Abstract
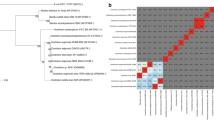
In typical acetone–butanol–ethanol (ABE) fermentation, acetone is the main by-product (50 % of butanol mass) of butanol production, resulting in a low yield of butanol. It is known that some Clostridium tetanomorphum strains are able to produce butanol without acetone in nature. Here, we described that C. tetanomorphum strain DSM665 can produce 4.16 g/L butanol and 4.98 g/L ethanol at pH 6.0, and 9.81 g/L butanol and 1.01 g/L ethanol when adding 1 mM methyl viologen. Butyrate and acetate could be reassimilated and no acetone was produced. Further analysis indicated that the activity of the acetate/butyrate:acetoacetyl-CoA transferase responsible for acetone production is lost in C. tetanomorphum DSM665. The genome of C. tetanomorphum DSM665 was sequenced and deposited in DDBJ, EMBL, and GenBank under the accession no. APJS00000000. Sequence analysis indicated that there are no typical genes (ctfA/B and adc) that are typically parts of an acetone synthesis pathway in C. tetanomorphum DSM665. This work provides new insights in the mechanism of clostridial butanol production and should prove useful for the design of a high-butanol-producing strain.



Similar content being viewed by others
References
Bramono SE, Lam YS, Ong SL, He J (2011) A mesophilic Clostridium species that produces butanol from monosaccharides and hydrogen from polysaccharides. Bioresour Technol 102(20):9558–9563
Carlos DC, Carmen F-S, Antonia R, Marta T, Daniel R, José LG (2013) Genome sequence of the butanol hyperproducer Clostridium saccharoperbutylacetonicum N1-4. Genome Announc 1(2):e00070–13
Clark SW, Bennett GN, Rudolph FB (1989) Isolation and characterization of mutants of Clostridium acetobutylicum ATCC 824 deficient in acetoacetyl-coenzyme A: acetate/butyrate: coenzyme A-transferase (EC 2.8.3.9) and in other solvent pathway enzymes. Appl Environ Microbiol 55(4):970–976
Cooksley CM, Zhang Y, Wang H, Redl S, Winzer K, Minton NP (2012) Targeted mutagenesis of the Clostridium acetobutylicum Acetone-Butanol-Ethanol fermentation pathway. Metab Eng 14(6):630–641
Dürre P (2007) Biobutanol: an attractive biofuel. Biotechnol J 2(12):1525–1534
Desai RP, Papoutsakis ET (1999) Antisense RNA strategies for metabolic engineering of Clostridium acetobutylicum. Appl Environ Microbiol 65(3):936–945
Gottwald M, Hippe H, Gottschalk G (1984) Formation of n-Butanol from D-Glucose by Strains of the “Clostridium tetanomorphum” Group. Appl Environ Microb 48(3):573–576
Jang Y-S, Lee JY, Lee J, Park JH, Im JA, Eom M-H, Lee J, Lee S-H, Song H, Cho J-H (2012) Enhanced butanol production obtained by reinforcing the direct butanol-forming route in Clostridium acetobutylicum. MBio 3(5):e00314–12
Jiang Y, Xu C, Dong F, Yang Y, Jiang W, Yang S (2009) Disruption of the acetoacetate decarboxylase gene in solvent-producing Clostridium acetobutylicum increases the butanol ratio. Metab Eng 11(4):284–291
Jones DT, Woods DR (1986) Acetone-butanol fermentation revisited. Microbiol Rev 50(4):484
Lee SY, Park JH, Jang SH, Nielsen LK, Kim J, Jung KS (2008) Fermentative butanol production by clostridia. Biotechnol Bioeng 101(2):209–228
Lehmann D, Honicke D, Ehrenreich A, Schmidt M, Weuster-Botz D, Bahl H, Lutke-Eversloh T (2012) Modifying the product pattern of Clostridium acetobutylicum: physiological effects of disrupting the acetate and acetone formation pathways. Appl Microbiol Biotechnol 94(3):743–754
Milne CB, Eddy JA, Raju R, Ardekani S, Kim P-J, Senger RS, Jin Y-S, Blaschek HP, Price ND (2011) Metabolic network reconstruction and genome-scale model of butanol-producing strain Clostridium beijerinckii NCIMB 8052. BMC Syst Biol 5(1):130
Nölling J, Breton G, Omelchenko MV, Makarova KS, Zeng Q, Gibson R, Lee HM, Dubois J, Qiu D, Hitti J (2001) Genome sequence and comparative analysis of the solvent-producing bacterium Clostridium acetobutylicum. J Bacteriol 183(16):4823–4838
Poehlein A, Hartwich K, Krabben P, Ehrenreich A, Liebl W, Dürre P, Gottschalk G, Daniel R (2013) Complete genome sequence of the solvent producer Clostridium saccharobutylicum NCP262 (DSM 13864). Genome Announc 1(6):e00997–13
Pruitt KD, Tatusova T, Klimke W, Maglott DR (2009) NCBI Reference Sequences: current status, policy and new initiatives. Nucleic Acids Res 37(Database issue):D32–D36
Qureshi N, Ezeji TC (2008) Butanol, ‘a superior biofuel’production from agricultural residues (renewable biomass): recent progress in technology. Biofuel Bioprod Bior 2(4):319–330
Tummala SB, Welker NE, Papoutsakis ET (2003) Design of antisense RNA constructs for downregulation of the acetone formation pathway of Clostridium acetobutylicum. J Bacteriol 185(6):1923–1934
Wiesenborn D, Rudolph F, Papoutsakis E (1988) Thiolase from Clostridium acetobutylicum ATCC 824 and its role in the synthesis of acids and solvents. Appl Environ Microbiol 54(11):2717–2722
Yarlagadda VN, Gupta A, Dodge CJ, Francis AJ (2012) Effect of exogenous electron shuttles on growth and fermentative metabolism in Clostridium sp. BC1. Bioresour Technol 108:295–299
Zerbino DR, Birney E (2008) Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18(5):821–829
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This study was funded by the National Natural Science Foundation of China (grant number 31270107), the Knowledge Innovation Program of the Chinese Academy of Sciences (grant number KSCX2-EW-J-6), and the National High Technology Research and Development Program of China (grant number 2011AA02A208).
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Fuyu Gong and Guanhui Bao contributed equally to this work.
Rights and permissions
About this article
Cite this article
Gong, F., Bao, G., Zhao, C. et al. Fermentation and genomic analysis of acetone-uncoupled butanol production by Clostridium tetanomorphum . Appl Microbiol Biotechnol 100, 1523–1529 (2016). https://doi.org/10.1007/s00253-015-7121-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7121-0