic regression model and the random forest both showed an AUC of 0.75. The support vector machine classifier obtained an AUC of 0.78 on the test set with a sensitivity of 64% and a specificity of 72%.
Conclusion
The study provides preliminary retrospective evidence demonstrating the feasibility of MRI radiomics in predicting MYCN amplification in neuroblastomas. Future studies are needed to explore the correlation between other imaging features and genetic markers and to develop multiclass predictive models.
Key Points
• MYCN amplification in neuroblastomas is an important determinant of disease prognosis.
• Radiomics analysis of pre-treatment MR examinations can be used to predict MYCN amplification in neuroblastomas.
• Radiomics machine learning models showed good generalisability to external test set, demonstrating reproducibility of the computational models.




Similar content being viewed by others
Abbreviations
- AUC-ROC:
-
Area under the receiver operator curve
- CT:
-
Computed tomography
- FST1:
-
(Non-contrast) fat-saturated T1-weighted images
- ICC:
-
Interclass correlation
- LR:
-
Logistic regression
- MRI:
-
Magnetic resonance imaging
- NFST2:
-
Non-fat-saturated T2-weighted images
- PACS:
-
Picture Archiving and Communications System
- ROI:
-
Region of interest
- SVM:
-
Support vector machine
- VOI:
-
Volume of interest
References
Li J, Thompson TD, Miller JW et al (2008) Cancer incidence among children and adolescents in the United States, 2001–2003. Pediatrics 121:e1470–e1477. https://doi.org/10.1542/peds.2007-2964
Nakagawara A, Li Y, Izumi H et al (2018) Neuroblastoma. Jpn J Clin Oncol 48:214–241. https://doi.org/10.1093/jjco/hyx176
Brodeur GM (2018) Spontaneous regression of neuroblastoma. Cell Tissue Res 372:277–286. https://doi.org/10.1007/s00441-017-2761-2
London WB, Castleberry RP, Matthay KK et al (2005) Evidence for an age cutoff greater than 365 days for neuroblastoma risk group stratification in the Children’s Oncology Group. J Clin Oncol 23:6459–6465. https://doi.org/10.1200/JCO.2005.05.571
Cohn SL, Pearson ADJ, London WB et al (2009) The International Neuroblastoma Risk Group (INRG) classification system: an INRG Task Force report. J Clin Oncol 27:289–297. https://doi.org/10.1200/JCO.2008.16.6785
Monclair T, Brodeur GM, Ambros PF et al (2009) The International Neuroblastoma Risk Group (INRG) staging system: an INRG Task Force report. J Clin Oncol 27:298–303. https://doi.org/10.1200/JCO.2008.16.6876
Ruiz-Pérez MV, Henley AB, Arsenian-Henriksson M (2017) The MYCN protein in health and disease. Genes 8(4):113. https://doi.org/10.3390/genes8040113
Tan E, Merchant K, Kn BP et al (2022) CT-based morphologic and radiomics features for the classification of MYCN gene amplification status in pediatric neuroblastoma. Childs Nerv Syst 38:1487–1495. https://doi.org/10.1007/s00381-022-05534-3
Chen X, Wang H, Huang K et al (2021) CT-based radiomics signature with machine learning predicts MYCN amplification in pediatric abdominal neuroblastoma. Front Oncol 11:687884. https://doi.org/10.3389/fonc.2021.687884
Wu H, Wu C, Zheng H et al (2021) Radiogenomics of neuroblastoma in pediatric patients: CT-based radiomics signature in predicting MYCN amplification. Eur Radiol 31:3080–3089. https://doi.org/10.1007/s00330-020-07246-1
Di Giannatale A, Di Paolo PL, Curione D et al (2021) Radiogenomics prediction for MYCN amplification in neuroblastoma: a hypothesis generating study. Pediatr Blood Cancer 68:e29110. https://doi.org/10.1002/pbc.29110
Ambros PF, Ambros IM, Brodeur GM et al (2009) International consensus for neuroblastoma molecular diagnostics: report from the International Neuroblastoma Risk Group (INRG) Biology Committee. Br J Cancer 100:1471–1482. https://doi.org/10.1038/sj.bjc.6605014
Theissen J, Boensch M, Spitz R et al (2009) Heterogeneity of the MYCN oncogene in neuroblastoma. Clin Cancer Res 15:2085–2090. https://doi.org/10.1158/1078-0432.CCR-08-1648
Qian L, Yang S, Zhang S et al (2022) Prediction of MYCN amplification, 1p and 11q aberrations in pediatric neuroblastoma via pre-therapy 18F-FDG PET/CT radiomics. Front Med (Lausanne) 9:840777. https://doi.org/10.3389/fmed.2022.840777
Feng L, Qian L, Yang S et al (2022) Prediction for mitosis-karyorrhexis index status of pediatric neuroblastoma via machine learning based 18F-FDG PET/CT radiomics. Diagnostics 12(2):262. https://doi.org/10.3390/diagnostics12020262
Feng L, Qian L, Yang S et al (2022) Clinical parameters combined with radiomics features of PET/CT can predict recurrence in patients with high-risk pediatric neuroblastoma. BMC Med Imaging 22:102. https://doi.org/10.1186/s12880-022-00828-z
Liu G, Poon M, Zapala MA et al (2022) Incorporating radiomics into machine learning models to predict outcomes of neuroblastoma. J Digit Imaging 35:605–612. https://doi.org/10.1007/s10278-022-00607-w
