Abstract
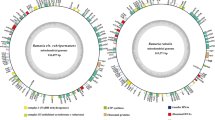
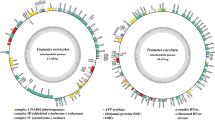
In the present study, the complete mitogenome of Turbinellus floccosus was sequenced, assembled, and compared with other basidiomycete mitogenomes. The mitogenome of T. floccosus consists of a circular DNA molecule, with a size of 62,846 bp. Gene arrangement analysis indicated that large-scale gene rearrangements occurred in the levels of family and genus of basidiomycete species, and the mitogenome of T. floccosus contained a unique gene order. A significant correlation between the number of introns and the mitochondrial genome size of Basidiomycota were detected (P < 0.01). A total of 896 introns were detected in the core protein-coding genes (PCGs) of 74 basidiomycete species, and the cox1 gene was the largest host gene of basidiomycete introns. Intron position class (Pcls) P383 in the cox1 gene was the most common intron in Basidiomycota, which distributed in 40 of 74 basidiomycete species. In addition, frequent intron loss/gain events were detected in basidiomycete species. More than 50% of bases around insertion sites (− 15 bp to 15 bp) of Pcls from different species were conservative, indicating site preferences of intron insertions in Basidiomycota. Further analysis showed that 76.09% of introns tended to insert downstream to a T base in Basidiomycota. Phylogenetic analysis for 74 basidiomycetes indicated mitochondrial genes are effective molecular markers for phylogeny of basidiomycetes. The study served as the first report on the mitogenome from the family Gomphaceae, which will help to understand the intron origin and evolution in Basidiomycota.
Key points
• The mitogenome of Turbinellus floccosus had a unique gene arrangement.
• Intron loss/gain events were detected in the 74 basidiomycete species.
• Introns tend to insert downstream of a T base in basidiomycete mitogenomes.









Similar content being viewed by others
Data availability
The complete mitogenome of T. floccosus was deposited in the GenBank database under the accession number MN623377.
References
Aguileta G, de Vienne DM, Ross ON, Hood ME, Giraud T, Petit E, Gabaldon T (2014) High variability of mitochondrial gene order among fungi. Genome Biol Evol 6(2):451–465. https://doi.org/10.1093/gbe/evu028
Allen JF (2015) Why chloroplasts and mitochondria retain their own genomes and genetic systems: colocation for redox regulation of gene expression. Proc Natl Acad Sci U S A 112(33):10231–10238. https://doi.org/10.1073/pnas.1500012112
Andersen MM, Balding DJ (2018) How many individuals share a mitochondrial genome? PLoS Genetics 14(11):e1007774. https://doi.org/10.1371/journal.pgen.1007774
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19(5):455–477. https://doi.org/10.1089/cmb.2012.0021
Bernt M, Donath A, Juhling F, Externbrink F, Florentz C, Fritzsch G, Putz J, Middendorf M, Stadler PF (2013) MITOS: improved de novo metazoan mitochondrial genome annotation. Mol Phylogenet Evol 69(2):313–319. https://doi.org/10.1016/j.ympev.2012.08.023
Bleasby AJ, Wootton JC (1990) Construction of validated, non-redundant composite protein sequence databases. Protein Eng 3(3):153–159
