Abstract
The ability to regulate the expression of genes is a central tool for the characterization of fungal genes. This is of particular interest to study genes required for specific processes or the effect of genes expressed only under specific conditions. Saccharomycopsis species show a unique property of necrotrophic mycoparasitism that is activated upon starvation. Here we describe the use of the MET17 promoter of S. schoenii as a tool to regulate gene expression based on the availability of methionine. Conditional expression was tested using lacZ and GFP reporter genes. Gene expression could be strongly down-regulated by the addition of methionine or cysteine to the growth medium and upregulated by starvation for methionine. We used X-gal (5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside) to detect lacZ-expression in plate assays and ONPG (ortho-nitrophenyl-β-galactopyranoside) as a substrate for β-galactosidase in liquid-phase assays. For in vivo expression analyses we used fluorescence microscopy for the detection and localization of a MET17-driven histone H4-GFP reporter gene. With these assays we demonstrated the usefulness of the MET17 promoter to regulate expression of genes based on methionine availability. In silico analyses revealed similar promoter motifs as found in MET3 genes of Saccharomyces cerevisiae and Ashbya gossypii. This suggests a regulation of the MET17 promoter by CBF1 and MET31/MET32 in conjunction with the transcriptional activator MET4, which were also identified in the S. schoenii genome.
Summary
This article describes the characterization of the S. schoenii MET17 promoter for regulated gene expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Growing interest in the biology of Saccharomycopsis species aims at the elucidation of its necrotrophic mycoparasitism. S. schoenii and other predator yeasts of this genus target prey cells by penetration pegs that lead to killing of the prey and uptake of prey nutrients by the predator (Lachance and Pang 1997; Junker et al. 2019). Recent comparative genomics studies indicated that Saccharomycopsis species lack several genes for the uptake and assimilation of sulphate (Junker et al. 2017, 2019; Hesselbart et al. 2018). Sulphate uptake mutants are resistant to otherwise toxic levels of selenate, which provides a convenient selection method for the isolation of Saccharomycopsis species (Lachance and Pang 1997; Dost et al. 2024). Starvation for sulphur compounds can easily be triggered by lack of methionine in growth media and starvation is a prerequisite for predacious activity.
Saccharomycopsis utilize an alternative codon usage as they translate CUG as serine instead of leucine (Krassowski et al. 2018; Muhlhausen et al. 2018; Junker et al. 2019). To initiate molecular genetic studies we development a tool-box for gene-function analyses in Saccharomycopsis (Kayacan et al. 2019; Kaimenyi et al. 2023). This tool-box contains a transformation protocol, several marker genes as well as several constitutively active promoters. However, a promoter for a regulated expression of target genes was not available.
In other systems regulatable promoters are derived from various sources. In Saccharomyces cerevisiae galactose-inducible promoters derived from the GAL1, GAL7 or GAL10 genes, have been used for the strong expression of genes (Brunelli and Pall 1993; Sil et al. 2000). Another sugar-inducible promoter available in S. cerevisiae is the MAL32 promoter that is activated by maltose and repressed by glucose (Meurer et al. 2017). Sugar-regulated promoters have been utilized in other systems as well, e.g. xylose-dependent gene expression in Chaetomium thermophilum (Reislöhner et al., 2023). Recombinant protein expression in Komagataella phaffii (formerly Pichia pastoris) relies on the methanol-inducibility of the AOX1 promoter, while in Aspergillus nidulans the alcA promoter is used for alcohol-induced gene expression (Waring et al. 1989; Raschke et al. 1996). Other modes of regulation employ the thiamine (vitamin B1)-repressible nmt1 promoter in Schizosaccharomyces pombe, the light-inducible vvd promoter in Neurospora crassa or doxycycline on/off-regulatable tet-promoters in S. cerevisiae and other fungi (Moreno et al. 2000; Hurley et al. 2012; Kluge et al. 2018). Feedback-inhibition of the methionine biosynthesis pathway via methionine allows regulatable gene expression by using MET3 or MET17 promoters, e.g. in Ashbya gossypii, Candida albicans or S. cerevisiae (Cherest et al. 1985; Thomas et al., 1989; Care et al. 1999; Dünkler and Wendland, 2007).
