Abstract
Purpose
Lymphoma lesion detection and segmentation on whole-body FDG-PET/CT are a challenging task because of the diversity of involved nodes, organs or physiological uptakes. We sought to investigate the performances of a three-dimensional (3D) convolutional neural network (CNN) to automatically segment total metabolic tumour volume (TMTV) in large datasets of patients with diffuse large B cell lymphoma (DLBCL).
Methods
The dataset contained pre-therapy FDG-PET/CT from 733 DLBCL patients of 2 prospective LYmphoma Study Association (LYSA) trials. The first cohort (n = 639) was used for training using a 5-fold cross validation scheme. The second cohort (n = 94) was used for external validation of TMTV predictions. Ground truth masks were manually obtained after a 41% SUVmax adaptive thresholding of lymphoma lesions. A 3D U-net architecture with 2 input channels for PET and CT was trained on patches randomly sampled within PET/CTs with a summed cross entropy and Dice similarity coefficient (DSC) loss. Segmentation performance was assessed by the DSC and Jaccard coefficients. Finally, TMTV predictions were validated on the second independent cohort.
Results
Mean DSC and Jaccard coefficients (± standard deviation) in the validations set were 0.73 ± 0.20 and 0.68 ± 0.21, respectively. An underestimation of mean TMTV by − 12 mL (2.8%) ± 263 was found in the validation sets of the first cohort (P = 0.27). In the second cohort, an underestimation of mean TMTV by − 116 mL (20.8%) ± 425 was statistically significant (P = 0.01).
Conclusion
Our CNN is a promising tool for automatic detection and segmentation of lymphoma lesions, despite slight underestimation of TMTV. The fully automatic and open-source features of this CNN will allow to increase both dissemination in routine practice and reproducibility of TMTV assessment in lymphoma patients.



Similar content being viewed by others
References
Kanoun S, Rossi C, Berriolo-Riedinger A, Dygai-Cochet I, Cochet A, Humbert O, et al. Baseline metabolic tumour volume is an independent prognostic factor in Hodgkin lymphoma. Eur J Nucl Med Mol Imaging. 2014;41:1735–43. https://doi.org/10.1007/s00259-014-2783-x.
Sasanelli M, Meignan M, Haioun C, Berriolo-Riedinger A, Casasnovas RO, Biggi A, et al. Pretherapy metabolic tumour volume is an independent predictor of outcome in patients with diffuse large B-cell lymphoma. Eur J Nucl Med Mol Imaging. 2014;41:2017–22. https://doi.org/10.1007/s00259-014-2822-7.
Ceriani L, Martelli M, Zinzani PL, Ferreri AJ, Botto B, Stelitano C, et al. Utility of baseline 18FDG-PET/CT functional parameters in defining prognosis of primary mediastinal (thymic) large B-cell lymphoma. Blood. 2015;126:950–6. https://doi.org/10.1182/blood-2014-12-616474.
Meignan M, Cottereau AS, Versari A, Chartier L, Dupuis J, Boussetta S, et al. Baseline metabolic tumor volume predicts outcome in high-tumor-burden follicular lymphoma: a pooled analysis of three multicenter studies. J Clin Oncol. 2016;34:3618–26. https://doi.org/10.1200/JCO.2016.66.9440.
Albano D, Bosio G, Bianchetti N, Pagani C, Re A, Tucci A, et al. Prognostic role of baseline 18F-FDG PET/CT metabolic parameters in mantle cell lymphoma. Ann Nucl Med. 2019;33:449–58. https://doi.org/10.1007/s12149-019-01354-9.
Albano D, Re A, Tucci A, Giubbini R, Bertagna F. Prognostic role of deltaMTV and deltaTLG in Burkitt lymphoma. Ann Nucl Med. 2019;33:280–7. https://doi.org/10.1007/s12149-019-01338-9.
Cottereau AS, El-Galaly TC, Becker S, Broussais F, Petersen LJ, Bonnet C, et al. Predictive value of PET response combined with baseline metabolic tumor volume in peripheral T-cell lymphoma patients. J Nucl Med. 2018;59:589–95. https://doi.org/10.2967/jnumed.117.193946.