Boore JL (1999) Animal mitochondrial genomes. Nucleic Acids Res 27(8):1767–1780
Bronstein O, Kroh A (2018) The first mitochondrial genome of the model echinoid Lytechinus variegatus and insights into Odontophoran phylogenetics. Genomics 111(4):710–718. https://doi.org/10.1016/j.ygeno.2018.04.008
Cameron SL (2014) Insect mitochondrial genomics: implications for evolution and phylogeny. Annu Rev Entomol 59:95–117. https://doi.org/10.1146/annurev-ento-011613-162007
Chen C, Li Q, Fu R, Wang J, Deng G, Chen X, Lu D (2021) Comparative mitochondrial genome analysis reveals intron dynamics and gene rearrangements in two Trametes species. Sci Rep 11(1):2569. https://doi.org/10.1038/s41598-021-82040-7
Cieslak M, Pruvost M, Benecke N, Hofreiter M, Morales A, Reissmann M, Ludwig A (2010) Origin and history of mitochondrial DNA lineages in domestic horses. PLoS One 5(12):e15311. https://doi.org/10.1371/journal.pone.0015311
Coordinators NR (2017) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 45(D1):D12–D17. https://doi.org/10.1093/nar/gkx1071
Costa GG, Cabrera OG, Tiburcio RA, Medrano FJ, Carazzolle MF, Thomazella DP, Schuster SC, Carlson JE, Guiltinan MJ, Bailey BA, Mieczkowski P, Pereira GA, Meinhardt LW (2012) The mitochondrial genome of Moniliophthora roreri, the frosty pod rot pathogen of cacao. Fungal Biol 116(5):551–562. https://doi.org/10.1016/j.funbio.2012.01.008
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14(6):1188–1190. https://doi.org/10.1101/gr.849004
du Toit Z, du Plessis M, Dalton DL, Jansen R, Paul Grobler J, Kotze A (2017) Mitochondrial genomes of African pangolins and insights into evolutionary patterns and phylogeny of the family Manidae. BMC Genomics 18(1):746. https://doi.org/10.1186/s12864-017-4140-5
Ferandon C, Moukha S, Callac P, Benedetto JP, Castroviejo M, Barroso G (2010) The Agaricus bisporus cox1 gene: the longest mitochondrial gene and the largest reservoir of mitochondrial group i introns. PLoS One 5(11):e14048. https://doi.org/10.1371/journal.pone.0014048
Ferandon C, Xu J, Barroso G (2013) The 135 kbp mitochondrial genome of Agaricus bisporus is the largest known eukaryotic reservoir of group I introns and plasmid-related sequences. Fungal Genet Biol 55:85–91. https://doi.org/10.1016/j.fgb.2013.01.009
Fourie G, Van der Merwe NA, Wingfield BD, Bogale M, Wingfield MJ, Steenkamp ET (2018) Mitochondrial introgression and interspecies recombination in the Fusarium fujikuroi species complex. IMA Fungus 9(1):37–48. https://doi.org/10.5598/imafungus.2018.09.01.04
Gan HM, Thomas BN, Cavanaugh NT, Morales GH, Mayers AN, Savka MA, Hudson AO (2017) Whole genome sequencing of Rhodotorula mucilaginosa isolated from the chewing stick (Distemonanthus benthamianus): insights into Rhodotorula phylogeny, mitogenome dynamics and carotenoid biosynthesis. PeerJ Preprints 5:e3219v2. https://doi.org/10.7287/peerj.preprints.3219v2
Giachini AJ, Camelini CM, Rossi MJ, Soares CRFS, Trappe JM (2012) Systematics of the Gomphales: the genus Gomphus sensu stricto. Mycotaxon 120:385–400. https://doi.org/10.5248/120.385