In this paper, we describe our efforts to establish a regulatable gene expression system in the predator yeast S. schoenii. Due to lack of homologs of the widely used GAL-, MAL- or MET3-promoters we targeted the S. schoenii MET17 promoter and employed lacZ and GFP as heterologous reporter genes. Regulatable expression via the SsMET17-promoter could be achieved via the addition or removal of methionine in the growth media. This adds a further tool to the functional analysis repertoire for Saccharomycopsis species.
Materials and methods
Strains and culture conditions
The Saccharomyces cerevisiae strain BY4741 (MATa; his3Δ1; leu2Δ0; met1Δ0; ura3Δ0; Euroscarf, Oberursel, Germany) and the Saccharomycopsis schoenii strain CBS 7425 (wild type, Westerdijk Institute, Utrecht, The Netherlands) and its derivatives were propagated in YPD (1% yeast extract, 2% casein peptone and 2% dextrose) at 30 °C. For the selection of transformants antibiotic resistance against G418 (200 μg/mL, Genaxxon, Ulm, Germany) was used. YPD full medium was supplemented with 10 mM methionine for the repression of MET17-promoter driven reporter gene expression. Minimal media (SD: 1.7 g/L yeast nitrogen base (YNB) without amino acids and NH4SO4, 20 g/L glucose, 2 g/L asparagine; and CSM: SD + 0.69 g/L CSM complete synthetic medium and CSM-Met: SD + 0.69 g/L CSM Single Drop-Out: -Met) were used for the induction of the MET17 promoter. Regulation of gene expression controlled via SsMET17p was tested in the presence or absence of 2mM methionine or cysteine in CSM-Met medium. Solid media were prepared by the addition of 2% agar prior to autoclaving. Escherichia coli DH5α was used as host for plasmids and propagated in 2xYT (1.6% bacto peptone, 1% yeast extract, 0.5% NaCl) with 100 μg/mL ampicillin at 37 °C.
Plasmid constructs
Plasmids pE065 (pRS417-SsTEFp-lacZ-SAK1) and pE074 (pRS417-SsMET17p-lacZ-SAK1) were described previously (Kayacan et al. 2019). Plasmid pE196 is a pUC57 based plasmid carrying a synthetic S. schoenii Histone H4-GFP fusion gene (Genscript, Piscataway NJ, USA). Plasmid pE284 (pRS417-SsMET17p-SsH4-GFP-SAK1) was generated by cleaving pE074 with EcoRI, which removes most of the lacZ gene and inserting the SsH4-GFP portion from pE196 via in vivo recombination in S. cerevisiae (Wendland 2003). To this end the SsH4-GFP-fragment was amplified from pE196 with primers #1140 (5’- gtgtaattatatcattttaccttcttttttactatactcgagtttATGTCCGGTCGTGGTAAAGG-3’); lower case letters indicate the homology region to the 3’-end of the MET17-promoter, upper case letters indicate the 5’-end of the S. schoenii Histone H4-gene) and #1141 (5’- tttgtgcaattttctctgaggaggctagatctggcgcgccggatcTATGCGTCCATCTTTACAGTC-3’); lower case letters indicate the homology region to SAK1, upper case letters correspond to the end of the ScURA3-terminator used as terminator for the SsH4-GFP-constuct) and co-transformed with the linear pE074 into BY4741. Correct plasmid assembly was verified by PCR and sequencing. DNA of S. cerevisiae transformants was prepared and plasmids were retrieved by transformation into E. coli as described (Kayacan et al. 2019). Plasmid DNA was prepared using the PureYield Plasmid Midiprep System (Promega, Walldorf, Germany).
Fungal transformation
S. cerevisiae was transformed using the LiAc/single-stranded carrier DNA/PEG method developed by Gietz and Woods (2002) with the addition of DMSO (Kawai et al. 2010). S. schoenii was transformed as described (Kaimenyi et al. 2023). S. schoenii integrates linear DNA preferentially at ectopic positions. Thus, plasmids E074 and E284 were cut with KpnI/SacI to release the SsMET17p-lacZ-SAK1 or SsMET17p-SsH4-GFP-SAK1 fragments from their vectors, respectively, while the SsTEF1p-lacZ-SAK1 fragment was released from pE065 via XhoI/BamHI. Linear DNAs were then transformed into S. schoenii. The SsTEF1p-lacZ fragment was co-transformed with YES1 PCR product amplified from pYES1 (Kayacan et al. 2019) using primers #9 (5’-gaagcttcgtacgctgcaggtc-3’) and #10 (5’-tctgatatcatcgatgaattcgag-3’). Transformants were selected based on G418 resistance and the presence of the constructs was verified by PCR and subsequent functional assays. This yielded S. schoenii transformants G218 (S. schoenii with SsTEF1p-lacZ, YES1), G427, G428 and G429 (S. schoenii with SsMET17p-lacZ-SAK1) and G447 and G448 (S. schoenii with SsMET17p-H4-GFP-SAK1).
