Abstract
Objective
To develop and validate a convolutional neural network (CNN) capable of predicting the anatomical landmarks used to calculate the hip-knee-ankle angles (HKAAs) from radiographs and thereby quantify lower extremity alignments in children.
Materials and methods
A search of the image archive at a large children’s hospital was conducted to identify full-length lower extremity radiographs performed in children (≤ 18 years old) for the indication of lower extremity alignment (7/2019–10/2019). A radiologist manually labeled each radiograph’s six requisite anatomical landmarks used to measure HKAAs (bilateral centers of the femoral head, tibial spine, and tibial plafond) and defined the resultant labels as ground truth. A 2D heatmap was generated for each ground truth landmark to encode the pseudo-probability of a landmark being at a particular location. A CNN was developed for indirect landmark localization by regressing across a collection of these heatmaps. The landmarks predicted from this model were used to calculate the HKAAs. Absolute prediction error and intraclass correlation were used to assess the accuracy of the HKAA estimates.
Results
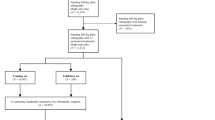
The study cohort consisted of 528 radiographs from 517 patients (mean age = 10.8 years, SD = 4.2 years). Evaluation of this CNN showed few HKAA prediction outliers (12/1056 [1.1%]), defined as having an absolute prediction error of > 10°. Excluding these outliers, the study cohort’s mean absolute prediction error for the HKAA was 0.94° ± 0.84°, and the intraclass correlation between the ground truth and prediction was 0.974.
Conclusion
The proposed CNN generated promising results and offers potential for using this model as a computer-aided diagnostic tool.





Similar content being viewed by others
References
Espandar R, Mortazavi SM, Baghdadi T. Angular deformities of the lower limb in children. Asian J Sports Med. 2009;1:46–53. https://doi.org/10.5812/asjsm.34871.
Greene WB. Genu varum and genu valgum in children: differential diagnosis and guidelines for evaluation. Comp Ther. 1996;22:22–9.
Shopfner CE, Coin CG. Genu varus and valgus in children. Radiology. 1969;92:723–32. https://doi.org/10.1148/92.4.723.
Brouwer GM, van Tol AW, Bergink AP, et al. Association between valgus and varus alignment and the development and progression of radiographic osteoarthritis of the knee. Arthritis Rheum. 2007;56:1204–11. https://doi.org/10.1002/art.22515.
Sabharwal S, Zhao C. The hip-knee-ankle angle in children: reference values based on a full-length standing radiograph. J Bone Joint Surg Am. 2009;91:2461–568. https://doi.org/10.2106/jbjs.i.00015.
Cherian JJ, Kapadia BH, Banerjee S, et al. Mechanical, anatomical, and kinematic axis in TKA: concepts and practical applications. Curr Rev Musculoskeleta Med. 2014;7:89–95. https://doi.org/10.1007/s12178-014-9218-y.
Moreland JR, Bassett LW, Hanker GJ. Radiographic analysis of the axial alignment of the lower extremity. J Bone Joint Surg. 1987;69:745–9. https://doi.org/10.2106/00004623-198769050-00016.
LeCun Y, Boser B, Denker JS, et al. Backpropagation applied to handwritten zip code recognition. Neural Comput. 1989;1:541–51. https://doi.org/10.1162/neco.1989.1.4.541.
LeCun Y, Bottou L, Bengio Y, Haffner P. Gradient-based learning applied to document recognition. Proc IEEE. 1998;86:2278–323. https://doi.org/10.1109/5.726791.
LeCun Y, Kavukcuoglu K, Farabet CC. Convolutional networks and applications in vision. In: Proc. IEEE International Symposium on Circuits and Systems 2010: 253–256. https://doi.org/10.1109/ISCAS.2010.5537907
Khan A, Sohail A, Zahoora U, Oureshi AS. A survey of the recent architectures of deep convolutional neural networks. Artif Intell Rev. 2020. https://doi.org/10.1007/s10462-020-09825-6.