Giachini AJ, Castellano MA (2011) A new taxonomic classification for species in Gomphus sensu lato. Mycotaxon 115:183–201. https://doi.org/10.5248/115.183
Giachini AJ, Hosaka K, Nouhra E, Spatafora J, Trappe JM (2010) Phylogenetic relationships of the Gomphales based on nuc-25S-rDNA, mit-12S-rDNA, and mit-atp6-DNA combined sequences. Fungal Biol 114(2-3):224–234. https://doi.org/10.1016/j.funbio.2010.01.002
Gonzalez-Avila PA, Luna-Vega I, Rios MV, Saade RL, Blanco JC (2013) Current knowledge and importance of the order Gomphales (Fungi: Basidiomycota) in Mexico. Nova Hedwigia 97(1-2):55–86. https://doi.org/10.1127/0029-5035/2013/0099
Gorman GS, Schaefer AM, Ng Y, Gomez N, Blakely EL, Alston CL, Feeney C, Horvath R, Yu-Wai-Man P, Chinnery PF, Taylor RW, Turnbull DM, McFarland R (2015) Prevalence of nuclear and mitochondrial DNA mutations related to adult mitochondrial disease. Ann Neurol 77(5):753–759. https://doi.org/10.1002/ana.24362
Gu X, Kang X, Liu J (2019) Mutation signatures in germline mitochondrial genome provide insights into human mitochondrial evolution and disease. Hum Genet 138(6):613–624. https://doi.org/10.1007/s00439-019-02009-5
Hahn C, Bachmann L, Chevreux B (2013) Reconstructing mitochondrial genomes directly from genomic next-generation sequencing reads--a baiting and iterative mapping approach. Nucleic Acids Res 41(13):e129. https://doi.org/10.1093/nar/gkt371
Hibbett DS (2006) A phylogenetic overview of the Agaricomycotina. Mycologia 98(6):917–925. https://doi.org/10.3852/mycologia.98.6.917
Hibbett DS, Binder M, Bischoff JF, Blackwell M, Cannon PF, Eriksson OE, Huhndorf S, James T, Kirk PM, Lucking R, Thorsten Lumbsch H, Lutzoni F, Matheny PB, McLaughlin DJ, Powell MJ, Redhead S, Schoch CL, Spatafora JW, Stalpers JA, Vilgalys R, Aime MC, Aptroot A, Bauer R, Begerow D, Benny GL, Castlebury LA, Crous PW, Dai YC, Gams W, Geiser DM, Griffith GW, Gueidan C, Hawksworth DL, Hestmark G, Hosaka K, Humber RA, Hyde KD, Ironside JE, Koljalg U, Kurtzman CP, Larsson KH, Lichtwardt R, Longcore J, Miadlikowska J, Miller A, Moncalvo JM, Mozley-Standridge S, Oberwinkler F, Parmasto E, Reeb V, Rogers JD, Roux C, Ryvarden L, Sampaio JP, Schussler A, Sugiyama J, Thorn RG, Tibell L, Untereiner WA, Walker C, Wang Z, Weir A, Weiss M, White MM, Winka K, Yao YJ, Zhang N (2007) A higher-level phylogenetic classification of the Fungi. Mycol Res 111(Pt 5):509–547. https://doi.org/10.1016/j.mycres.2007.03.004
Himmelstrand K, Olson A, Brandstrom Durling M, Karlsson M, Stenlid J (2014) Intronic and plasmid-derived regions contribute to the large mitochondrial genome sizes of Agaricomycetes. Curr Genet 60(4):303–313. https://doi.org/10.1007/s00294-014-0436-z
Huang W, Feng H, Tu W, Xiong C, Jin X, Li P, Wang X, Li Q (2021) Comparative mitogenomic analysis reveals dynamics of intron within andbetween Tricholoma species and phylogeny of Basidiomycota. Front Genet 12: 534871. https://doi.org/10.3389/fgene.2021.534871
