Abstract
Background
Measurement of limb-length discrepancy (LLD) from a radiograph is a cognitively simple but time-consuming task.
Objective
To develop a convolutional neural network (CNN) to localize anatomical landmarks from full-length lower-extremity radiographs in predicting LLD.
Materials and methods
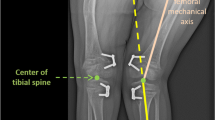
The author searched a hospital’s image database to identify studies performed between Feb. 1, 2016, and Sept. 30, 2019. Inclusion criteria were: (1) patients ≤21 years old, (2) study indication of LLD, (3) full-length lower-extremity anteroposterior radiographs performed on the EOS system, and (4) imaging field-of-view that included entire bilateral femurs and tibias. The six requisite ground truth anatomical landmarks for measuring LLD from each radiograph — bilateral top of femoral heads, medial femoral condyles, and center of tibial plafonds — were manually labeled by a pediatric radiologist. For each landmark, a two-dimensional heatmap was generated to encode the pseudo-probability of a landmark being at a particular spatial location. A CNN was developed that regressed across a collection of these heatmaps for landmark localization and bone length predictions.
Results
The study cohort consisted of 504 full-length lower-extremity radiographs from 359 patients with wide ranging skeletal deformities and in situ hardware. Evaluation of this CNN showed that the mean point-error for the predicted top of femoral head, medial femoral condyle, and center of tibial plafond were 0.37 cm, 0.39 cm and 0.42 cm, respectively. The mean absolute error for the predicted femoral, tibial and limb lengths, and LLD were 0.33 cm, 0.30 cm, 0.30 cm, and 0.36 cm, respectively. Predicted bone lengths correlated strongly with ground truth.
Conclusion
This prototype CNN delivered promising results in predicting bone lengths from full-length lower-extremity radiographs and offers the potential use of a computer algorithm to predict LLD.












Similar content being viewed by others
References
Khamis S, Carmeli E (2017) Relationship and significance of gait deviations associated with limb length discrepancy: a systematic review. Gait Posture 57:115–123
Raczkowski JW, Daniszewska B, Zolynski K (2010) Functional scoliosis caused by leg length discrepancy. Arch Med Sci 6:393–398
Sheha ED, Steinhaus ME, Kim HJ et al (2018) Leg-length discrepancy, functional scoliosis, and low back pain. JBJS Rev 6:e6
Rannisto S, Okuloff A, Uitti J et al (2015) Leg-length discrepancy is associated with low back pain among those who must stand while working. BMC Musculoskelet Disord 16:11
Murray KJ, Azari MF (2015) Leg length discrepancy and osteoarthritis in the knee, hip and lumbar spine. J Can Chiropr Assoc 59:226–237
Cleveland RH, Kushner DC, Ogden MC et al (1988) Determination of leg length discrepancy. A comparison of weight-bearing and supine imaging. Investig Radiol 23:301–304
Lampe HI, Swierstra BA, Diepstraten AF (1996) Measurement of limb length inequality. Comparison of clinical methods with orthoradiography in 190 children. Acta Orthop Scand 67:242–244
Terry MA, Winell JJ, Green DW et al (2005) Measurement variance in limb length discrepancy: clinical and radiographic assessment of interobserver and intraobserver variability. J Pediatr Orthop 25:197–201
Sabharwal S, Zhao C, McKeon J et al (2006) Computed radiographic measurement of limb length discrepancy — full length standing antero-posterior radiograph versus scanograms. J Bone Joint Surg Am 88:2243–2251
Escott BG, Ravi B, Weathermon AC et al (2013) EOS low-dose radiography: a reliable and accurate upright assessment of lower-limb lengths. J Bone Joint Surg Am 95:e1831–e1837
Garner MR, Dow M, Bixby E et al (2016) Evaluating length: the use of low-dose biplanar radiography (EOS) and tantalum bead implantation. J Pediatr Orthop 36:e6–e9
LeCun Y, Boser B, Denker JS et al (1989) Backpropagation applied to handwritten ZIP code recognition. Neural Comput 1:541–551
LeCun Y, Bottou L, Bengio Y, Haffner P (1998) Gradient-based learning applied to document recognition. Proc IEEE 86:2278–2323
LeCun Y, Kavukcuoglu K, Farabet CC (2010) Convolutional networks and applications in vision. Proc IEEE Int Symp Circ Syst 2010:253–256
Khan A, Sohail A, Zahoora U, Oureshi AS (2020) A survey of the recent architectures of deep convolutional neural networks. Artif Intell Rev 53:5455–5516
Ciresan D, Meier U, Masci J, Schmidhuber J (2012) Multi-column deep neural network for traffic sign classification. Neural Netw 32:333–338
Liu X, Deng Z, Yang Y (2019) Recent progress in semantic image segmentation. Artif Intell Rev 52:1089–1106
