Abstract
Key message
We have developed and validated an efficient protocol for producing gene-edited carrot plants that do not result in the stable incorporation of foreign DNA in the edited plant’s genome.
Abstract
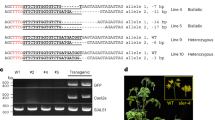
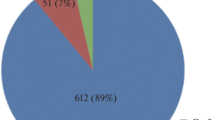
We report here a method for producing transgene-free, gene-edited carrot (Daucus carota subs. sativus) plants. With this approach, PEG-mediated transformation is used to transiently express a cytosine base editor and a guide RNA in protoplasts to induce targeted mutations in the carrot genome. These protoplasts are then cultured under conditions that lead to the production of somatic embryos which subsequently develop into carrot plants. For this study, we used the Centromere-Specific Histone H3 (CENH3) gene as a target for evaluating the efficiency with which regenerated, edited plants could be produced. After validating sgRNA performance and protoplast transformation efficiency using transient assays, we performed two independent editing experiments using sgRNAs targeting different locations within CENH3. In the first experiment, we analyzed 184 regenerated plants and found that 22 of them (11.9%) carried targeted mutations within CENH3, while in the second experiment, 28 out of 190 (14.7%) plants had mutations in CENH3. Of the 50 edited carrot lines that we analyzed, 43 were homozygous or bi-allelic for mutations in CENH3. No evidence of the base editor expression plasmid was found in the edited lines tested, indicating that this approach is able to produce transgene-free, gene-edited lines. The protocol that we describe provides an efficient method for easily generating large numbers of transgene-free, gene-edited carrot plants.






Similar content being viewed by others
Data availability
The data underlying this article are available in the article and in its online supplementary material.
References
Ahn MJ, Noah SA, Ha SH, Back K, Lee SW, Bae JM (2012) Production of ketocarotenoids in transgenic carrot plants with an enhanced level of β-carotene. Plant Biotechnol Rep 6:133–140. https://doi.org/10.1007/s11816-011-0206-z
Andersson M, Turesson H, Nicolia A, Fält AS, Samuelsson M, Hofvander P (2017) Efficient targeted multiallelic mutagenesis in tetraploid potato (Solanum tuberosum) by transient CRISPR-Cas9 expression in protoplasts. Plant Cell Rep 36:117–128. https://doi.org/10.1007/s00299-016-2062-3
Annon A, Rathore K, Crosby K (2014) Overexpression of a tobacco osmotin gene in carrot (Daucus carota L.) enhances drought tolerance. In Vitro Cell Dev Biol Plant 50:299–306. https://doi.org/10.1007/s11627-013-9590-0
Baranski R, Klocke E, Ryschka U (2007) Monitoring the expression of green fluorescent protein in carrot. Acta Physiol Plant 29:239–246. https://doi.org/10.1007/s11738-007-0030-9
Cheng H, Hao M, Ding B, Mei D, Wang W, Wang H et al (2021) Base editing with high efficiency in allotetraploid oilseed rape by A3A-PBE system. Plant Biotechnol J 19(1):87–97. https://doi.org/10.1111/pbi.13444
Clasen B, Stoddard T, Luo S, Demorest Z, Li J, Cedrone F et al (2016) Improving cold storage and processing traits in potato through targeted gene knockout. Plant Biotechnol J 14(1):169–176. https://doi.org/10.1111/pbi.12370
De Bruyn C, Ruttink T, Eeckhaut T, Jacobs T, De Keyser E, Goossens A et al (2020) Establishment of CRISPR/Cas9 genome editing in witloof (Cichorium intybus var. foliosum). Front Genome Ed 2:604876. https://doi.org/10.3389/fgeed.2020.604876
Dirks R, Sidorov V, Tulmans C (1996) A new protoplast culture system in Daucus carota L. and its applications for mutant selection and transformation. Theor Appl Genet 93:809–815. https://doi.org/10.1007/BF00224080
Eeckhaut T, Lakshmanan PS, Deryckere D, Bockstaele EV, Huylenbroeck JV (2013) Progress in plant protoplast research. Plants 238:991–1003. https://doi.org/10.1007/s00425-013-1936-7
Fossi M, Amundson K, Kuppu S, Britt A, Comai L (2019) Regeneration of Solanum tuberosum plants from protoplasts induces widespread genome instability. Plant Physiol 180:78–86. https://doi.org/10.1104/pp.18.00906
Gautheret R (1939) Sur la possibilité de réaliser la culture indéfinie des tissues de tubercules de carotte. C R Soc Biol Paris 208:118–120
Grzebelus E, Szklarczyk M, Baranski R (2012) An improved protocol for plant regeneration from leaf- and hypocotyl-derived protoplasts of carrot. PCTOC 109:101–109. https://doi.org/10.1007/s11240-011-0078-5