James TY, Kauff F, Schoch CL, Matheny PB, Hofstetter V, Cox CJ, Celio G, Gueidan C, Fraker E, Miadlikowska J, Lumbsch HT, Rauhut A, Reeb V, Arnold AE, Amtoft A, Stajich JE, Hosaka K, Sung GH, Johnson D, O'Rourke B, Crockett M, Binder M, Curtis JM, Slot JC, Wang Z, Wilson AW, Schussler A, Longcore JE, O'Donnell K, Mozley-Standridge S, Porter D, Letcher PM, Powell MJ, Taylor JW, White MM, Griffith GW, Davies DR, Humber RA, Morton JB, Sugiyama J, Rossman AY, Rogers JD, Pfister DH, Hewitt D, Hansen K, Hambleton S, Shoemaker RA, Kohlmeyer J, Volkmann-Kohlmeyer B, Spotts RA, Serdani M, Crous PW, Hughes KW, Matsuura K, Langer E, Langer G, Untereiner WA, Lucking R, Budel B, Geiser DM, Aptroot A, Diederich P, Schmitt I, Schultz M, Yahr R, Hibbett DS, Lutzoni F, McLaughlin DJ, Spatafora JW, Vilgalys R (2006) Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature 443(7113):818–822. https://doi.org/10.1038/nature05110
Johri P, Marinov GK, Doak TG, Lynch M (2019) Population genetics of Paramecium mitochondrial genomes: recombination, mutation spectrum, and efficacy of selection. Genome Biol Evol 11(5):1398–1416. https://doi.org/10.1093/gbe/evz081
Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20(4):1160–1166. https://doi.org/10.1093/bib/bbx108
Kolesnikova AI, Putintseva YA, Simonov EP, Biriukov VV, Oreshkova NV, Pavlov IN, Sharov VV, Kuzmin DA, Anderson JB, Krutovsky KV (2019) Mobile genetic elements explain size variation in the mitochondrial genomes of four closely-related Armillaria species. BMC Genomics 20(1):351. https://doi.org/10.1186/s12864-019-5732-z
Lamus V, Franco S, Montoya L, Endara AR, Caballero LA, Bandala VM (2015) Mycorrhizal synthesis of the edible mushroom Turbinellus floccosus with Abies religiosa from central Mexico. Mycoscience 56(6):622–626. https://doi.org/10.1016/j.myc.2015.07.001
Lanfear R, Frandsen PB, Wright AM, Senfeld T, Calcott B (2017) PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol Biol Evol 34(3):772–773. https://doi.org/10.1093/molbev/msw260
Lang BF, Gray MW, Burger G (1999) Mitochondrial genome evolution and the origin of eukaryotes. Annu Rev Genet 33:351–397. https://doi.org/10.1146/annurev.genet.33.1.351
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Lavrov DV, Boore JL, Brown WM (2002) Complete mtDNA sequences of two millipedes suggest a new model for mitochondrial gene rearrangements: duplication and nonrandom loss. Mol Biol Evol 19(2):163–169. https://doi.org/10.1093/oxfordjournals.molbev.a004068
Li Q, Chen C, Xiong C, Jin X, Chen Z, Huang W (2018a) Comparative mitogenomics reveals large-scale gene rearrangements in the mitochondrial genome of two Pleurotus species. Appl Microbiol Biotechnol 102(14):6143–6153. https://doi.org/10.1007/s00253-018-9082-6
Li Q, He X, Ren Y, Xiong C, Jin X, Peng L, Huang W (2020a) Comparative mitogenome analysis reveals mitochondrial genome differentiation in ectomycorrhizal and asymbiotic Amanita species. Front Microbiol 11:1382. https://doi.org/10.3389/fmicb.2020.01382
Li Q, Liao M, Yang M, Xiong C, Jin X, Chen Z, Huang W (2018b) Characterization of the mitochondrial genomes of three species in the ectomycorrhizal genus Cantharellus and phylogeny of Agaricomycetes. Int J Biol Macromol 118(Pt A):756–769. https://doi.org/10.1016/j.ijbiomac.2018.06.129