Tompson J, Jain A, LeCun Y, Bregler C. Joint training of a convolutional network and a graphical model for human pose estimation. In: Adv Neural Inf Process Syst 2014: 1799–1807. https://doi.org/10.5555/2968826.2969027
Pfister T, Charles J, Zisserman A. Flow ConvNets for human pose estimation in videos. In: Proc Int Conf Comput Vis 2015: 1913–1921. https://doi.org/10.1109/ICCV.2015.222
Newell A, Yang K, Deng J. Stacked hourglass networks for human pose estimation. In: Proc European Conference on Computer Vision 2018: 734–750. https://doi.org/10.1007/978-3-319-46484-8_29
Mahpod S, Das R, Maiorana E, Keller Y, Campisi P. Facial landmark point localization using coarse-to-fine deep recurrent neural network. ArXiv 2018, abs/1805.01760. https://arxiv.org/abs/1805.01760v2
Earp SWF, Samacoits A, Jain S, Noinongyao P, Boonpunmongkol S. Sub-pixel face landmarks using heatmaps and a bag of tricks. ArXiv 2021, abs/2103.03059. https://arxiv.org/abs/2103.03059v2
Law H, Deng J. Cornernet: detecting objects as paired keypoints. Int J Comput Vis. 2019;128:642–56. https://doi.org/10.1007/s11263-019-01204-1.
Yi J, Wu P, Huang Q, Qu H, Metaxas DN. Vertebra-focused landmark detection for scoliosis assessment. In: IEEE International Symposium on Biomedical Imaging 2020: 736–740. https://doi.org/10.1109/ISBI45749.2020.9098675
Tsai A. Anatomical landmark localization via convolutional neural networks for limb length discrepancy measurements. Pediatr Radiol. 2021;51:1431-1447. https://doi.org/10.1007/s00247-021-05004-z
Payer C, Stern D, Bischof H, Urschler M. Integrating spatial configuration into heatmap regression based CNNs for landmark localization. Med Image Anal. 2019;54:207–19. https://doi.org/10.1016/j.media.2019.03.007.
Shorten C, Khoshgoftaar TM. A survey on image data augmentation for deep learning. J Big Data. 2019;6:1–48. https://doi.org/10.1186/s40537-019-0197-0.
Raschka S. Model evaluation, model selection, and algorithm selection in machine learning. CoRR 2018: abs/1811.12808. http://arxiv.org/abs/1811.12808
Cicchetti DV. Guidelines, criteria, and rules of thumb for evaluating normed and standardized assessment instruments in psychology. Psychol Assess. 1994;6:284–90. https://doi.org/10.1037/1040-3590.6.4.284.
Schock J, Truhn D, Abrar DB, et al. Automated analysis of alignment in long-leg radiographs using a fully automated support system based on artificial intelligence. Radiology: Artificial Intelligence 2021: Epub ahead of print. https://doi.org/10.1148/ryai.2020200198
Acknowledgements
The author would like to thank Ms. Nancy Drinan for her help in the editing of the manuscript. The author would also like to acknowledge the use of Boston Children’s Hospital's HighPerformance Computing Resources BCH HPC Cluster Enkefalos 2 (E2) which has been crucial to the research reported in this publication. The software used in the project was installed and configured by BioGrids.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
Approval was obtained by the local institutional review board at Boston Children’s Hospital, Harvard Medical School in Boston, Massachusetts, USA (protocol number IRB- P00037876).
Informed consent
Written informed consent was waived by the institutional review board at Boston Children’s Hospital.
Conflict of interest
The author declares no competing interests.
Study subjects or cohorts overlap
The study subjects and cohorts have not been previously reported.
Guarantor
The scientific guarantor of this publication is Andy Tsai. He agrees to be accountable for all aspects of this work.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tsai, A. A deep learning approach to automatically quantify lower extremity alignment in children. Skeletal Radiol 51, 381–390 (2022). https://doi.org/10.1007/s00256-021-03844-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00256-021-03844-2