Setio AA, Ciompi F, Litjens G et al (2016) Pulmonary nodule detection in CT images: false positive reduction using multi-view convolutional networks. IEEE Trans Med Imaging 35:1160–1169
Yasaka K, Akai H, Abe O, Kiryu S (2018) Deep learning with convolutional neural network for differentiation of liver masses at dynamic contrast-enhanced CT: a preliminary study. Radiology 286:887–896
Kurata Y, Nishio M, Kido A et al (2019) Automatic segmentation of the uterus on MRI using a convolutional neural network. Comput Biol Med 114:103438
Milletari F, Navab N, Ahmadi S (2016) V-net: fully convolutional neural networks for volumetric medical image segmentation. In: Proceedings of the Fourth International Conference on 3D Vision, pp 565–571
Lu F, Wu F, Hu P et al (2017) Automatic 3D liver location and segmentation via convolutional neural network and graph cut. Int J Comput Assist Radiol Surg 12:171–182
Lakhani P, Sundaram B (2017) Deep learning at chest radiography: automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology 284:574–582
Kooi T, Litjens G, van Ginneken B et al (2017) Large scale deep learning for computer aided detection of mammographic lesions. Med Image Anal 35:303–312
Tompson J, Jain A, LeCun Y, Bregler C (2014) Joint training of a convolutional network and a graphical model for human pose estimation. Proc Adv Neural Inf Proc Syst 1:1799–1807
Newell A, Yang K, Deng J (2018) Stacked hourglass networks for human pose estimation. In: Proceedings of the European Conference on Computer Vision, pp 734–750
Law H, Deng J (2019) Cornernet: detecting objects as paired keypoints. Int J Comput Vis 128:642–656
Yi J, Wu P, Huang Q, Qu H, Metaxas DN (2020) Vertebra-focused landmark detection for scoliosis assessment. In: IEEE International Symposium on Biomedical Imaging, pp 736–740
Toshev A, Szegedy C (2014) DeepPose: human pose estimation via deep neural networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp 1653–1660
Zhang J, Liu M, Shen D (2017) Detecting anatomical landmarks from limited medical image data using two-stage task-oriented deep neural networks. IEEE Trans Image Proc 26:4753–4764
Pfister T, Charles J, Zisserman A (2015) Flowing ConvNets for human pose estimation in videos. In: Proceedings of the IEEE International Conference on Computer Vision, pp 1913–1921
Payer C, Stern D, Bischof H, Urschler M (2019) Integrating spatial configuration into heatmap regression based CNNs for landmark localization. Med Image Anal 54:207–219
He K, Zhang X, Ren S, Sun J (2015) Delving deep into rectifiers: surpassing human-level performance on ImageNet classification. In: Proceedings of the IEEE International Conference on Computer Vision: pp 1026–1034
Qian N (1999) On the momentum term in gradient descent learning algorithms. Neural Netw 12:145–151
Raschka S (2018) Model evaluation, model selection, and algorithm selection in machine learning. http://arxiv.org/abs/1811.12808
Zheng Q, Shellikeri S, Huang H et al (2020) Deep learning measurement of leg length discrepancy in children based on radiographs. Radiology 296:152–158
Roh Y, Heo G, Whang SE (2019) A survey on data collection for machine learning: a big data–AI integration perspective. IEEE Trans Knowl Data Eng. https://doi.org/10.1109/TKDE.2019.2946162
Shorten C, Khoshgoftaar TM (2019) A survey on image data augmentation for deep learning. J Big Data 6:1–48
Liu D, Zhou KS, Bernhardt D, Comaniciu D (2010) Search strategies for multiple landmark detection by submodular maximization. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp 2831–2838
Ebner T, Stern D, Donner R et al (2014) Towards automatic bone age estimation from MRI: localization of 3D anatomical landmarks. Med Image Comput Assist Interv 2014:421–428
Pan SJ, Yang Q (2018) A survey on transfer leaning. IEEE Trans Knowl Data Eng 22:1345–1359
Acknowledgments
The author would like to thank Ms. Nancy Drinan for her help in the editing of the manuscript. The author would also like to acknowledge the use of Boston Children’s Hospital’s High-Performance Computing Resources BCH HPC Cluster Enkefalos 2 (E2), which has been crucial to the research reported in this publication. Software used in the project was installed and configured by BioGrids.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tsai, A. Anatomical landmark localization via convolutional neural networks for limb-length discrepancy measurements. Pediatr Radiol 51, 1431–1447 (2021). https://doi.org/10.1007/s00247-021-05004-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00247-021-05004-z