Li Q, Ren Y, Shi X, Peng L, Zhao J, Song Y, Zhao G (2019a) Comparative mitochondrial genome analysis of two ectomycorrhizal fungi (Rhizopogon) reveals dynamic changes of intron and phylogenetic relationships of the subphylum Agaricomycotina. Int J Mol Sci 20(20):5167. https://doi.org/10.3390/ijms20205167
Li Q, Ren Y, Xiang D, Shi X, Zhao J, Peng L, Zhao G (2020b) Comparative mitogenome analysis of two ectomycorrhizal fungi (Paxillus) reveals gene rearrangement, intron dynamics, and phylogeny of basidiomycetes. IMA Fungus 11:12. https://doi.org/10.1186/s43008-020-00038-8
Li Q, Wang Q, Chen C, Jin X, Chen Z, Xiong C, Li P, Zhao J, Huang W (2018c) Characterization and comparative mitogenomic analysis of six newly sequenced mitochondrial genomes from ectomycorrhizal fungi (Russula) and phylogenetic analysis of the Agaricomycetes. Int J Biol Macromol 119:792–802. https://doi.org/10.1016/j.ijbiomac.2018.07.197
Li Q, Wang Q, Jin X, Chen Z, Xiong C, Li P, Liu Q, Huang W (2019b) Characterization and comparative analysis of six complete mitochondrial genomes from ectomycorrhizal fungi of the Lactarius genus and phylogenetic analysis of the Agaricomycetes. Int J Biol Macromol 121:249–260. https://doi.org/10.1016/j.ijbiomac.2018.10.029
Li Q, Wang Q, Jin X, Chen Z, Xiong C, Li P, Zhao J, Huang W (2019c) Characterization and comparison of the mitochondrial genomes from two Lyophyllum fungal species and insights into phylogeny of Agaricomycetes. Int J Biol Macromol 121:364–372. https://doi.org/10.1016/j.ijbiomac.2018.10.037
Li Q, Wang Q, Jin X, Chen Z, Xiong C, Li P, Zhao J, Huang W (2019d) The first complete mitochondrial genome from the family Hygrophoraceae (Hygrophorus russula) by next-generation sequencing and phylogenetic implications. Int J Biol Macromol 122:1313–1320. https://doi.org/10.1016/j.ijbiomac.2018.09.091
Li Q, Xiang D, Wan Y, Wu Q, Wu X, Ma C, Song Y, Zhao G, Huang W (2019e) The complete mitochondrial genomes of five important medicinal Ganoderma species: features, evolution, and phylogeny. Int J Biol Macromol 139:397–408. https://doi.org/10.1016/j.ijbiomac.2019.08.003
Li Q, Yang L, Xiang D, Wan Y, Wu Q, Huang W, Zhao G (2020c) The complete mitochondrial genomes of two model ectomycorrhizal fungi (Laccaria): features, intron dynamics and phylogenetic implications. Int J Biol Macromol 145:974–984. https://doi.org/10.1016/j.ijbiomac.2019.09.188
Li X, Li L, Bao Z, Tu W, He X, Zhang B, Ye L, Wang X, Li Q (2020d) The 287,403 bp mitochondrial genome of ectomycorrhizal fungus Tuber calosporum reveals intron expansion, tRNA loss, and gene rearrangement. Front Microbiol 11:591453. https://doi.org/10.3389/fmicb.2020.591453
Liu W, Cai Y, Zhang Q, Chen L, Shu F, Ma X, Bian Y (2019) The mitochondrial genome of Morchella importuna (272.2 kb) is the largest among fungi and contains numerous introns, mitochondrial non-conserved open reading frames and repetitive sequences. Int J Biol Macromol 143:373–381. https://doi.org/10.1016/j.ijbiomac.2019.12.056
Li Q, Wu P, Li L, Feng H, Tu W, Bao Z, Xiong C, Gui M, Huang W (2021) The first eleven mitochondrial genomes from the ectomycorrhizal fungal genus (Boletus) reveal intron loss and gene rearrangement. Int J Biol Macromol 172: 560–572. https://doi.org/10.1016/j.ijbiomac.2021.01.087