Microscopy
Saccharomycopsis cells were propagated as described and imaged using an Axio Imager microscope equipped with a pco.edge 4.2 camera. Images were acquired and processed using VisiView 5 software (Visitron Systems GmbH, Puchheim, Germany) and Fiji (Schindelin et al. 2012).
Testing of β-galactosidase activity
LacZ expression of strains G427, G428, G429 and G218 was assayed using plate- or liquid phase ONPG (ο-nitrophenyl galactopyranoside)-assays. For plate assays, strains were grown in YPD o/n, washed once with H20 and then 2.5 μL of culture (∼ 1 × 106 cells/mL) were spotted on YPD + 10 mM methionine plates or 2.5 μL of culture (∼ 1 × 107 cells/mL) were spotted on minimal medium plates (CSM, CSM-Met and CSM-Met supplemented with either 2 mM methionine or 2 mM cysteine). Plates were incubated at 30 °C for three days. Then 5 μl of X-gal (10 mg/mL) were spotted onto each colony and plates were incubated for up to two hours before photography. The ONPG-assay was carried out as described previously (Ravasio et al. 2014). Briefly, cells were grown o/n in either YPD + 10 mM methionine or incubated in minimal medium (SD) at 30 °C. The OD600 was measured and samples were prepared in technical duplicates: 2 × 100 μL cell suspensions were centrifuged, washed and resuspended in 50 μl H2O each. Cells were disrupted by three cycles of freezing in liquid nitrogen and thawing. To these samples 75 μL Z- buffer (60 mM Na2HPO4 × 2H2O, 40 mM NaH2PO4 × 2H2O, 10 mM KCl, 1 mM MgSO4 × 7H2O) containing 4 mg/mL ONPG was added. Samples were incubated at 37 °C for 30 min. The enzymatic reaction was stopped by adding 125 μl 1 M Na2CO3. Samples were centrifuged to remove the cell debris and 200 μl of twofold diluted supernatant were measured three times at A405 and A550 (Multiskan FC Microplate Photometer and SkanIT RE Software 6.1.1, ThermoFisher Scientific) to calculate Miller units (MU = 1000 x (OD405 – (1, 75 x OD550) / (t [min] x V [ml] x OD600).
Results
Regulatable promoters in Saccharomycopsis schoenii
Comparative genomics has shown that Saccharomycopsis species lack 10 genes required for the uptake and assimilation of sulphate (Fig. 1). Yet, starvation for methionine still results in upregulation of sulfur metabolic processes including the genes CYS3, MET2 and MET17; particularly, MET17 was found to be highly upregulated under methionine deprivation (Junker et al. 2019). In S. cerevisiae MET-pathway genes are regulated by basic loop–helix–loop (bHLH) protein Cbf1, which recognizes the E-box consensus sequence CACGTG, and the leucine-zipper transcriptional activator Met4 (Kuras et al. 1997; Blaiseau and Thomas 1998). Homologs of these S. cerevisiae genes were found in the S. schoenii genome, suggesting that these proteins take part in the regulation of e.g. SsMET17 during starvation. Additionally, two Cbf1 binding sites were found in the SsMET17 promoter at -371 to -377 (CACGTG) and − 611 to -617 (CATGTG). As MET17 is highly upregulated under methionine starvation we tested the promoter of this gene - encompassing a 1 kb upstream region - for regulated heterologous gene expression.