Grzebelus E, Kruk M, Macko-Podgórni A, Grzebelus D (2013) Response of carrot protoplasts and protoplast-derived aggregates to selection using a fungal culture filtrate of Alternaria radicina. Plant Cell Tissue Organ Cult 115:209–222
Guo W, Duan Y, Olivares-Fuster O, Wu Z, Arias CR, Burns J et al (2005) Protoplast transformation and regeneration of transgenic valencia sweet orange plants containing a juice quality-related pectin methylesterase gene. Plant Cell Rep 24:482–486. https://doi.org/10.1007/s00299-005-0952-x
Henikoff S, Dalal Y (2005) Centromeric chromatin: what makes it unique? Curr Opin Genet Dev 15:177–184. https://doi.org/10.1016/j.gde.2005.01.004
Hsu CT, Cheng YJ, Yuan YH, Hung WF, Cheng QW, Wu FH et al (2019) Application of Cas12a and nCas9-activation-induced cytidine deaminase for genome editing and as a non-sexual strategy to generate homozygous/multiplex editing plants in the allotetraploid genome of tobacco. Plant Mol Biol 101(4–5):355–371. https://doi.org/10.1007/s11103-019-00907-w
Hsu CT, Lee WC, Cheng YJ, Yuan YH, Wu FH, Lin CS (2021) Genome editing and protoplast regeneration to study plant-pathogen interactions in the model plant Nicotiana bentamiana. Front Genome Ed 2:627803. https://doi.org/10.3389/fgeed.2020.627803
Huang TK, Puchta H (2019) CRISPR/Cas-mediated gene targeting in plants: finally a turn for the better for homologous recombination. Plant Cell Rep 38:443–453. https://doi.org/10.1007/s00299-019-02379-0
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna J (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 17:816–821. https://doi.org/10.1126/science.1225829
Klimek-Chodacka M, Oleszkiewicz T, Lowder LG, Qi Y, Baranski R (2018) Efficient CRISPR/Cas9-based genome editing in carrot cells. Plant Cell Rep 37:575–586. https://doi.org/10.1007/s00299-018-2252-2
Klimek-Chodacka M, Oleszkiewicz T, Baranski R (2019) Visual assay for gene editing using a CRISPR/Cas9 system in carrot cells. In: Qi Y (ed) Plant Genome Editing with CRISPR systems. Humana Press, New Jersey
Kluesner M, Nedveck DA, Lahr WS, Garbe JR, Abrahante JE, Webber BR et al (2018) EditR: a method to quantify base editing from Sanger sequencing. CRISPR J 1:3. https://doi.org/10.1089/crispr.2018.0014
Kuppu S, Tan EK, Nguyen H, Rodger A, Comai L, Chan SW et al (2015) Point mutations in centromeric histone induce post-zygotic incompatibility and uniparental inheritance. PLoS Genet 11(9):e1005494. https://doi.org/10.1371/journal.pgen.1005494
Li JF, Norville JE, Aach J, Mccormack M, Zhang D, Bush J et al (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31:688–691. https://doi.org/10.1038/nbt.2654
Li J, Sun Y, Du J, Zhao Y, Xia L (2017) Generation of targeted point mutations in rice by a modified CRISPR/Cas9 system. Mol Plant 10:526–529. https://doi.org/10.1016/j.molp.2016.12.001
Lin CS, Hsu CT, Yang LH, Lee LY, Fu J-Y, Cheng Q-W et al (2018) Application of protoplast technology to CRISPR/Cas9 mutagenesis: from single-cell mutation section to mutant plant regeneration. Plant Biotechnol J 16(7):1295–1310. https://doi.org/10.1111/pbi.12870
Modrzejewski D, Hartung F, Lehnert H, Sprink T, Kohl C, Keilwagen J et al (2020) Which factors affect the occurrence of off-target effects caused by the use of CRISPR/Cas: a systematic review in plants. Front Plant Sci 11:574959. https://doi.org/10.3389/fpls.2020.574959
Nadakuduti SS, Enciso-Rodriguez F (2021) Advances in genome editing with CRISPR systems and transformation technologies for plant DNA manipulation. Front Plant Sci 11:637159. https://doi.org/10.3389/fpls.2020.637159
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31:691–693. https://doi.org/10.1038/nbt.2655
Nobécourt, P. (1939) Sur la pérennité et l'augmentation de volume des cultures de tissues végétaux. Compt. Rendus Soc. Biol. Lyon 130:1270–1271
Oleszkiewicz T, Klimek-Chodacka M, Kruczek M, Godel-Jedrychowska K, Sala K, Milewska-Hendel A et al (2021) Inhibition of carotenoid biosynthesis by CRISPR/Cas9 triggers cell wall remodeling in carrot. Int J Mol Sci 22:6516. https://doi.org/10.3390/ijms22126516
Osakabe Y, Liang Z, Ren C, Nishitani C, Osakabe K, Wada M et al (2018) CRISPR-Cas9-mediated genome editing in apple and grapevine. Nat Protoc 13:2844–2863. https://doi.org/10.1038/s41596-018-0067-9
