Abstract
Key message
A sodium hydrogen exchanger (NHX) gene from the date palm enhances tolerance to salinity in Arabidopsis plants.
Abstract
Plant sodium hydrogen exchangers/antiporters (NHXs) are pivotal regulators of intracellular Na+/K+ and pH homeostasis, which is essential for salt stress adaptation. In this study, a novel orthologue of Na+/H+ antiporter was isolated from date palm (PdNHX6) and functionally characterized in mutant yeast cells and Arabidopsis plants to assess the behavior of the transgenic organisms in response to salinity. Genetically transformed yeast cells with PdNHX6 were sensitive to salt stress when compared to the empty vector (EV) yeast cells. Besides, the acidity value of the vacuoles of the transformant yeast cells has significantly (p ≤ 0.05) increased, as indicated by the calibrated fluorescence intensity measurements and the fluorescence imagining analyses. This observation supports the notion that PdNHX6 might regulate proton pumping into the vacuole, a crucial salt tolerance mechanism in the plants. Consistently, the transient overexpression and subcellular localization revealed the accumulation of PdNHX6 in the tonoplast surrounding the central vacuole of Nicotiana benthamiana leaf epidermal cells. Stable overexpression of PdNHX6 in Arabidopsis plants enhanced tolerance to salt stress and retained significantly higher chlorophyll, water contents, and increased seed germination under salinity when compared to the wild-type plants. Despite the significant increase of Na+, transgenic Arabidopsis lines maintained a balanced Na+/K+ ratio under salt stress conditions. Together, the results obtained from this study imply that PdNHX6 is involved in the salt tolerance mechanism in plants by controlling K+ and pH homeostasis of the vacuoles.











Similar content being viewed by others
References
Al-Harrasi I, Al-Yahyai R, Yaish MW (2018) Differential DNA methylation and transcription profiles in date palm roots exposed to salinity. PLoS ONE 13:e0191492. https://doi.org/10.1371/journal.pone.0191492
Al Harrasi I, Al-Yahyai R, Yaish MW (2017) Detection of differential DNA methylation under stress conditions using bisulfite sequence analysis. In: Plant stress tolerance. Springer, New York, pp 121–137
Al Kharusi L, Al Yahyai R, Yaish MW (2019) Antioxidant response to salinity in salt-tolerant and salt-susceptible cultivars of date palm. Agriculture 9:8. https://doi.org/10.3390/agriculture9010008
Al Kharusi L, Assaha DV, Al-Yahyai R, Yaish MW (2017) Screening of date palm (Phoenix dactylifera L.) cultivars for salinity tolerance. Forests 8:136
Ali R, Brett CL, Mukherjee S, Rao R (2004) Inhibition of sodium/proton exchange by a Rab-GTPase-activating protein regulates endosomal traffic in yeast. J Biol Chem 279:4498–4506
Arnon DI (1949) Copper enzymes in isolated chloroplasts polyphenoloxidase in Beta vulgaris. Plant Physiol 24:1–15. https://doi.org/10.1104/pp.24.1.1
Barragán V et al (2012) Ion exchangers NHX1 and NHX2 mediate active potassium uptake into vacuoles to regulate cell turgor and stomatal function in Arabidopsis. Plant Cell 24:1127–1142
Bassil E, Coku A, Blumwald E (2012) Cellular ion homeostasis: emerging roles of intracellular NHX Na+/H+ antiporters in plant growth and development. J Exp Bot 63:5727–5740
Bonales-Alatorre E, Pottosin I, Shabala L, Chen Z-H, Zeng F, Jacobsen S-E, Shabala S (2013) Differential activity of plasma and vacuolar membrane transporters contributes to genotypic differences in salinity tolerance in a halophyte species, Chenopodium quinoa. Int J Mol Sci 14:9267–9285. https://doi.org/10.3390/ijms14059267
Brett CL, Donowitz M, Rao R (2005) Evolutionary origins of eukaryotic sodium/proton exchangers. Am J Physiol-Cell Physiol 288:C223–C239
Chow C-N et al (2015) PlantPAN 2.0: an update of plant promoter analysis navigator for reconstructing transcriptional regulatory networks in plants. Nucleic Acids Res 44:D1154–D1160. https://doi.org/10.1093/nar/gkv1035
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Craig Plett D, Møller IS (2010) Na+ transport in glycophytic plants: what we know and would like to know. Plant Cell Environ 33:612–626