Sulphate uptake and assimilation pathway. Genes, proteins and intermediate products are shown. MET genes in light grey are absent in Saccharomycopsis schoenii and other Saccharomycopsis species resulting in methionine auxotrophy requiring uptake of sulphur containing amino acids via permeases encoded e.g. by MUP1
MET17-promoter driven reporter gene constructs
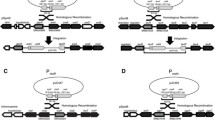
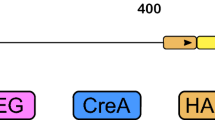
As reporter genes to be placed under the control of SsMET17 we chose lacZ and GFP. To localize the green fluorescence protein in S. schoenii we employed a histone H4-GFP fusion construct (Fig. 2). Since linear DNA is preferentially integrated at ectopic loci the integration of the cassettes was untargeted. Transformants were selected based on their resistance to G418 provided by the selection markers. Integrity of the cassette after integration was verified by PCR (not shown). This yielded the transformants G218 with the SsTEF1p-lacZ construct to register expression of lacZ via a strong constitutive promoter, G427/G428/G429 carrying the SsMET17p-lacZ construct and G447/G448 with the SsMET17p-H4-GFP construct.
Reporter gene constructs used in this study. lacZ and H4-GFP reporter gene construct under the control of either the S. schoenii TEF1 (plasmid E065) or MET17 promoters (plasmids E074 and E284) were used. Plasmids were assembled in S. cerevisiae using in vivo recombination (for E065 and E074 see Kayacan et al. 2019). E284 was generated by exchanging the lacZ gene for the H4-GFP (see materials and methods). For transformation the cassettes were released from the vector backbone by restriction digest with the indicated enzymes. S. schoenii transformants were selected for G418 resistance provided by SAK1 and YES1. Approximate sizes (in bp) of the constructs are shown
Plate assays revealing methionine-regulatable β-galactosidase expression
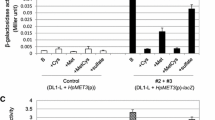
To analyze the regulation of heterologous gene expression via the SsMET17-promoter we spotted transformants on either YPD medium supplemented with additional methionine or on CSM minimal medium plates and incubated them for three days at 30 °C. Then X-gal was applied on the colonies to monitor β-galactosidase activity (Fig. 3): The SsMET17 promoter was shut down effectively on medium containing 10 mM methionine, although we reported some leakiness after a prolonged incubation (120 min, Fig. 3A). On the other hand, expression of the reporter by the SsMET17 promoter was strongly induced on minimal CSM medium (Fig. 3B). To investigate if the SsMET17-promoter can be shut off by organic sulphur compounds we added either 2 mM methionine or cysteine to the solid medium (Fig. 3C). While the control spots showed LacZ-expression, addition of methionine/cysteine blocked its expression. The expression of β-galactosidase via the SsTEF1 promoter was very strong and constitutive under both conditions as expected. This indicates that the SsMET17 promoter is a useful tool for regulated gene expression in S. schoenii.
Colony level analysis of SsMET17-regulated lacZ gene expression in S. schoenii. Cells of strains with the indicated reporter gene constructs were spotted either on YPD + methionine plates and on minimal media CSM-plates and incubated for three days at 30 °C. To visualize β-galactosidase activity 5 μL of an X-gal solution (10 mg/mL) was spotted on each colony. Methionine was sufficient to down regulate SsMET17-dependent LacZ activity (A), while starvation induced expression of lacZ via the SsMET17 promoter (B). Addition of methionine or cysteine to minimal medium blocked lacZ expression (C). SsTEF1-driven lacZ expression in contrast was constitutive
Liquid phase analysis of methionine-regulatable β-galactosidase expression
We used a liquid-phase ONPG assay to quantify the SsMET17-promoter-regulated β-galactosidase activity in either induced (minimal medium) or repressed (YPD) conditions (Fig. 4). The average absolute Miller Units ranged between 7.7 and 10.7 under repressed (YPD) and 396.2 to 639.7 under induced conditions (SD) for SsMET17p-LacZ mutants and reached 3087.4 (YPD) and 1560.5 (SD) for the SsTEFp-LacZ mutant. Normalization of average Miller Units (relative to SsTEFp-LacZ) allowed comparison of SsMET17-driven β-galactosidase activity under repressive and inductive conditions: Under inducing conditions in minimal medium the SsMET17-driven β-galactosidase activity reached ∼ 41% of that obtained via expression of lacZ from the SsTEF1-promoter, while this relative activity was ∼ 0.3% under repressing conditions. The normalized β-galactosidase activity in minimal medium compared to YPD was increased on average by > 100-fold. (Fig. 4).
