Abstract
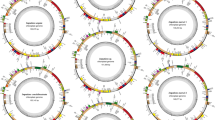
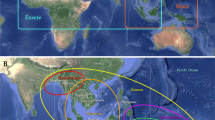
The complete plastome sequencing is an efficient option for increasing phylogenetic resolution and evolutionary studies, as well as may greatly facilitate the use of plastid DNA markers in plant population genetic studies. Merostachys and Guadua stand out as the most common and the highest potential utilization bamboos indigenous of Brazil. Here, we sequenced the complete plastome sequences of the Brazilian Guadua chacoensis and Merostachys sp. to perform full plastome phylogeny and characterize the occurrence, type, and distribution of SRRs using 20 Bambuseae species. The determined plastome sequence of Merostachys sp. and G. chacoensis is 136,334 and 135,403 bp in size, respectively, with an identical gene content and typical quadripartite structure consisting of a pair of IRs separated by the LSC and SSC regions. The Maximum Likelihood and Bayesian Inference analyses produced phylogenomic trees identical in topology. These trees supported monophyly of Paleotropical and Neotropical Bamboos clades. The Neotropical bamboos segregated into three well-supported lineages, Chusqueinae, Guaduinae, and Arthrostylidiinae, with the last two forming a well-supported sister relationship. Paleotropical bamboos segregated into two well-supported lineages, Hickeliinae and Bambusinae + Melocanninae. We identified 141.8 cpSSR in Bambuseae plastomes and an inferior value (38.15) for plastome coding sequences. Among them, we identified 16 polymorphic SSR loci, with number of alleles varying from 3 to 10. These 16 polymorphic cpSSR loci in Bambuseae plastome can be assessed for the intraspecific level of polymorphism, leading to innovative highly sensitive phylogeographic and population genetics studies for this tribe.



Similar content being viewed by others
References
Bamboo Phylogeny Group (2006) The bamboo phylogeny project. Bamboo 27:11–14
Bamboo Phylogeny Group (2012) An updated tribal and subtribal classification for the Bambusoideae (Poaceae). In: Gielis J, Potters G (eds) Proceedings of the 9th World Bamboo Congress. Antwerp, Belgium: World Bamboo Organization, pp 3–27
Bock R (2006) Extranuclear inheritance: gene transfer out of plastids. Prog Bot 67:75–100. doi:10.1007/3-540-27998-9_4
Burke SV, Colin PG, Melvin RD (2012) Plastome sequences of two New World bamboos—Arundinaria gigantea and Cryptochloa strictiflora (Poaceae)—extend phylogenomic understanding of Bambusoideae. Am J Bot 99:1951–1961. doi:10.3732/ajb.1200365
Burke SV, Clark LG, Triplett JK, Grennan CP, Duvall MR (2014) Biogeography and phylogenomics of new world Bambusoideae (Poaceae), revisited. Am J Bot 101:886–891. doi:10.3732/ajb.1400063
Clark LG, Dransfield S, Triplett JK, Sánchez-Ken JG (2007) Phylogenetic relationships among the one-flowered, determinate genera of Bambuseae (Poaceae: Bambusoideae). Aliso 23:315–332
Clark LG, Londoño X, Ruiz-Sanchez E (2015) Bamboo taxonomy and habitat. In: Liese W, Kohl M (eds) Bamboo, Tropical forestry. Springer International Publishing, Switzerland, pp 1–30
Delplancke M, Alvarez N, Espíndola A, Joly H, Benoit L, Brouck E, Arrigo N (2012) Gene flow among wild and domesticated almond species: insights from chloroplast and nuclear markers. Evol Appl 5:317–329. doi:10.1111/j.1752-4571.2011.00223.x
Fisher AE, Clark LG, Kelchner SA (2014) Molecular phylogeny estimation of the bamboo genus Chusquea (Poaceae: Bambusoideae: Bambuseae) and description of two new subgenera. Syst Bot 39:829–844. doi:10.1600/036364414X681554
George B, Bhatt BS, Awasthi M, George B, Singh AK (2015) Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants. Curr Genet. doi:10.1007/s00294-015-0495-9
Greco TM, Pinto MM, Tombolato FC, Xia N (2015) Diversity of bamboo in Brazil. J Trop Subtrop Botany 23:1–16. doi:10.11926/j.issn.1005-3395.2015.01.001
Guala G, Bogler D, Sadle J, Francisco-Ortega J (2000) Molecular evidence for polyphyly in the genus Apoclada (Poaceae: Bambusoideae). Bamboo Sci Cult 14:15–20
Huang CY, Ayliffe MA, Timmis JN (2003) Direct measurement of the transfer rate of chloroplast DNA into the nucleus. Nature 422:72–76. doi:10.1038/nature01435