Sun K, Zhu H, Chai W, Yan F (2022) TP53 mutation estimation based on MRI radiomics analysis for breast cancer. J Magn Reson Imaging. https://doi.org/10.1002/jmri.28323
Liu Y-F, Shu X, Qiao X-F et al (2022) Radiomics-based machine learning models for predicting P504s/P63 immunohistochemical expression: a noninvasive diagnostic tool for prostate cancer. Front Oncol 12:911426. https://doi.org/10.3389/fonc.2022.911426
Meißner A-K, Gutsche R, Galldiks N et al (2021) Radiomics for the non-invasive prediction of the BRAF mutation status in patients with melanoma brain metastases. Neuro Oncol. https://doi.org/10.1093/neuonc/noab294
Lee J, Steinmann A, Ding Y et al (2021) Radiomics feature robustness as measured using an MRI phantom. Sci Rep 11:3973. https://doi.org/10.1038/s41598-021-83593-3
Bernatz S, Zhdanovich Y, Ackermann J et al (2021) Impact of rescanning and repositioning on radiomic features employing a multi-object phantom in magnetic resonance imaging. Sci Rep 11:14248. https://doi.org/10.1038/s41598-021-93756-x
Saltybaeva N, Tanadini-Lang S, Vuong D et al (2022) Robustness of radiomic features in magnetic resonance imaging for patients with glioblastoma: multi-center study. Phys Imaging Radiat Oncol 22:131–136. https://doi.org/10.1016/j.phro.2022.05.006
Crombé A, Kind M, Fadli D et al (2020) Intensity harmonization techniques influence radiomics features and radiomics-based predictions in sarcoma patients. Sci Rep 10:15496. https://doi.org/10.1038/s41598-020-72535-0
Nyúl LG, Udupa JK, Zhang X (2000) New variants of a method of MRI scale standardization. IEEE Trans Med Imaging 19:143–150. https://doi.org/10.1109/42.836373
Davatzikos C, Rathore S, Bakas S et al (2018) Cancer imaging phenomics toolkit: quantitative imaging analytics for precision diagnostics and predictive modeling of clinical outcome. J Med Imaging (Bellingham) 5:011018. https://doi.org/10.1117/1.JMI.5.1.011018
Pati S, Singh A, Rathore S et al (2020) The cancer imaging phenomics toolkit (captk): technical overview. Brainlesion 11993:380–394. https://doi.org/10.1007/978-3-030-46643-5_38
Diciotti S, Ciulli S, Mascalchi M et al (2013) The “peeking” effect in supervised feature selection on diffusion tensor imaging data. AJNR Am J Neuroradiol 34:E107–E107. https://doi.org/10.3174/ajnr.A3685
An Introduction to ‘glmnet’ • glmnet. https://glmnet.stanford.edu/articles/glmnet.html. Accessed 6 Dec 2022
Ghosh A, Yekeler E, Dalal D et al (2022) Whole-tumour apparent diffusion coefficient (ADC) histogram analysis to identify MYCN-amplification in neuroblastomas: preliminary results. Eur Radiol 32:8453–8462. https://doi.org/10.1007/s00330-022-08750-2
Huang M, Weiss WA (2013) Neuroblastoma and MYCN. Cold Spring Harb Perspect Med 3:a014415. https://doi.org/10.1101/cshperspect.a014415
Lee JW, Son MH, Cho HW et al (2018) Clinical significance of MYCN amplification in patients with high-risk neuroblastoma. Pediatr Blood Cancer 65:e27257. https://doi.org/10.1002/pbc.27257
Brisse HJ, Blanc T, Schleiermacher G et al (2017) Radiogenomics of neuroblastomas: relationships between imaging phenotypes, tumor genomic profile and survival. PLoS ONE 12:e0185190. https://doi.org/10.1371/journal.pone.0185190
Shur JD, Doran SJ, Kumar S et al (2021) Radiomics in oncology: a practical guide. Radiographics 41:1717–1732. https://doi.org/10.1148/rg.2021210037
Rožanec J, Poštuvan T, Fortuna B, Mladenić D (2021) Tackling class imbalance in radiomics: the COVID-19 use case. In Ljubljana ’21: Slovenian KDD Conference on Data Mining and Data Warehouses, October, 2021, Ljubljana, Slovenia. ACM, New York, NY, USA, p 4. https://ailab.ijs.si/dunja/SiKDD2021/Papers/Ro%C5%BEanecetal.pdf
Vabalas A, Gowen E, Poliakoff E, Casson AJ (2019) Machine learning algorithm validation with a limited sample size. PLoS ONE 14:e0224365. https://doi.org/10.1371/journal.pone.0224365
Funding
The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Dr Lisa States.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
One of the authors has significant statistical expertise.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
A part of the data (n = 77) has been reported in the manuscript: Whole-tumour apparent diffusion coefficient (ADC) histogram analysis to identify MYCN amplification in neuroblastomas: preliminary results https://doi.org/10.1007/s00330-022-08750-2
Methodology
• retrospective
• diagnostic
• performed at one institution, but data obtained retrospectively from multiple sites
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ghosh, A., Yekeler, E., Teixeira, S.R. et al. Role of MRI radiomics for the prediction of MYCN amplification in neuroblastomas. Eur Radiol 33, 6726–6735 (2023). https://doi.org/10.1007/s00330-023-09628-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-09628-7