Abstract
As an important semi-essential amino acid, l-arginine (l-Arg) has important application prospects in medicine and health care. However, it remains a challenge to efficiently produce l-Arg by Escherichia coli (E. coli). In the present study, we obtained an E. coli A1 with l-Arg accumulation ability, and carried out a series of metabolic engineering on it, and finally obtained an E. coli strain A7 with high l-Arg production ability. First, genome analysis of strain A1 was performed to explore the related genes affecting l-Arg accumulation. We found that gene speC and gene speF played an important role in the accumulation of l-Arg. Second, we used two strategies to solve the feedback inhibition of the l-Arg pathway in E. coli. One was the combination of a mutation of the gene argA and the deletion of the gene argR, and the other was the combination of a heterologous insertion of the gene argJ and the deletion of the gene argR. The combination of exogenous argJ gene insertion and argR gene deletion achieved higher titer accumulation with less impact on strain growth. Finally, we inserted the gene cluster argCJBDF of Corynebacterium glutamicum (C. glutamicum) to enhance the metabolic flux of the l-Arg pathway in E. coli. The final strain obtained 70.1 g/L l-Arg in a 5-L bioreactor, with a yield of 0.326 g/g glucose and a productivity of 1.17 g/(L· h). This was the highest level of l-Arg production by E. coli ever reported. Collectively, our findings provided valuable insights into the possibility of the industrial production of l-Arg by E. coli.
Key points
• Genetic background of E. coli A1 genome analysis.
• Heterologous argJ substitution of argA mutation promoted excessive accumulation of L-Arg in E. coli A1.
• The overexpression of L-Arg synthesis gene cluster argCJBDF of Corynebacterium glutamicum (C. glutamate) promoted the accumulation of L-Arg, and 70.1 g/L L-Arg was finally obtained in fed-batch fermentation.
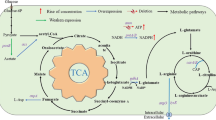
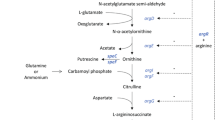
Graphical abstract







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in the published article. Construction of recombinant plasmids and strains; primer pairs used in this study; Analyzing of l-Arg production by shake-flask fermentation.
References
Applebaum D, Sabo DL, Fischer EH, Morris DR (1975) Biodegradative ornithine decarboxylase of Escherichia coli. Purification, properties, and pyridoxal 5’-phosphate binding site. Biochemistry 14(16):3675–3681. https://doi.org/10.1021/bi00687a025
Applebaum DM, Dunlap JC, Morris DR (1977) Comparison of the biosynthetic and biodegradative ornithine decarboxylases of Escherichia coli. Biochemistry 16(8):1580–1584. https://doi.org/10.1021/bi00627a008
Alexander S, Christopher S, Sandra Z, Philipp S, Gottfried U (2018) Dcua of aerobically grown Escherichia coli serves as a nitrogen shuttle (l-aspartate/fumarate) for nitrogen uptake. Mol Microbiol 109(6):801–811. https://doi.org/10.1111/mmi.14074
Charlier D, Bervoets I (2019) Regulation of arginine biosynthesis, catabolism and transport in Escherichia coli. Amino Acids 51(8):1103–1127. https://doi.org/10.1007/s00726-019-02757-8
Cherney LT, Cherney MM, Garen CR, James MNG (2010) Crystal structure of the intermediate complex of the arginine repressor from mycobacterium tuberculosis bound with its dna operator reveals detailed mechanism of arginine repression. J Mol Biol 399(2):240–254. https://doi.org/10.1016/j.jmb.2010.03.065
Caldara M, Dupont G, Leroy F, Goldbeter A, Vuyst L, Cunin R (2008) Arginine biosynthesis in Escherichia coli: experimental perturbation and mathematical modeling. J Biol Chem. 283(10):6347–6358. https://doi.org/10.1074/jbc.M705884200
Celis RT (1999) Repression and activation of arginine transport genes in Escherichia coli K12 by the ArgP protein. J Mol Biol 294(5):1087–1095. https://doi.org/10.1006/jmbi.1999.3308