SsMET17-promoter activity in liquid-phase β-galactosidase assays. The different Saccharomycopsis strains shown were processed as described in Materials and methods. The β-galactosidase activities were calculated as Miller units shown as relative values with respect to constitutive lacZ expression via the SsTEF1 promoter. Error bars represent relative standard deviations of experiments performed in technical duplicates
Histone H4-GFP expression regulated by the S. schoenii MET17-promoter
We also analyzed heterologous gene expression in vivo using a SsMET17-driven histone H4-GFP reporter gene. We chose an H4-GFP fusion construct instead of SsMET17-promoter driven GFP expression to be able to localize the GFP-signal to the nucleus and avoid that low-level expression would be scattered in the cytoplasm. Cells grown o/n in YPD + 10 mM methionine showed no GFP-signal indicating that expression of the reporter gene was practically shut off (Fig. 5A). Conversely, o/n incubation in minimal SD medium strongly induced the reporter gene and resulted in bright nuclear GFP-fluorescence (Fig. 5B). Similar results were obtained with CSM-methionine drop out medium (CSM-Met). Addition of 2 mM methionine or 2 mM cysteine to CSM-Met efficiently down-regulated H4-GFP expression providing a range of media to control SsMET17-promoter activity (Fig. 5C-E).
Regulation of SsMET17-dependent H4-GFP reporter gene activity. Cells of two independent transformants were incubated overnight (for 22 h) under repressive conditions (A) or inducing conditions (B, C). Addition of methionine or cysteine blocks expression of the GFP-reporter even under inducing conditions (D, E). Representative images of cells are shown. Brightfield images (DIC, upper part) and images showing GFP-fluorescence (GFP, center) were merged in an overlay (Merge, lower part). Size bar is 5 μm
To monitor the time required to lose the nuclear GFP signal we used time-lapse microscopy. Cells induced o/n in minimal medium were placed on deep-well microscopy slides containing YPD + methionine solidified with agarose. Then GFP-fluorescence was monitored over time. Resting cells maintained their nuclear GFP signal even after more than eight hours suggesting that fluorescence is not diminished by influx of untagged histone H4 (upper cell in Fig. 6). Dividing cells, however, lost their GFP-signal after two cell generations. Signal intensity in the dividing cell at timepoint 92’ compared to 248’ indicates that the amount of H4-GFP is distributed equally between mother and daughter nuclei suggesting that SsMET17-driven H4-GFP production is rapidly turned off after the switch to methionine-containing medium and wild type untagged histone H4 is incorporated into replicated DNA. (lower cells in Fig. 6).
Time-lapse analysis of H4-GFP localization. A sequence of four time points is shown with S. schoenii cells of strain G447 (SsMET17p-H4-GFP) that were transferred from MET-promoter inducing to repressive conditions. At the indicated time points (in min) Brightfield (top row) and fluorescent images (middle row) were acquired and merged (bottom row). Size bar is 5 μm
Discussion
Regulatable promoters are very useful tools to either up-regulate or down-regulate the expression of genes, particularly of essential genes or genes required for specific processes (Delic et al. 2013). A number of promoters have been established in various systems including sugar-, amino-acid and vitamin regulatable promoters. Often this requires metabolic changes to regulate these promoters, which places some constraints on the conditions under which cells can then be studied (Fuller et al. 2018). However, metabolic promoters are often quite robust in that they provide a means to shut-down promoter activity (almost) completely, e.g. in the case of MAL- and GAL-promoters for S. cerevisiae (Mumberg et al. 1994; Meurer et al. 2017). Other regulatable promoters that do not require changes in the medium composition are tet-regulatable or light-regulatable promoter systems (Wishart et al. 2005; Hurley et al. 2012).
Previously, a set of constitutive Saccharomycopsis promoters was identified via reporter gene analyses in S. cerevisiae (Kayacan et al. 2019). This study now advances the repertoire for gene-function analyses in S. schoenii with the characterization of the SsMET17 promoter. In other yeasts, use of the regulatable MET3 promoter has been described (Cherest et al. 1985; Care et al. 1999; Dünkler and Wendland, 2007). Since the MET3 gene is not present in Saccharomycopsis, we picked the SsMET17 promoter based on available RNAseq expression data (Junker et al. 2019).
Our data corroborate the RNAseq results since under inducing conditions SsMET17 expression was found to be very strong. Importantly, expression can be turned off conveniently by an overnight incubation in methionine containing medium. This allows liquid-phase or microscopic studies. Plate assays, however, revealed that after prolonged incubation under restrictive conditions some leakiness occurs (see Fig. 3A). This may be due to colony dynamics in that cells in the center of a colony where cells may relatively quickly experience starvation conditions and thus induce MET-promoter activity (Cap et al. 2015).