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW--a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res 41(Web Server issue):W575-W581. https://doi.org/10.1093/nar/gkt289
Losada L, Pakala SB, Fedorova ND, Joardar V, Shabalina SA, Hostetler J, Pakala SM, Zafar N, Thomas E, Rodriguez-Carres M, Dean R, Vilgalys R, Nierman WC, Cubeta MA (2014) Mobile elements and mitochondrial genome expansion in the soil fungus and potato pathogen Rhizoctonia solani AG-3. FEMS Microbiol Lett 352(2):165–173. https://doi.org/10.1111/1574-6968.12387
Lowe TM, Chan PP (2016) tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res 44(W1):W54–W57. https://doi.org/10.1093/nar/gkw413
Munoz-Gomez SA, Wideman JG, Roger AJ, Slamovits CH (2017) The origin of mitochondrial cristae from Alphaproteobacteria. Mol Biol Evol 34(4):943–956. https://doi.org/10.1093/molbev/msw298
Pacheco-Cobos L, Rosetti MF, Esquivel AM, Hudson R (2015) Towards a traditional ecological knowledge-based monitoring scheme: a proposal for the case of edible mushrooms. Biodiversity And Conservation 24(5):1253–1269. https://doi.org/10.1007/s10531-014-0856-6
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Hohna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542. https://doi.org/10.1093/sysbio/sys029
Salavirta H, Oksanen I, Kuuskeri J, Makela M, Laine P, Paulin L, Lundell T (2014) Mitochondrial genome of Phlebia radiata is the second largest (156 kbp) among fungi and features signs of genome flexibility and recent recombination events. PLoS One 9(5):e97141. https://doi.org/10.1371/journal.pone.0097141
Schubert M, Lindgreen S, Orlando L (2016) AdapterRemoval v2: rapid adapter trimming, identification, and read merging. BMC Res Notes 9:88. https://doi.org/10.1186/s13104-016-1900-2
Slater GS, Birney E (2005) Automated generation of heuristics for biological sequence comparison. BMC Bioinformatics 6:31. https://doi.org/10.1186/1471-2105-6-31
Stajich JE, Wilke SK, Ahren D, Au CH, Birren BW, Borodovsky M, Burns C, Canback B, Casselton LA, Cheng CK, Deng J, Dietrich FS, Fargo DC, Farman ML, Gathman AC, Goldberg J, Guigo R, Hoegger PJ, Hooker JB, Huggins A, James TY, Kamada T, Kilaru S, Kodira C, Kues U, Kupfer D, Kwan HS, Lomsadze A, Li W, Lilly WW, Ma LJ, Mackey AJ, Manning G, Martin F, Muraguchi H, Natvig DO, Palmerini H, Ramesh MA, Rehmeyer CJ, Roe BA, Shenoy N, Stanke M, Ter-Hovhannisyan V, Tunlid A, Velagapudi R, Vision TJ, Zeng Q, Zolan ME, Pukkila PJ (2010) Insights into evolution of multicellular fungi from the assembled chromosomes of the mushroom Coprinopsis cinerea (Coprinus cinereus). Proc Natl Acad Sci U S A 107(26):11889–11894. https://doi.org/10.1073/pnas.1003391107
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stone CL, Buitrago ML, Boore JL, Frederick RD (2010) Analysis of the complete mitochondrial genome sequences of the soybean rust pathogens Phakopsora pachyrhizi and P. meibomiae. Mycologia 102(4):887–897. https://doi.org/10.3852/09-198
Stothard P (2000) The sequence manipulation suite: JavaScript programs for analyzing and formatting protein and DNA sequences. Biotechniques 28(6):1102, 1104.