Huang CY, Grunheit N, Ahmadinejad N, Timmis JN, Martin W (2005) Mutational decay and age of chloroplast and mitochondrial genomes transferred recently to angiosperm nuclear chromosomes. Plant Physiol 138:1723–1733. doi:10.1104/pp.105.060327
Judziewicz EJ, Clark LG, Londono X, Stern M (1999) American bamboos. Smithsonian Institution Press, Washington DC
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. doi:10.1093/molbev/mst010
Kelchner SA, Group BP (2013) Higher level phylogenetic relationships within the bamboos (Poaceae: Bambusoideae) based on five plastid markers. Mol phylogenetics Evol 67:404–413. doi:10.1016/j.ympev.2013.02.005
Khadivi-Khub A, Zamani Z, Fattahi R, Wünsch A (2013) Genetic variation in wild Prunus L. subgen. Cerasus germplasm from Iran characterized by nuclear and chloroplast SSR markers. Trees 28:471–485. doi:10.1007/s00468-013-0964-z
Kurtz S, Choudhuri JV, Ohlebusch E, Schleiermacher C, Stoye J, Giegerich R (2001) REPuter: the manifold applications of repeat analysis on a genomic scale. Nucl Acids Res 29:4633–4642
Li W, Feng Y, Sun H, Deng Y, Yu H, Chen H (2014) Analysis of simple sequence repeats in the Gaeumannomyces graminis var. tritici genome and the development of microsatellite markers. Curr Genet 60:237–245. doi:10.1007/s00294-014-0428-z
List of Species of the Brazilian Flora (2015) Rio de Janeiro Botanical Garden. http://floradobrasil.jbrj.gov.br. Accessed 04 July 2015
Lizarazu MA, Agrasar ZER, Vega AS (2011) A new species of Merostachys (Poaceae, Bambusoideae, Bambuseae) and synopsis of the genus in Argentina and neighboring regions. Sist Bot 36:896–906. doi:10.1600/036364411X604912
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucl Acids Res. doi:10.1093/nar/gkt289
Londoño X, Peterson PM (1992) Guadua chacoensis (Poaceae: Bambuseae), its taxonomic identity, morphology, and affinities. Novon 2:41–47
Ma PF, Zhang YX, Guo ZH, Li DZ (2015) Evidence for horizontal transfer of mitochondrial DNA to the plastid genome in a bamboo genus. Sci Rep 5:11608. doi:10.1038/srep11608
Myers N, Mittermeier RA, Mittermeier CG, Fonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858. doi:10.1038/35002501
Nazareno AG, Carlsen M, Lohmann LG (2015) Complete Chloroplast Genome of Tanaecium tetragonolobum: The First Bignoniaceae Plastome. PLoS One 10(6):e0129930. doi:10.1371/journal.pone.0129930
Ogihara Y, Isono K, Kojima T, Endo A, Hanaoka M, Shiina T, Terachi T, Utsugi S, Murata M, Mori N, Takumi S, Ikeo K, Gojobori T, Murai R, Murai K, Matsuoka Y, Ohnishi Y, Tajiri H, Tsunewaki K (2002) Structural features of a wheat plastome as revealed by complete sequencing of chloroplast DNA. Mol Genet Genomics 266:740–746. doi:10.1007/s00438-001-0606-9
Palmer JD (1985) Chloroplast DNA and molecular phylogeny. BioEssays 2:263–267
Powell W, Morgantet M, Andre C, McNicol JW, Machray GC, Doyle JJ, Tingey SV, Rafalski JA (1995) Hypervariable microsatellites provide a general source of polymorphic DNA markers for the chloroplast genome. Curr Biol 5:1023–1029
Provan J, Powell W, Hollingsworth PM (2001) Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol 16:142–147. doi:10.1016/s0169-5347(00)02097-8
Rogalski M, Vieira LN, Fraga HPF, Guerra MP (2015) Plastid genomics in horticultural species: importance and applications for plant population genetics, evolution, and biotechnology. Front Plant Sci 6:586. doi:10.3389/fpls.2015.00586
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinform 19:1572–1574. doi:10.1093/bioinformatics/btg180
Roullier C, Rossel G, Tay D, McKey D, Lebot V (2011) Combining chloroplast and nuclear microsatellites to investigate origin and dispersal of New World sweet potato landraces. Mol Ecol 20:3963–3977. doi:10.1111/j.1365-294X.2011.05229.x
Santos-Gonçalves APS, Carvalho-Okano RMD, Filgueiras TS (2012) A new species of Merostachys (Poaceae: Bambusoideae) from southeastern Brazil. Syst Bot 37:938–940. doi:10.1600/036364412X656400
Saski C, Lee SB, Fjellheim S, Guda C, Jansen RK, Luo H, Tomkins J, Rognli OA, Daniell H, Clarke JL (2007) Complete chloroplast genome sequences of Hordeum vulgare, Sorghum bicolor and Agrostis stolonifera, and comparative analyses with other grass genomes. Theor Appl Genet 115:571–590. doi:10.1007/s00122-007-0567-4