The regulation of MET-promoters in S. cerevisiae requires protein complexes with Cbf1, the centromere binding factor 1. Cbf1 binds as a homodimer to CACGTG consensus sequences present in S. cerevisiae MET gene promoters and the centromere DNA element CDEI (Cai and Davies, 1990). CBF1 is conserved in yeasts and Cbf1-binding sites were also identified in the SsMET17 promoter as well as in the promoters of other sulphur metabolism genes, e.g. SsCYS3 and SsMET2.
The ability to induce Saccharomycopsis genes under starvation conditions provides an important addition to the molecular toolbox for this genus. It will be useful to discern specific functions of genes during starvation, especially since under these conditions Saccharomycopsis cells trigger a predacious behavior that results in penetration peg formation and the attack and subsequent killing of prey cells (Lachance et al., 1997; Junker et al. 2018, 2019). The SsMET17 promoter allows the upregulation of genes during starvation, i.e. at the onset of predacious behavior.
Data availability
No datasets were generated or analysed during the current study.
References
Blaiseau PL, Thomas D (1998) Multiple transcriptional activation complexes tether the yeast activator Met4 to DNA. EMBO J 17:6327–6336
Brunelli JP, Pall ML (1993) A series of yeast shuttle vectors for expression of cDNAs and other DNA sequences. Yeast 9:1299–1308
Cai M, Davis RW (1990) Yeast centromere binding protein CBF1, of the helix-loop-helix protein family, is required for chromosome stability and methionine prototrophy. Cell 61:437–446
Cap M, Vachova L, Palkova Z (2015) Longevity of U cells of differentiated yeast colonies grown on respiratory medium depends on active glycolysis. Cell Cycle 14:3488–3497
Care RS, Trevethick J, Binley KM, Sudbery PE (1999) The MET3 promoter: a new tool for Candida albicans molecular genetics. Mol Microbiol 34:792–798
Cherest H, Nguyen NT, Surdin-Kerjan Y (1985) Transcriptional regulation of the MET3 gene of Saccharomyces cerevisiae. Gene 34:269–281
Delic M, Mattanovich D, Gasser B (2013) Repressible promoters - a novel tool to generate conditional mutants in Pichia pastoris. Microb Cell Fact 12:6
Dost C, Michling F, Kaimenyi D, Rij M, Wendland J (2024) Isolation of Saccharomycopsis species from plant material. Microbiol Res 283:127691. https://doi.org/10.1016/j.micres.2024.127691
Dunkler A, Wendland J (2007) Use of MET3 promoters for regulated gene expression in Ashbya Gossypii. Curr Genet 52:1–10
Fuller KK, Dunlap JC, Loros JJ (2018) Light-regulated promoters for tunable, temporal, and affordable control of fungal gene expression. Appl Microbiol Biotechnol 102:3849–3863
Gietz RD, Woods RA (2002) Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol 350:87–96
Hesselbart A, Junker K, Wendland J (2018) Draft genome sequence of Saccharomycopsis fermentans CBS 7830, a predacious yeast belonging to the Saccharomycetales. Genome Announc 6
Hurley JM, Chen CH, Loros JJ, Dunlap JC (2012) Light-inducible system for tunable protein expression in Neurospora Crassa. G3 (Bethesda) 2:1207–1212
Junker K, Hesselbart A, Wendland J (2017) Draft genome sequence of Saccharomycopsis fodiens CBS 8332, a necrotrophic mycoparasite with biocontrol potential. Genome Announc 5
Junker K, Bravo Ruiz G, Lorenz A, Walker L, Gow NAR, Wendland J (2018) The mycoparasitic yeast Saccharomycopsis Schoenii predates and kills multi-drug resistant Candida Auris. Sci Rep 8:14959
Junker K, Chailyan A, Hesselbart A, Forster J, Wendland J (2019) Multi-omics characterization of the necrotrophic mycoparasite Saccharomycopsis Schoenii. PLoS Pathog 15:e1007692
Kaimenyi D, Rij M, Wendland J (2023) Improved gene-targeting efficiency upon starvation in Saccharomycopsis. Fungal Genet Biol 167:103809
Kawai S, Hashimoto W, Murata K (2010) Transformation of Saccharomyces cerevisiae and other fungi: methods and possible underlying mechanism. Bioeng Bugs 1:395–403