Zhou Z, Sehn LH, Rademaker AW, Gordon LI, Lacasce AS, Crosby-Thompson A, et al. An enhanced International Prognostic Index (NCCN-IPI) for patients with diffuse large B-cell lymphoma treated in the rituximab era. Blood. 2014;123:837–42. https://doi.org/10.1182/blood-2013-09-524108.
Barrington SF, Meignan M. Time to prepare for risk adaptation in lymphoma by standardizing measurement of metabolic tumor burden. J Nucl Med. 2019;60:1096–102. https://doi.org/10.2967/jnumed.119.227249.
Decazes P, Becker S, Toledano MN, Vera P, Desbordes P, Jardin F, et al. Tumor fragmentation estimated by volume surface ratio of tumors measured on 18F-FDG PET/CT is an independent prognostic factor of diffuse large B-cell lymphoma. Eur J Nucl Med Mol Imaging. 2018;45:1672–9. https://doi.org/10.1007/s00259-018-4041-0.
Ilyas H, Mikhaeel NG, Dunn JT, Rahman F, Moller H, Smith D, et al. Is there an optimal method for measuring baseline metabolic tumor volume in diffuse large B cell lymphoma? Eur J Nucl Med Mol Imaging. 2019;46:520–1. https://doi.org/10.1007/s00259-018-4200-3.
Biggi A, Gallamini A, Chauvie S, Hutchings M, Kostakoglu L, Gregianin M, et al. International validation study for interim PET in ABVD-treated, advanced-stage hodgkin lymphoma: interpretation criteria and concordance rate among reviewers. J Nucl Med. 2013;54:683–90. https://doi.org/10.2967/jnumed.112.110890.
Itti E, Meignan M, Berriolo-Riedinger A, Biggi A, Cashen AF, Vera P, et al. An international confirmatory study of the prognostic value of early PET/CT in diffuse large B-cell lymphoma: comparison between Deauville criteria and deltaSUVmax. Eur J Nucl Med Mol Imaging. 2013;40:1312–20. https://doi.org/10.1007/s00259-013-2435-6.
Capobianco N, Meignan MA, Cottereau AS, Vercellino L, Sibille L, Spottiswoode B, et al. Deep learning FDG uptake classification enables total metabolic tumor volume estimation in diffuse large B-cell lymphoma. J Nucl Med. 2020. https://doi.org/10.2967/jnumed.120.242412.
Hatt M, Lee JA, Schmidtlein CR, Naqa IE, Caldwell C, De Bernardi E, et al. Classification and evaluation strategies of auto-segmentation approaches for PET: report of AAPM task group no. 211. Med Phys. 2017;44:e1–e42. https://doi.org/10.1002/mp.12124.
Nioche C, Orlhac F, Boughdad S, Reuze S, Goya-Outi J, Robert C, et al. LIFEx: a freeware for radiomic feature calculation in multimodality imaging to accelerate advances in the characterization of tumor heterogeneity. Cancer Res. 2018;78:4786–9. https://doi.org/10.1158/0008-5472.CAN-18-0125.
Wahl RL, Jacene H, Kasamon Y, Lodge MA. From RECIST to PERCIST: evolving considerations for PET response criteria in solid tumors. J Nucl Med. 2009;50(Suppl 1):122S–50S. https://doi.org/10.2967/jnumed.108.057307.
Isensee F, Jäger PF, Kohl SAA, Petersen J, Maier-Hein KH. Automated design of deep learning methods for biomedical image segmentation. arXiv:190408128v2 [csCV]. [last revised 2 Apr 2020] Available from: http://arxiv.org/abs/1904.08128.
McKinney W. Data structures for statistical computing in Python. Proceedings of the 9th Python in Science Conference (SCIPY 2010). Available from: https://conference.scipy.org/proceedings/scipy2010/; 2010; p. 56–61.
Hunter JD. Matplotlib: a 2D graphics environment. Comput Sci Eng. 2007;9:90–5. https://doi.org/10.1109/MCSE.2007.55.
Bi L, Kim J, Kumar A, Wen L, Feng D, Fulham M. Automatic detection and classification of regions of FDG uptake in whole-body PET-CT lymphoma studies. Comput Med Imaging Graph. 2016;60:3–10. https://doi.org/10.1016/j.compmedimag.2016.11.008.
Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv:14091556v6 [csCV]. [last revised 10 Apr 2015] Available from: http://arxiv.org/abs/1409.1556.
Yu Y, Decazes P, Lapuyade-Lahorgue J, Gardin I, Vera P, Ruan S. Semi-automatic lymphoma detection and segmentation using fully conditional random fields. Comput Med Imaging Graph. 2018;70:1–7. https://doi.org/10.1016/j.compmedimag.2018.09.001.
Grossiord E, Talbot H, Passat N, Meignan M, Najman L. Automated 3D lymphoma lesion segmentation from PET/CT characteristics. IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017); 2017; p. 174–8.
Li H, Jiang H, Li S, Wang M, Wang Z, Lu G, et al. DenseX-net: an end-to-end model for lymphoma segmentation in whole-body PET/CT images. IEEE Access. 2020;8:8004–18. https://doi.org/10.1109/ACCESS.2019.2963254.
Sibille L, Seifert R, Avramovic N, Vehren T, Spottiswoode B, Zuehlsdorff S, et al. (18)F-FDG PET/CT uptake classification in lymphoma and lung cancer by using deep convolutional neural networks. Radiology. 2020;294:445–52. https://doi.org/10.1148/radiol.2019191114.
Pfaehler E, Burggraaff C, Kramer G, Zijlstra J, Hoekstra OS, Jalving M, et al. PET segmentation of bulky tumors: strategies and workflows to improve inter-observer variability. PLoS One. 2020;15:e0230901. https://doi.org/10.1371/journal.pone.0230901.
Meignan M, Sasanelli M, Casasnovas RO, Luminari S, Fioroni F, Coriani C, et al. Metabolic tumour volumes measured at staging in lymphoma: methodological evaluation on phantom experiments and patients. Eur J Nucl Med Mol Imaging. 2014;41:1113–22. https://doi.org/10.1007/s00259-014-2705-y.
Mettler J, Muller H, Voltin CA, Baues C, Klaeser B, Moccia A, et al. Metabolic tumour volume for response prediction in advanced-stage Hodgkin lymphoma. J Nucl Med. 2018. https://doi.org/10.2967/jnumed.118.210047.
Boellaard R, O'Doherty MJ, Weber WA, Mottaghy FM, Lonsdale MN, Stroobants SG, et al. FDG PET and PET/CT: EANM procedure guidelines for tumour PET imaging: version 1.0. Eur J Nucl Med Mol Imaging. 2010;37:181–200. https://doi.org/10.1007/s00259-009-1297-4.
Acknowledgements
We thank the Fondation AP-HP pour la Recherche, Assistance Publique–Hôpitaux de Paris, for their financial support on the deep learning workstation, Romain Ricci (LYSA Image) for technical support and Kathryn Schutte (Owkin) for English support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Simon Jegou (SJ) is an employee at Owkin. SJ advised us on the choice of the algorithm and image processing but did not have access neither to data nor to the training process. All other authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. NCT00498043 and NCT01659099 were approved by the ethics committees of Lyon University Hospital (2007) and Brest University Hospital (2012), respectively.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Advanced Image Analyses (Radiomics and Artificial Intelligence).
Electronic supplementary material
ESM 1
(DOCX 27 kb)
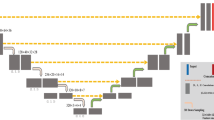
Supplemental Figure 1
Architecture of the 3D U-net. Patch size of 112 × 96 × 224 with batch size 2 is fed to the neural network. Weights are updated with the rectified Adam optimizer. (PPTX 109 kb)
Supplemental Figure 2
Regression analyses between predicted SUVmax from pTMTV mask and the reference SUVmax (from each clinical trial) in the validation sets of the first cohort (A, n = 639) and second cohort (B, n = 94). (PPTX 54 kb)
Rights and permissions
About this article
Cite this article
Blanc-Durand, P., Jégou, S., Kanoun, S. et al. Fully automatic segmentation of diffuse large B cell lymphoma lesions on 3D FDG-PET/CT for total metabolic tumour volume prediction using a convolutional neural network.. Eur J Nucl Med Mol Imaging 48, 1362–1370 (2021). https://doi.org/10.1007/s00259-020-05080-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-020-05080-7