Abstract
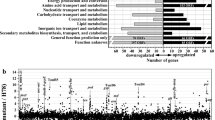
The rhizobacterium Pseudomonas protegens H78 biosynthesizes a number of antibiotic compounds, including pyoluteorin, 2,4-diacetylphloroglucinol, and pyrrolnitrin. Here, we investigated the global regulatory function of the nitrogen metabolism-related sigma factor RpoN in P. protegens H78 through RNA-seq and phenotypic analysis. During the mid- to late-log growth phase, transcriptomic profiling revealed that 562 genes were significantly upregulated, and 502 genes were downregulated by at least twofold at the RNA level in the rpoN deletion mutant in comparison with the wild-type strain H78. With respect to antibiotics, Plt biosynthesis and the expression of its operon were positively regulated, while Prn biosynthesis and the expression of its operon were negatively regulated by RpoN. RpoN is responsible for the global activation of operons involved in flagellar biogenesis and assembly, biofilm formation, and bacterial mobility. In contrast, RpoN was shown to negatively control a number of secretion system operons including one type VI secretion system operon (H1-T6SS), two pilus biogenesis operons (Flp/Tad-T4b pili and Csu-T1 pili), and one polysaccharide biosynthetic operon (psl). In addition, two operons that are involved in mannitol and inositol utilization are under the positive regulation of RpoN. Consistent with this result, the ability of H78 to utilize mannitol or inositol as a sole carbon source is positively influenced by RpoN. Taken together, the RpoN-mediated global regulation is mainly involved in flagellar biogenesis and assembly, bacterial mobility, biofilm formation, antibiotic biosynthesis, secretion systems, and carbon utilization in P. protegens H78.







Similar content being viewed by others
References
Abbas A, Morrissey JP, Marquez PC, Sheehan MM, Delany IR, O'Gara F (2002) Characterization of interactions between the transcriptional repressor PhlF and its binding site at the phlA promoter in Pseudomonas fluorescens F113. J Bacteriol 184(11):3008–3016. https://doi.org/10.1128/JB.184.11.3008-3016.2002
Anderson DK, Ohta N, Wu J, Newton A (1995) Regulation of the Caulobacter crescentus rpoN gene and function of the purified σ54 in flagellar gene transcription. Mol Gen Genet 246(6):697–706. https://doi.org/10.1007/BF00290715
Brunker P, Altenbuchner J, Mattes R (1998) Structure and function of the genes involved in mannitol, arabitol and glucitol utilization from Pseudomonas fluorescens DSM50106. Gene 206(1):117–126. https://doi.org/10.1016/S0378-1119(97)00574-X
Dasgupta N, Wolfgang MC, Goodman AL, Arora SK, Jyot J, Lory S, Ramphal R (2003) A four-tiered transcriptional regulatory circuit controls flagellar biogenesis in Pseudomonas aeruginosa. Mol Microbiol 50(3):809–824. https://doi.org/10.1046/j.1365-2958.2003.03740.x
Dong TG, Mekalanos JJ (2012) Characterization of the RpoN regulon reveals differential regulation of T6SS and new flagellar operons in Vibrio cholerae O37 strain V52. Nucleic Acids Res 40(16):7766–7775. https://doi.org/10.1093/nar/gks567
Dong T, Yu R, Schellhorn H (2011) Antagonistic regulation of motility and transcriptome expression by RpoN and RpoS in Escherichia coli. Mol Microbiol 79(2):375–386. https://doi.org/10.1111/j.1365-2958.2010.07449.x
Fazli M, Rybtke M, Steiner E, Weidel E, Berthelsen J, Groizeleau J, Bin W, Zhi BZ, Yaming Z, Kaever V, Givskov M, Hartmann RW, Eberl L, Tolker-Nielsen T (2017) Regulation of Burkholderia cenocepacia biofilm formation by RpoN and the c-di-GMP effector BerB. MicrobiologyOpen 6(4):e00480. https://doi.org/10.1002/mbo3.480
Giltner CL, Nguyen Y, Burrows LL (2012) Type IV pilin proteins: versatile molecular modules. Microbiol Mol Biol Rev 76(4):740–772. https://doi.org/10.1128/MMBR.00035-12
Haas D, Keel C (2003) Regulation of antibiotic production in root-colonizing Peudomonas spp. and relevance for biological control of plant disease. Annu Rev Phytopathol 41:117–153. https://doi.org/10.1146/annurev.phyto.41.052002.095656