Rees H, Liu D (2018) Base editing: precision chemistry on the genome and transcriptome of living cells. Nat Rev 19:770–778. https://doi.org/10.1038/s41576-018-0059-1
Rojas-Anaya E, Loza-Rubio E, Olivera-Flores MT, GomezLim M (2009) Expression of rabies virus G protein in carrots (Daucus carota). Transgenic Res 18:911–919. https://doi.org/10.1007/s11248-009-9278-8
Scott RJ, Draper J (1987) Transformation of carrot tissues derived from pro- embryogenic suspension cells: a useful model system for gene expression studies in plants. Plant Mol Biol 8:265–274
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z et al (2013) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31:686–688. https://doi.org/10.1038/nbt.2650
Simon P (2000) Domestication, historical development, and modern breeding of carrot. Plant Breed Rev 19:157–189. https://doi.org/10.1002/9780470650172.ch5
Tang Xu, Ren Q, Yang L, Bao Y, Zhong Z, He Y et al (2019) Single transcript unit CRISPR 2.0 systems for robust Cas9 and Cas12a mediated plant genome editing. Plant Biotechnol J 17(7):1431–1445. https://doi.org/10.1111/pbi.13068
Thomas JC, Guiltinan MJ, Bustos S, Thomas T, Nessler C (1989) Carrot (Daucus carota) hypocotyls transformation using Agrobacterium tumefaciens. Plant Cell Rep 8:354–357. https://doi.org/10.1007/BF00716672
Wally O, Punja Z (2010) Enhanced disease resistance in transgenic carrot (Daucus carota L.) plants over-expressing a rice cationic peroxidase. Planta 232:1229–1239. https://doi.org/10.1007/s00425-010-1252-4
Woo JW, Kim J, Kwon S, Corvalàn C, Cho SW, Kim H et al (2015) DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33:1162–1164. https://doi.org/10.1038/nbt.3389
Wu S, Zhu H, Liu J, Yang Q, Shao X, Bi F et al (2020) Establishment of a PEG-mediated protoplast transformation system based on DNA and CRISPR/Cas9 ribonucleoprotein complexes for banana. BMB Plant Biol 20:425. https://doi.org/10.1186/s12870-020-02609-8
Xu ZS, Feng K, Xiong AS (2019a) CRISPR/Cas9-mediated multiply targeted mutagenesis in orange and purple carrot plants. Mol Biotech 61:191–199. https://doi.org/10.1007/s12033-018-00150-6
Xu ZS, Yang QQ, Feng K, Xiong AS (2019b) Changing carrot color: insertions in DcMYB7 alter the regulation of anthocyanin biosynthesis and modification. Plant Physiol 181:195–207. https://doi.org/10.1104/pp.19.00523
Yu J, Tu L, Subburaj S, Bae S, Lee S (2021) Simultaneous targeting of duplicated genes in petunia protoplasts for flower color modifications via CRISPR-Cas9 ribonucleoproteins. Plant Cell Rep 40:1037–1045. https://doi.org/10.1007/s00299-020-02593-1
Zaman N, Seitz K, Kabir M, George-Schreder L, Shepstone I, Liu YL (2018) A förster resonance energy transfer sensor for live-cell imaging of mitogen-activated protein kinase activity in Arabidopsis. Plant J 97:970–983. https://doi.org/10.1111/tpj.14164
Zong Y, Song Q, Chao L, Jin S, Zhang D, Wang Y et al (2018) Efficient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nat Biotech 36:950–953. https://doi.org/10.1038/nbt.4261
Acknowledgements
We thank Aneta Malec for instruction in the method of carrot protoplast regeneration. We thank Phillip Simon and Richard Amasino for their assistance with vernalization of the mutant carrot plants.
Funding
This work was supported by the AFRI Exploratory Research Program [GRANT12854173 /project accession no. 1021280] from the USDA National Institute of Food and Agriculture, United States of America.
Author information
Authors and Affiliations
Contributions
All authors participated in planning the experiments. All authors provided critical feedback and helped shape the research, analysis and manuscript. CMM carried out the majority of the experiments with PJK also contributing. CMM collected and analyzed the data. CMM produced the figures for the manuscript with input from PJK. CMM took the lead in writing the manuscript with input from PJK. All authors participated in review and revision of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Zhanyuan Jon Zhang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Meyer, C.M., Goldman, I.L., Grzebelus, E. et al. Efficient production of transgene-free, gene-edited carrot plants via protoplast transformation. Plant Cell Rep 41, 947–960 (2022). https://doi.org/10.1007/s00299-022-02830-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-022-02830-9