Kayacan Y, Griffiths A, Wendland J (2019) A script for initiating molecular biology studies with non-conventional yeasts based on Saccharomycopsis Schoenii. Microbiol Res 229:126342
Kluge J, Terfehr D, Kuck U (2018) Inducible promoters and functional genomic approaches for the genetic engineering of filamentous fungi. Appl Microbiol Biotechnol 102:6357–6372
Krassowski T, Coughlan AY, Shen XX et al (2018) Evolutionary instability of CUG-Leu in the genetic code of budding yeasts. Nat Commun 9:1887
Kuras L, Barbey R, Thomas D (1997) Assembly of a bZIP-bHLH transcription activation complex: formation of the yeast Cbf1-Met4-Met28 complex is regulated through Met28 stimulation of Cbf1 DNA binding. EMBO J 16:2441–2451
Lachance MA, Pang WM (1997) Predacious yeasts. Yeast 13:225–232
Meurer M, Chevyreva V, Cerulus B, Knop M (2017) The regulatable MAL32 promoter in Saccharomyces cerevisiae: characteristics and tools to facilitate its use. Yeast 34:39–49
Moreno MB, Duran A, Ribas JC (2000) A family of multifunctional thiamine-repressible expression vectors for fission yeast. Yeast 16:861–872
Muhlhausen S, Schmitt HD, Pan KT, Plessmann U, Urlaub H, Hurst LD, Kollmar M (2018) Endogenous stochastic decoding of the CUG Codon by competing ser- and Leu-tRNAs in Ascoidea Asiatica. Curr Biol 28:2046–2057 e2045
Mumberg D, Muller R, Funk M (1994) Regulatable promoters of Saccharomyces cerevisiae: comparison of transcriptional activity and their use for heterologous expression. Nucleic Acids Res 22:5767–5768
Raschke WC, Neiditch BR, Hendricks M, Cregg JM (1996) Inducible expression of a heterologous protein in Hansenula polymorpha using the alcohol oxidase 1 promoter of Pichia pastoris. Gene 177:163–167
Ravasio D, Walther A, Trost K, Vrhovsek U, Wendland J (2014) An indirect assay for volatile compound production in yeast strains. Sci Rep 4:3707
Reislohner S, Schermann G, Kilian M, Santamaria-Munoz D, Zimmerli C, Kellner N, Bassler J, Brunner M, Hurt E (2023) Identification and characterization of sugar-regulated promoters in Chaetomium Thermophilum. BMC Biotechnol 23:19
Schindelin J, Arganda-Carreras I, Frise E et al (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682
Sil AK, Xin P, Hopper JE (2000) Vectors allowing amplified expression of the Saccharomyces cerevisiae Gal3p-Gal80p-Gal4p transcription switch: applications to galactose-regulated high-level production of proteins. Protein Expr Purif 18:202–212
Thomas D, Cherest H, Surdin-Kerjan Y (1989) Elements involved in S-adenosylmethionine-mediated regulation of the Saccharomyces cerevisiae MET25 gene. Mol Cell Biol 9:3292–3298
Waring RB, May GS, Morris NR (1989) Characterization of an inducible expression system in aspergillus nidulans using alcA and tubulin-coding genes. Gene 79:119–130
Wendland (2003) PCR-based methods facilitate targeted gene manipulations and cloning procedures. Curr Genet 44:115–123. https://doi.org/10.1007/s00294-003-0436-x
Wishart JA, Hayes A, Wardleworth L, Zhang N, Oliver SG (2005) Doxycycline, the drug used to control the tet-regulatable promoter system, has no effect on global gene expression in Saccharomyces cerevisiae. Yeast 22:565–569
Acknowledgements
This project was funded in part by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – – project 448656174.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.W. and M.R.; methodology, M.R. and J.W.; validation, M.R. and J.W.; formal analysis, M.R. investigation, M.R. and J.W.; resources, J.W., data curation, M.R.; writing—original draft preparation, J.W.; writing—review and editing, J.W. and M.R.; visualization, M.R. and J.W; supervision, J.W.; project administration, J.W.; funding acquisition, J.W. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by Alexander Idnurm.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rij, M., Wendland, J. Use of the Saccharomycopsis schoenii MET17 promoter for regulated heterologous gene expression. Curr Genet 70, 9 (2024). https://doi.org/10.1007/s00294-024-01294-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00294-024-01294-6