Chen XL, Zhang B, Tang L, Jiao HT, Yi XH, Feng XU, Hong X, Hua W, Hua XY (2014) Expression and characterization of ArgR, an arginine regulatory protein in Corynebacterium crenatum. Biomed Environ Sci 27(6):436–443. https://doi.org/10.3967/bes2014.072
Chen Y, Ho J, Shis DL, Gupta C, Long J, Wagner DS, Ott W, Josić K, Bennett MR (2018) Tuning the dynamic range of bacterial promoters regulated by ligand-inducible transcription factors. Nat Commun 9(1):64. https://doi.org/10.1038/s41467-017-02473-5
Dou W, Xu M, Cai D, Zhang X, Rao Z, Xu Z (2011) Improvement of L-arginine production by overexpression of a bifunctional ornithine acetyltransferase in Corynebacterium crenatum. Biotechnol Appl Biochem 165(3–4):845–855. https://doi.org/10.1007/s12010-011-9302-3
Ginesy M, Belotserkovsky J, Enman J, Isaksson L, Rova U (2015) Metabolic engineering of Escherichia coli for enhanced arginine biosynthesis. Microb Cell Fact 14(1):1–11. https://doi.org/10.1186/s12934-015-0211-y
Ginésy M, Rusanova-Naydenova D, Rova U (2017) Tuning of the carbon-to-nitrogen ratio for the roduction of L-arginine by Escherichia coli. Fermentation 3(4):60. https://doi.org/10.3390/fermentation3040060
Gruer MJ, Guest JR (1994) Two genetically-distinct and differentially-regulated aconitases (acna and acnb) in Escherichia coli. Microbiology 140(10):2531–2541. https://doi.org/10.1099/00221287-140-10-2531
Jiang Y, Sheng Q, Wu XY, Ye BC, Zhang B (2020) L-arginine production in Corynebacterium glutamicum: manipulation and optimization of the metabolic process. Crit Rev Biotechnol 245(3):1–14. https://doi.org/10.1080/07388551.2020.1844625
Kaya H, Bulut O, Kamali AR, Ege D (2018) L-arginine modified multi-walled carbon nanotube/sulfonated poly (ether ether ketone) nanocomposite films for biomedical applications. Appl Surf Sci 444:168–176. https://doi.org/10.1016/j.apsusc.2018.03.046
Krzystek-Korpacka M, Wiśniewski J, Fleszar MG, Bednarz-Misa I, Bronowicka-Szydełko A, Gacka M, Masłowski L, Kędzior K, Witkiewicz W (2019) Gamian A (2019) Metabolites of the Nitric Oxide (NO) Pathway are altered and indicative of reduced NO and arginine bioavailability in patients with cardiometabolic diseases complicated with chronic wounds of lower extremities: targeted metabolomics approach (LC-MS/MS). Oxid Med Cell Longev 10:1–13. https://doi.org/10.1155/2019/5965721
Kathrin P, Frederik W, Marcus P, Christian R, Jörn K (2013) A novel type of N-acetylglutamate synthase is involved in the first step of arginine biosynthesis in Corynebacterium glutamicum. BMC Genomics 14(1):1–15. https://doi.org/10.1186/1471-2164-14-713
Kim SK, Seong W, Han GH, Le DH, Lee SG (2017) Crispr interference-guided multiplex repression of endogenous competing pathway genes for redirecting metabolic flux in Escherichia coli. Microb Cell Fact 16(1):1–15. https://doi.org/10.1186/s12934-017-0802-x
Lu CD (2006) Pathways and regulation of bacterial arginine metabolism and perspectives for obtaining arginine overproducing strains. Appl Microbiol Biotechnol 70(3):261–272. https://doi.org/10.1007/s00253-005-0308-z
Long M, Xu M, Ma Z, Pan X, You J, Hu M, Shao Y, Yang T, Zhang X, Rao Z (2020) Significantly enhancing production of trans -4-hydroxy-l-proline by integrated system engineering in Escherichia coli. Sci Adv 6(21):eaba2383. https://doi.org/10.1126/sciadv.aba2383
Li Y, Lin Z, Huang C, Zhang Y, Wang Z, Tang YJ, Chen T, Zhao X (2015) Metabolic engineering of Escherichia coli using CRISPR-Cas9 meditated genome editing. Metab Eng 31:13–21. https://doi.org/10.1016/j.ymben.2015.06.006
Min L, Jin Z, Caldovic L, Morizono H, Allewell NM, Tuchman M, Shi D (2009) Mechanism of allosteric inhibition of N -acetyl-L-glutamate synthase by L-arginine. J Biol Chem 284(8):4873–4880. https://doi.org/10.1074/jbc.M805348200