Vaidya G, Lohman DL, Meier R (2011) SequenceMatrix: concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 27(2):171–180
Valach M, Burger G, Gray MW, Lang BF (2014) Widespread occurrence of organelle genome-encoded 5S rRNAs including permuted molecules. Nucleic Acids Res 42(22):13764–13777. https://doi.org/10.1093/nar/gku1266
van Creveld SG, Rosenwasser S, Schatz D, Koren I, Vardi A (2015) Early perturbation in mitochondria redox homeostasis in response to environmental stress predicts cell fate in diatoms. ISME J 9(2):385–395. https://doi.org/10.1038/ismej.2014.136
Wang X, Jia LH, Wang MD, Yang H, Chen MY, Li X, Liu HY, Li Q, Liu N (2020a) The complete mitochondrial genome of medicinal fungus Taiwanofungus camphoratus reveals gene rearrangements and intron dynamics of Polyporales. Sci Rep 10(1):16500. https://doi.org/10.1038/S41598-020-73461-X
Wang X, Song A, Wang F, Chen M, Li X, Li Q, Liu N (2020b) The 206 kbp mitochondrial genome of Phanerochaete carnosa reveals dynamics of introns, accumulation of repeat sequences and plasmid-derived genes. Int J Biol Macromol 162:209–219. https://doi.org/10.1016/j.ijbiomac.2020.06.142
Wang X, Wang YJ, Yao W, Shen JW, Chen MY, Gao M, Ren JN, Li Q, Liu N (2020c) The 256 kb mitochondrial genome of Clavaria fumosa is the largest among phylum Basidiomycota and is rich in introns and intronic ORFs. IMA Fungus 11(1):26. https://doi.org/10.1186/S43008-020-00047-7
Wang Y, Xu J (2020) Mitochondrial genome polymorphisms in the human pathogenic fungus Cryptococcus neoformans. Front Microbiol 11:706. https://doi.org/10.3389/fmicb.2020.00706
Wu P, Bao Z, Tu W, Li L, Xiong C, Jin X, Li P, Gui M, Huang W, Li Q (2021) The mitogenomes of two saprophytic Boletales species (Coniophora) reveals intron dynamics and accumulation of plasmid-derived and non-conserved genes. Comput Struct Biotechnol J 19:401–414. https://doi.org/10.1016/j.csbj.2020.12.041
Xia Y, Zheng Y, Murphy RW, Zeng X (2016) Intraspecific rearrangement of mitochondrial genome suggests the prevalence of the tandem duplication-random loss (TDLR) mechanism in Quasipaa boulengeri. BMC Genomics 17(1):965. https://doi.org/10.1186/s12864-016-3309-7
Yadav V, Sun S, Billmyre RB, Thimmappa BC, Shea T, Lintner R, Bakkeren G, Cuomo CA, Heitman J, Sanyal K (2018) RNAi is a critical determinant of centromere evolution in closely related fungi. Proc Natl Acad Sci U S A 115(12):3108–3113. https://doi.org/10.1073/pnas.1713725115
Ye J, Cheng J, Ren Y, Liao W, Li Q (2020) The first mitochondrial genome for Geastrales (Sphaerobolus stellatus) reveals intron dynamics and large-scale gene rearrangements of Basidiomycota. Front Microbiol 11:1970. https://doi.org/10.3389/fmicb.2020.01970
Yoon H, You YH, Woo JR, Park YJ, Kong WS, Lee BM, Kim JG (2012) The mitochondrial genome of the white-rot fungus Flammulina velutipes. J Gen Appl Microbiol 58(4):331–337
Zhong L, Wang M, Li D, Tang S, Zhang T, Bian W, Chen X (2018) Complete mitochondrial genome of Odontobutis haifengensis (Perciformes, Odontobutiae): a unique rearrangement of tRNAs and additional non-coding regions identified in the genus Odontobutis. Genomics 110(6):382–388. https://doi.org/10.1016/j.ygeno.2017.12.008
Zubaer A, Wai A, Hausner G (2018) The mitochondrial genome of Endoconidiophora resinifera is intron rich. Sci Rep 8(1):17591. https://doi.org/10.1038/s41598-018-35926-y
Funding
This study was supported by the start-up funds for talent introduction of the Chengdu University.
Author information
Authors and Affiliations
Contributions
Conceived and designed experiments: Q.L., and J.C.. Performed the experiments: X.W., and Z.L.. Analyzed the data: Q.L., H.P., W.L., and Y.Z.. Wrote and revised the paper: Q.L., and J.C..
Corresponding authors
Ethics declarations
Ethical approval
This research does not contain any studies with human participants performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 2749 kb)
Rights and permissions
About this article
Cite this article
Cheng, J., Luo, Q., Ren, Y. et al. Panorama of intron dynamics and gene rearrangements in the phylum Basidiomycota as revealed by the complete mitochondrial genome of Turbinellus floccosus. Appl Microbiol Biotechnol 105, 2017–2032 (2021). https://doi.org/10.1007/s00253-021-11153-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11153-w