Hao B, Mo ZL, Xiao P, Pan HJ, Lan X, Li GY (2013) Role of alternative sigma factor 54 (RpoN) from Vibrio anguillarum M3 in protease secretion, exopolysaccharide production, biofilm formation, and virulence. Appl Microbiol Biotechnol 97(6):2575–2585. https://doi.org/10.1007/s00253-012-4372-x
Hill DS, Stein JI, Torkewitz NR, Morse AM, Howell CR, Pachlatko JP, Becker JO, Ligon JM (1994) Cloning of genes involved in the synthesis of pyrrolnitrin from Pseudomonas fluorescens and role of pyrrolnitrin synthesis in biological control of plant disease. Appl Environ Microbiol 60(1):78–85
Huang X, Zhu D, Ge Y, Hu H, Zhang X, Xu Y (2004) Identification and characterization of pltZ, a gene involved in the repression of pyoluteorin biosynthesis in Pseudomonas sp. M18. FEMS Microbiol Lett 232(2):197–202. https://doi.org/10.1016/S0378-1097(04)00074-6
Huang X, Wang Z, Liu Y, Zhang X (2017) Complete genome sequence of Pseudomonas protegens H78, a plant growth-promoting rhizobacterium. Genome Announc 5(16):e00233-17. https://doi.org/10.1128/genomeA.00233-17
Hunt TP, Magasanik B (1985) Transcription of glnA by purified Escherichia coli components: core RNA polymerase and the products of glnF, glnG, and glnL. Proc Natl Acad Sci U S A 82(24):8453–8457. https://doi.org/10.1073/pnas.82.24.8453
Ishimoto KS, Lory S (1989) Formation of pilin in Pseudomonas aeruginosa requires the alternative sigma factor (RpoN) of RNA polymerase. Proc Natl Acad Sci U S A 86(6):1954–1957. https://doi.org/10.1073/pnas.86.6.1954
Jones J, Studholme DJ, Knight CG, Preston GM (2007) Integrated bioinformatic and phenotypic analysis of RpoN-dependent traits in the plant growth-promoting bacterium Pseudomonas fluorescens SBW25. Environ Microbiol 9(12):3046–3064. https://doi.org/10.1111/j.1462-2920.2007.01416.x
Kawagishi I, Nakada M, Nishioka N, Homma M (1997) Cloning of a Vibrio alginolyticus rpoN gene that is required for polar flagellar formation. J Bacteriol 179(21):6851–6854. https://doi.org/10.1128/jb.179.21.6851-6854.1997
King EO, Ward MK, Raney DE (1954) Two simple media for the demonstration of pyocyanin and fluorescin. J Lab Clin Med 44(2):301–307
Kroger C, Fuchs TM (2009) Characterization of the myo-inositol utilization island of Salmonella enterica serovar Typhimurium. J Bacteriol 191(2):545–554. https://doi.org/10.1128/JB.01253-08
Leang C, Krushkal J, Ueki T, Puljic M, Sun J, Juarez K, Nunez C, Reguera G, DiDonato R, Postier B, Adkins RM, Lovley DR (2009) Genome-wide analysis of the RpoN regulon in Geobacter sulfurreducens. BMC Genomics 10:331. https://doi.org/10.1186/1471-2164-10-331
Li Y, Jiang H, Xu Y, Zhang X (2008) Optimization of nutrient components for enhanced phenazine-1-carboxylic acid production by gacA-inactivated Pseudomonas sp. M18G using response surface method. Appl Microbiol Biotechnol 77(6):1207–1217. https://doi.org/10.1007/s00253-007-1213-4
Li S, Huang X, Wang G, Xu Y (2012) Transcriptional activation of pyoluteorin operon mediated by the LysR-type regulator PltR bound at a 22 bp lys box in Pseudomonas aeruginosa M18. PLoS One 7(6):e39538. https://doi.org/10.1371/journal.pone.0039538
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621–628. https://doi.org/10.1038/nmeth.1226
O’Toole R, Milton DL, Horstedt P, Wolf-Watz H (1997) RpoN of the fish pathogen Vibrio (Listonella) anguillarum is essential for flagellum production and virulence by the water-borne but not intraperitoneal route of inoculation. Microbiology 143(Pt 12):3849–3859. https://doi.org/10.1099/00221287-143-12-3849
Pallen M (1999) RpoN-dependent transcription of rpoH? Mol Microbiol 31(1):393–395. https://doi.org/10.1046/j.1365-2958.1999.01148.x
Pechy-Tarr M, Bottiglieri M, Mathys S, Lejbolle KB, Schnider-Keel U, Maurhofer M, Keel C (2005) RpoN (σ54) controls production of antifungal compounds and biocontrol activity in Pseudomonas fluorescens CHA0. Mol Plant-Microbe Interact 18(3):260–272. https://doi.org/10.1094/MPMI-18-0260
Porrua O, Garcia-Gonzalez V, Santero E, Shingler V, Govantes F (2009) Activation and repression of a σN-dependent promoter naturally lacking upstream activation sequences. Mol Microbiol 73(3):419–433. https://doi.org/10.1111/j.1365-2958.2009.06779.x