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:W686–W689. doi:10.1093/nar/gki366
Shaw J, Lickey EB, Beck JT, Farmer SB, Liu W, Miller J, Siripun KC, Winder CT, Schilling EE, Small RL (2005) The tortoise and the hare II: relative utility of 21 noncoding chloroplast DNA sequences for phylogenetic analysis. Am J Bot 92:142–166. doi:10.3732/ajb.92.1.142
Shaw J, Lickey EB, Schilling EE, Small RL (2007) Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare III. Am J Bot 94:275–288. doi:10.3732/ajb.94.3.275
Soreng RJ, Peterson PM, Romaschenko K, Davidse G, Zuloaga FO, Judziewicz EJ, Filgueiras TS, Davis JI, Morrone O (2015) A worldwide phylogenetic classification of the Poaceae (Gramineae). J Syst Evol 53:117–137. doi:10.1111/jse.12150
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinform 22:2688–2690. doi:10.1093/bioinformatics/btl446
Stegemann S, Bock R (2006) Experimental reconstruction of functional gene transfer from the tobacco plastid genome to the nucleus. Plant Cell 18:2869–2878. doi:10.1105/tpc.106.046466
Stegemann S, Hartmann S, Ruf S, Bock R (2003) High-frequency gene transfer from the chloroplast genome to the nucleus. Proc Natl Acad Sci USA 100:8828–8833. doi:10.1073/pnas.1430924100
Sungkaew S, Stapleton CM, Salamin N, Hodkinson TR (2009) Non-monophyly of the woody bamboos (Bambuseae; Poaceae): a multi-gene region phylogenetic analysis of Bambusoideae ss. J Plant Res 122:95–108. doi:10.1007/s10265-008-0192-6
Viana PL, Filgueiras TS (2014) Three new species of Aulonemia (Poaceae: Bambusoideae) from the Brazilian Atlantic rainforest. Phytotaxa 156:235–249. doi:10.11646/phytotaxa.156.4.6
Viana PL, Filgueiras TS, Clark LG (2013) Cambajuva (Poaceae: Bambusoideae: Bambuseae: Arthrostylidiinae), a new woody bamboo genus from southern Brazil. Syst Bot 38:97–103. doi:10.1600/036364413X662097
Vieira LN, Faoro H, Fraga HPF, Rogalski M, Souza EM et al (2014) An improved protocol for intact chloroplasts and cpDNA isolation in conifers. PLoS One 9(1):e84792. doi:10.1371/journal.pone.0084792
Wheeler GL, McGlaughlin ME, Wallace LE (2012) Variable length chloroplast markers for population genetic studies in Acmispon (Fabaceae). Am J Bot 99:e408–e410. doi:10.3732/ajb.1200129
Wheeler GL, Dorman HE, Buchanan A, Challagundla L, Wallace LE (2014) A review of the prevalence, utility, and caveats of using chloroplast simple sequence repeats for studies of plant biology. Appl Plant Sci 2:1400059. doi:10.3732/apps.1400059
Wu FH, Kan DP, Lee SB, Daniell H, Lee YW, Lin CC, Lin NS, Lin CS (2009) Complete nucleotide sequence of Dendrocalamus latiflorus and Bambusa oldhamii chloroplast genomes. Tree Physiol tpp015:1–10. doi:10.1093/treephys/tpp015
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255. doi:10.1093/bioinformatics/bth352
Wysocki WP, Clark LG, Attigala L, Ruiz-Sanchez E, Duvall MR (2015) Evolution of the bamboos (Bambusoideae; Poaceae): a full plastome phylogenomic analysis. BMC Evol Biol 15:50. doi:10.1186/s12862-015-0321-5
Zhang W-P, Clark LG (2000) Phylogeny and classification of the bamboos (Poaceae: Bambusoideae). In: Jacobs SWL, Everett J (eds) Grasses: systematics and evolution. CSIRO Publishing, Collingwood, Victoria, pp 35–42
Zhang Y-J, Ma P-F, Li D-Z (2011) High-throughput sequencing of six bamboo chloroplast genomes: phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS One 6(5):e20596. doi:10.1371/journal.pone.0020596
Acknowledgments
This work was supported by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq Proc. 457726/2013-0, and 306126/2013-3). The authors thank to Instituto Çarakura for kindly providing plant material.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Vieira, L.d., dos Anjos, K.G., Faoro, H. et al. Phylogenetic inference and SSR characterization of tropical woody bamboos tribe Bambuseae (Poaceae: Bambusoideae) based on complete plastid genome sequences. Curr Genet 62, 443–453 (2016). https://doi.org/10.1007/s00294-015-0549-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-015-0549-z