Dai X, Xu Y, Ma Q, Xu W, Wang T, Xue Y, Chong K (2007) Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol 143:1739–1751. https://doi.org/10.1104/pp.106.094532
Dragwidge JM, Ford BA, Ashsnest JR, Das P, Gendall AR (2018) Two endosomal NHX-type Na+/H+ antiporters are involved in auxin-mediated development in Arabidopsis thaliana. Plant Cell Physiol 59:1660–1669. https://doi.org/10.1093/pcp/pcy090
Fan L et al (2018) Na+, K+/H+ antiporters regulate the pH of endoplasmic reticulum and auxin-mediated development. Plant Cell Environ 41:850–864
Feng D-F, Doolittle RF (1987) Progressive sequence alignment as a prerequisitetto correct phylogenetic trees. J Mol Evol 25:351–360. https://doi.org/10.1007/bf02603120
Ford BA, Ernest JR, Gendall AR (2012) Identification and characterization of orthologs of AtNHX5 and AtNHX6 in Brassica napus. Front Plant Sci 3:208
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
Gong X, Zhang J, Liu J-H (2014) A stress responsive gene of Fortunella crassifolia FcSISP functions in salt stress resistance. Plant Physiol Biochem 83:10–19
Guan B, Hu Y, Zeng Y, Wang Y, Zhang F (2011) Molecular characterization and functional analysis of a vacuolar Na+/H+ antiporter gene (HcNHX1) from Halostachys caspica. Mol Biol Rep 38:1889–1899
Hamam AM, Britto DT, Flam-Shepherd R, Kronzucker HJ (2016) Measurement of differential Na+ efflux from apical and bulk root zones of intact barley and Arabidopsis plants. Front Plant Sci 7:272
Himabindu Y, Chakradhar T, Reddy MC, Kanygin A, Redding KE, Chandrasekhar T (2016) Salt-tolerant genes from halophytes are potential key players of salt tolerance in glycophytes. Environ Exp Bot 124:39–63
Jana GA, Al Kharusi L, Sunkar R, Al-Yahyai R, Yaish MW (2019) Metabolomic analysis of date palm seedlings exposed to salinity and silicon treatments. Plant Signal Behav 14:1663112
Jiang X, Leidi EO, Pardo JM (2010) How do vacuolar NHX exchangers function in plant salt tolerance? Plant Signal Behav 5:792–795
Krishnamurthy P, Vishal B, Khoo K, Rajappa S, Loh C-S, Kumar PP (2019) Expression of AoNHX1 increases salt tolerance of rice and Arabidopsis, and bHLH transcription factors regulate AtNHX1 and AtNHX6 in Arabidopsis. Plant Cell Rep 38:1–17
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Latz A et al (2013) Salt stress triggers phosphorylation of the Arabidopsis vacuolar K+ channel TPK1 by calcium-dependent protein kinases (CDPKs). Mol Plant 6:1274–1289
Leidi EO et al (2010) The AtNHX1 exchanger mediates potassium compartmentation in vacuoles of transgenic tomato. Plant J 61:495–506
Li M, Li Y, Li H, Wu G (2011) Overexpression of AtNHX5 improves tolerance to both salt and drought stress in Broussonetia papyrifera (L.) Vent. Tree Physiol 31:349–357
Liu C et al (2014) OsbZIP71, a bZIP transcription factor, confers salinity and drought tolerance in rice. Plant Mol Biol 84:19–36
Liu L, Zeng Y, Pan X, Zhang F (2012) Isolation, molecular characterization, and functional analysis of the vacuolar Na+/H+ antiporter genes from the halophyte Karelinia caspica. Mol Biol Rep 39:7193–7202. https://doi.org/10.1007/s11033-012-1551-x
Ma Y, Wang J, Zhong Y, Geng F, Cramer GR, Cheng Z-MM (2015) Subfunctionalization of cation/proton antiporter 1 genes in grapevine in response to salt stress in different organs. Hortic Res 2:15031
Marchler-Bauer A et al (2014) CDD: NCBI's conserved domain database. Nucleic Acids Res 43:D222–D226
Mishra A, Tanna B (2017) Halophytes: potential resources for salt stress tolerance genes and promoters. Front Plant Sci 8:829
Mishra S, Alavilli H, Lee B-H, Panda SK, Sahoo L (2014) Cloning and functional characterization of a vacuolar Na+/H+ antiporter gene from mungbean (VrNHX1) and its ectopic expression enhanced salt tolerance in Arabidopsis thaliana. PLoS ONE 9:e106678. https://doi.org/10.1371/journal.pone.0106678
Mullan D, Pietragalla J (2012) Leaf relative water content physiological breeding II: a field guide to wheat phenotyping. CIMMYT, Mexico, pp 25–27
Munns R, Gilliham M (2015) Salinity tolerance of crops—what is the cost? New Phytol 208:668–673
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Munns R, Wallace PA, Teakle NL, Colmer TD (2010) Measuring soluble ion concentrations (Na+, K+, Cl−) in salt-treated plants. In: Plant stress tolerance. Springer, New York, pp 371–382