Potvin E, Sanschagrin F, Levesque RC (2008) Sigma factors in Pseudomonas aeruginosa. FEMS Microbiol Rev 32(1):38–55. https://doi.org/10.1111/j.1574-6976.2007.00092.x
Saldias MS, Lamothe J, Wu R, Valvano MA (2008) Burkholderia cenocepacia requires the RpoN sigma factor for biofilm formation and intracellular trafficking within macrophages. Infect Immun 76(3):1059–1067. https://doi.org/10.1128/IAI.01167-07
Sambrook J, Russell DW (2001) Molecular cloning : a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sana TG, Soscia C, Tonglet CM, Garvis S, Bleves S (2013) Divergent control of two type VI secretion systems by RpoN in Pseudomonas aeruginosa. PLoS One 8(10):e76030. https://doi.org/10.1371/journal.pone.0076030
Sarniguet A, Kraus J, Henkels MD, Muehlchen AM, Loper JE (1995) The sigma factor σS affects antibiotic production and biological control activity of Pseudomonas fluorescens Pf-5. Proc Natl Acad Sci U S A 92(26):12255–12259. https://doi.org/10.1073/pnas.92.26.12255
Schnider U, Keel C, Blumer C, Troxler J, Defago G, Haas D (1995) Amplification of the housekeeping sigma factor in Pseudomonas fluorescens CHA0 enhances antibiotic production and improves biocontrol abilities. J Bacteriol 177(18):5387–5392. https://doi.org/10.1128/jb.177.18.5387-5392.1995
Smith AH, Blevins JS, Bachlani GN, Yang XF, Norgard MV (2007) Evidence that RpoS (σS) in Borrelia burgdorferi is controlled directly by RpoN (σ54/σN). J Bacteriol 189(5):2139–2144. https://doi.org/10.1128/JB.01653-06
Solovyev V, Salamov A (2011) Automatic annotation of microbial genomes and metagenomic sequences. In: Li RW (ed) Metagenomics and its applications in agriculture, biomedicine and environmental studies. Nova Science Publishers, New York, pp 61–78
Taylor M, Butler R, Chambers S, Casimiro M, Badii F, Merrick M (1996) The RpoN-box motif of the RNA polymerase sigma factor σN plays a role in promoter recognition. Mol Microbiol 22(5):1045–1054
Tomaras AP, Dorsey CW, Edelmann RE, Actis LA (2003) Attachment to and biofilm formation on abiotic surfaces by Acinetobacter baumannii: involvement of a novel chaperone-usher pili assembly system. Microbiology 149(Pt 12):3473–3484. https://doi.org/10.1099/mic.0.26541-0
Totten PA, Lara JC, Lory S (1990) The rpoN gene product of Pseudomonas aeruginosa is required for expression of diverse genes, including the flagellin gene. J Bacteriol 172(1):389–396. https://doi.org/10.1128/jb.172.1.389-396.1990
Wang Z, Huang X, Liu Y, Yang G, Liu Y, Zhang X (2017) GacS/GacA activates pyoluteorin biosynthesis through Gac/Rsm-RsmE cascade and RsmA/RsmE-driven feedback loop in Pseudomonas protegens H78. Mol Microbiol 105(6):968–985. https://doi.org/10.1111/mmi.13749
Wei X, Huang X, Tang L, Wu D, Xu Y (2013) Global control of GacA in secondary metabolism, primary metabolism, secretion systems, and motility in the rhizobacterium Pseudomonas aeruginosa M18. J Bacteriol 195(15):3387–3400. https://doi.org/10.1128/JB.00214-13
Yu D, Xia M, Zhang L, Song Y, Duan Y, Yuan T, Yao M, Wu L, Tian C, Wu Z, Li X, Zhou J, Qiu D (2017) RpoN (σ54) is required for floc formation but not for extracellular polysaccharide biosynthesis in a floc-forming Aquincola tertiaricarbonis strain. Appl Environ Microbiol 83(14):e00709–e00717. https://doi.org/10.1128/AEM.00709-17
Zhou Q, Su J, Jiang H, Huang X, Xu Y (2010) Optimization of phenazine-1-carboxylic acid production by a gacA/qscR-inactivated Pseudomonas sp. M18GQ harboring pME6032Phz using response surface methodology. Appl Microbiol Biotechnol 86(6):1761–1773. https://doi.org/10.1007/s00253-010-2464-z
Funding
This study was supported by the National Natural Science Foundation of China (31470196, 31270083).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants by any of the authors.
Electronic supplementary material
ESM 1
(PDF 2918 kb)
Rights and permissions
About this article
Cite this article
Liu, Y., Shi, H., Wang, Z. et al. Pleiotropic control of antibiotic biosynthesis, flagellar operon expression, biofilm formation, and carbon source utilization by RpoN in Pseudomonas protegens H78. Appl Microbiol Biotechnol 102, 9719–9730 (2018). https://doi.org/10.1007/s00253-018-9282-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9282-0