Majumdar R, Barchi B, Turlapati SA, Gagne M, Minocha R, Long S, Minocha SC (2016) Glutamate, ornithine, arginine, proline, and polyamine metabolic interactions: the pathway is regulated at the post-transcriptional level. In Front Plant Sci 7(175):475. https://doi.org/10.3389/fpls.2016.00078
Nandineni MR, Gowrishankar J (2004) Evidence for an arginine exporter encoded by yggA (ArgO) that is regulated by the LysR-Type transcriptional regulator ArgP in Escherichia coli. J Bacteriol 186(11):3539–3546. https://doi.org/10.1128/JB.186.11.3539-3546.2004
Park SH, Kim HU, Kim TY, Park JS, Kim SS, Lee SY (2014) Metabolic engineering of Corynebacterium glutamicum for L-arginine production. Nat Commun 5(1):1–9. https://doi.org/10.1038/ncomms5618
Pandey SK, Melichercik M, Carey J (2020) Conserved dynamic mechanism of allosteric response to L-arg in divergent bacterial arginine repressors. Molecules 25(9):2247. https://doi.org/10.3390/molecules25092247
Rajagopal BS, Deponte J, Tuchman M, Malamy MH (1998) Use of inducible feedback-resistant n-acetylglutamate synthetase (argA) genes for enhanced arginine biosynthesis by genetically engineered Escherichia coli K12 strains. Appl Environ Microbiol 64(5):1805–1811. https://doi.org/10.1016/S1383-5718(98)00029-1
Shi D, Norma A, Mendel T (2015) The N-Acetylglutamate synthase family: structures, function and mechanisms. Int J Mol Sci 16(12):13004–13022. https://doi.org/10.3390/ijms160613004
Schneider J, Niermann K, Wendisch VF (2011) Production of the amino acids L-glutamate, L-lysine, L-ornithine and L-arginine from arabinose by recombinant Corynebacterium glutamicum. J Biotechnol 154(2–3):191–198. https://doi.org/10.1016/j.jbiotec.2010.07.009
Sander T, Farke N, Diehl C, Kuntz M, Glatter T, Link H (2019a) Allosteric feedback inhibition enables robust amino acid biosynthesis in E. coli by enforcing enzyme overabundance. Cell Syst 8(1):66–75. https://doi.org/10.1016/j.cels.2018.12.005
Sander T, Wang CY, Glatter T, Link H (2019b) Crispri-based downregulation of transcriptional feedback improves growth and metabolism of arginine overproducing E. coli. ACS Synth Biol 8(9):1983–1990. https://doi.org/10.1021/acssynbio.9b00183
Utagawa T (2004) Production of arginine by fermentation. J Nutr 134(10):2854S-2857S. https://doi.org/10.1093/jn/134.10.2854S
Vyas S, Maas WK (1994) The arginine repressor of Escherichia coli. Arch Biochem Biophys 100(3):542–546. https://doi.org/10.1016/0003-9861(63)90124-3
Wade M (2015) High-throughput silencing using the crispr-cas9 system: a revie of the benefits and challenges. J Biomol Screen 20(8):1027–1039. https://doi.org/10.1177/1087057115587916
Wang T, Tague N, Whelan SA, Dunlop MJ (2021) Programmable gene regulation for metabolic engineering using decoy transcription factor binding sites. Rucleic Acids Res 49(2):1163–1172. https://doi.org/10.1093/nar/gkaa1234
Xingyue B, Feifei W, Rendi Y, Yan Z, Linglin F, Yanbo W (2020) Ornithine decarboxylation (odc) system of shewanella baltica regulates putrescine production and acid resistance. J Food Prot 84(2):303–309. https://doi.org/10.4315/JFP-20-227
Yoshida H, Araki K, Nakayama K (1981) L-arginine production by arginine analog-resistant mutants of microorganisms. Agri Biol Chem 45(4):959–963. https://doi.org/10.1080/00021369.1981.10864636
Funding
This work was supported by the Green Bio-fabrication Program of China [No. 2021YFC2100900], the Top-Notch Academic Programs Project of Jiangsu Higher Education Institutions, the 111 Project (Grant number 111–2-06).
Author information
Authors and Affiliations
Contributions
JX and HW conceived the experiments. HW and WZ designed and performed the experiments and analyzed the data. HW and JX wrote the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, HD., Xu, JZ. & Zhang, WG. Metabolic engineering of Escherichia coli for efficient production of l-arginine. Appl Microbiol Biotechnol 106, 5603–5613 (2022). https://doi.org/10.1007/s00253-022-12109-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12109-4