Omasits U, Ahrens CH, Müller S, Wollscheid B (2013) Protter: interactive protein feature visualization and integration with experimental proteomic data. Bioinformatics 30:884–886
Patankar HV, Al-Harrasi I, Al-Yahyai R, Yaish MW (2018) Identification of candidate genes involved in the salt tolerance of date palm (Phoenix dactylifera L.) based on a yeast functional bioassay DNA and cell biology
Patankar HV, Al-Harrasi I, Al-Yahyai R, Yaish MW (2019a) Functional characterization of date palm aquaporin gene PdPIP1;2 confers drought and salinity tolerance to yeast and Arabidopsis. Genes 10:390. https://doi.org/10.3390/genes10050390
Patankar HV, Al-Harrasi I, Al Kharusi L, Jana GA, Al-Yahyai R, Sunkar R, Yaish MW (2019b) Overexpression of a metallothionein 2A gene from date palm confers abiotic stress tolerance to yeast and Arabidopsis thaliana. Int J Mol Sci 20:2871
Patankar HV, Assaha DV, Al-Yahyai R, Sunkar R, Yaish MW (2016) Identification of reference genes for quantitative real-time PCR in date palm (Phoenix dactylifera L.) subjected to drought and salinity. PLoS ONE 11:e0166216
Petrezselyova S, Kinclova-Zimmermannova O, Sychrova H (2013) Vhc1, a novel transporter belonging to the family of electroneutral cation–Cl−cotransporters, participates in the regulation of cation content and morphology of Saccharomyces cerevisiae vacuoles. Biochim Biophys Acta (BBA) Biomembr 1828:623–631
Qiu Q-S (2016) AtNHX5 and AtNHX6: Roles in protein transport. Plant Signal Behav 11:e1184810. https://doi.org/10.1080/15592324.2016.1184810
Reguera M et al (2015) pH regulation by NHX-type antiporters is required for receptor-mediated protein trafficking to the vacuole in Arabidopsis. Plant Cell 27:1200–1217
Rodríguez-Rosales MP, Gálvez FJ, Huertas R, Aranda MN, Baghour M, Cagnac O, Venema K (2009) Plant NHX cation/proton antiporters. Plant signal Behav 4:265–276
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151:257–279. https://doi.org/10.1111/ppl.12165
Tester M, Davenport R (2003) Na+ tolerance and Na+ transport in higher plants. Ann Bot 91:503–527
Wang L, Wu X, Liu Y, Qiu Q-S (2015) AtNHX5 and AtNHX6 control cellular K+ and pH homeostasis in Arabidopsis: three conserved acidic residues are essential for K+ transport. PLoS ONE 10:e0144716. https://doi.org/10.1371/journal.pone.0144716
Xu H et al (2018) Arabidopsis thaliana trihelix transcription factor AST1 mediates salt and osmotic stress tolerance by binding to a novel AGAG-box and some GT motifs. Plant Cell Physiol 59:946–965. https://doi.org/10.1093/pcp/pcy032
Xu Y et al (2013) Functional characterization of a wheat NHX antiporter gene TaNHX2 that encodes a K+/H+ exchanger. PLoS ONE 8:e78098. https://doi.org/10.1371/journal.pone.0078098
Yaish M (2015) Proline accumulation is a general response to abiotic stress in the date palm tree (Phoenix dactylifera L.). Genet Mol Res 14:9943–9950
Yaish MW, El-Kereamy A, Zhu T, Beatty PH, Good AG, Bi Y-M, Rothstein SJ (2010) The APETALA-2-like transcription factor OsAP2–39 controls key interactions between abscisic acid and gibberellin in rice. PLoS Genet 6:e1001098
Yaish MW, Patankar HV, Assaha DVM, Zheng Y, Al-Yahyai R, Sunkar R (2017) Genome-wide expression profiling in leaves and roots of date palm (Phoenix dactylifera L.) exposed to salinity. BMC Genomics 18:1. https://doi.org/10.1186/s12864-017-3633-6
Yaish MW, Sunkar R, Zheng Y, Ji B, Al-Yahyai R, Farooq SA (2015) A genome-wide identification of the miRNAome in response to salinity stress in date palm (Phoenix dactylifera L.). Front Plant Sci 6:946
Zaid A, De Wet P (1999) Climatic requirements of date palm. Date palm cultivation. FAO, Roma
Zhang G, Chen M, Li L, Xu Z, Chen X, Guo J, Ma Y (2009) Overexpression of the soybean GmERF3 gene, an AP2/ERF type transcription factor for increased tolerances to salt, drought, and diseases in transgenic tobacco. J Exp Bot 60:3781–3796
Acknowledgements
The authors would like to thank Professor Hana Sychrova, Institute of Physiology Academy of Sciences of the Czech Republic, Prague, Czech Republic, for donating the salt-sensitive mutant S. cerevisiae BYT458 strain, which was used in this study.
Funding
This study is supported by the generous grant number RC/RG-SCI/BIOL/18/01 from the research council (TRC), Oman to MWY.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. All authors revised and approved the final manuscript.
Additional information
Communicated by Neal Stewart.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2020_2549_MOESM2_ESM.tif
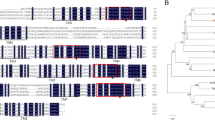
Supplementary file2 Multiple sequence alignment of the deduced amino acid sequence of date palm (PdNHX6) and seven NHX6 isoforms from different plant species. PdNHX6 protein conserved regions (dark grey-colored highlights) that are shared with other NHX6 members. All the shaded areas expand the common conserved domain of NHX family known as sodium/hydrogen exchanger 3 (TIF 5439 kb)
299_2020_2549_MOESM3_ESM.tif
Supplementary file3 The putative topology of date palm (PdNHX6) and Arabidopsis (AtNHX6), using Protter database. The toplogy of PdNHX6 shows 11 transmembrane domains and signal peptide (a). However, AtNHX6 putative toplogy shows 12 transmembrane domains and no signal peptide was predicted (b) (TIF 2393 kb)
299_2020_2549_MOESM4_ESM.tif
Supplementary file4 The abundance of transcription factor binding sites (TFBSs) within 2000 bp promoter region of PdNHX6 computationally analyzed using PlantPAN 2.0 database (TIF 711 kb)
299_2020_2549_MOESM5_ESM.tif
Supplementary file5 The calibration curve of normalized fluorescent intensity (NI490) plotted against pH ranging from 4.0–8.0 (TIF 582 kb)
299_2020_2549_MOESM6_ESM.tif
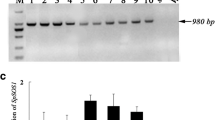
Supplementary file6 PCR-based genotyping of PdNHX6 transgenic Arabidopsis plants using different primer pairs; 35S promoter and NHX6RA (a), NHX6FA, and OCS terminator (b) NHX6FA and NHX6RA (c) and 35S promoter and OCS terminator (d) (TIF 996 kb)
299_2020_2549_MOESM7_ESM.tif
Supplementary file7 Expression analysis of three independent homozygous PdNHX6 transgenic Arabidopsis lines (TL1, TL2, and TL3) and the WT using semi-quantitative RT-PCR and shown on an ethidium bromide-stained 1% agarose gel. The Arabidopsis actin 2 (AtACT2) (AT3G18780) was used as a loading control (TIF 3180 kb)
299_2020_2549_MOESM8_ESM.tif
Supplementary file8 Assay of the salinity tolerance ability of PdNHX6 Arabidopsis lines in vitro. The effect of PdNHX6 on root length, fresh weight, and dry weight of Arabidopsis plantlets grown on MS-agar plates (a) and MS-agar plates supplemented with 100 mM NaCl (b). Each genotype is denoted by color code, as shown on top of the scanning images. The bars represent the mean root length ± SE of three independent replicates. The bars with different letters are significantly different at p < 0.05 (TIF 1588 kb)
Rights and permissions
About this article
Cite this article
Al-Harrasi, I., Jana, G.A., Patankar, H.V. et al. A novel tonoplast Na+/H+ antiporter gene from date palm (PdNHX6) confers enhanced salt tolerance response in Arabidopsis. Plant Cell Rep 39, 1079–1093 (2020). https://doi.org/10.1007/s00299-020-02549-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-020-02549-5