Abstract
Our knowledge of the lagomorph immune system remains largely based upon studies of the European rabbit (Oryctolagus cuniculus), a major model for studies of immunology. Two important and devastating viral diseases, rabbit hemorrhagic disease and myxomatosis, are affecting European rabbit populations. In this context, we discuss the genetic diversity of the European rabbit immune system and extend to available information about other lagomorphs. Regarding innate immunity, we review the most recent advances in identifying interleukins, chemokines and chemokine receptors, Toll-like receptors, antiviral proteins (RIG-I and Trim5), and the genes encoding fucosyltransferases that are utilized by rabbit hemorrhagic disease virus as a portal for invading host respiratory and gut epithelial cells. Evolutionary studies showed that several genes of innate immunity are evolving by strong natural selection. Studies of the leporid CCR5 gene revealed a very dramatic change unique in mammals at the second extracellular loop of CCR5 resulting from a gene conversion event with the paralogous CCR2. For the adaptive immune system, we review genetic diversity at the loci encoding antibody variable and constant regions, the major histocompatibility complex (RLA) and T cells. Studies of IGHV and IGKC genes expressed in leporids are two of the few examples of trans-species polymorphism observed outside of the major histocompatibility complex. In addition, we review some endogenous viruses of lagomorph genomes, the importance of the European rabbit as a model for human disease studies, and the anticipated role of next-generation sequencing in extending knowledge of lagomorph immune systems and their evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction and historical perspective
During the twentieth century, the European rabbit (Oryctolagus cuniculus) was widely used in laboratories studying immunology and infectious diseases. Serologically detectable allelic forms of rabbit immunoglobulins (Igs) were discovered when isoantibodies developed after immunizations of individual rabbits with immunoglobulins from another rabbit in the form of antigen–antibody complexes, antibodies bound to bacteria, or purified immunoglobulins with Freund’s adjuvant (Oudin 1956, 1960; Dubiski et al. 1959; Dray and Young 1959). By 1962, a nomenclature for rabbit Ig allotypes had been agreed upon (Dray et al. 1962). A series of reviews in a volume “The Rabbit in Contemporary Immunological Research” that was edited by S. Dubiski in 1987 provides an excellent overview of research progress up to the time when a transition to recombinant DNA technology had begun to reveal a more detailed view of immunogenetics of rabbits. In addition to Ig allotypes, rabbit blood groups (Cohen 1987), genetic polymorphisms and deficiencies in components of the complement system (Komatsu 1987), T cell receptors (“T cells” section), and major histocompatibility antigens (RLA) (“MHC or RLA” section) were being characterized. Anti-allotype antisera had already contributed to understanding of B cell development and detection of Igs in pre-B cells, on B lymphocyte surfaces (B cell receptors), and revealed that individual B cells expressed only one of two alleles (allelic exclusion) (reviewed in Cedar and Bergman 2008). In addition, demonstration that genetic recombination occurred between IGHV and IGHC genes was an early indicator that Igs were encoded by more than one genetic unit (Mage et al. 1971). It is unfortunate that the Jackson Laboratory discontinued breeding colonies of several different inbred rabbit strains. Although some were distributed, they are no longer available with the exception of the EIII/JC inbred strain of New Zealand White rabbits (Peng et al. 2015). This strain is maintained by the laboratory of ND Christensen along with a transgenic line carrying human HLA-A2.1 (Hu et al. 2006) used for studies of papilloma virus infection and vaccine development (see “Infectious diseases” section). Similarly, allotype-defined rabbit colonies maintained by Andrew Kelus in Basel and by Rose Mage at NIAID, NIH were not maintained after each retired.
In the twenty-first century, rabbits are still a major source of highly specific polyclonal and monoclonal antibodies of high affinity. They are key models used for studies of human infectious, autoimmune, and cardiovascular diseases. Studies of IGHV (Su and Nei 1999; Esteves et al. 2005; Pinheiro et al. 2013a, 2014a) and IGKC genes (Landucci Tosi et al. 1976; Landucci Tosi et al. l976b; Bouton and van der Loo 1997) expressed in leporids have already provided two of the few examples of trans-species polymorphism observed outside of the major histocompatibility complex. Immunogenetic studies have progressed to include analyses of draft genome assemblies of O. cuniculus and one of ∼30 species of pika (Ochotona princeps). Although these are the only two whole genomes currently available, additional genomic sequencing of members of the entire order Lagomorpha is now planned or already in progress (“Autoimmune disease models” section)
Lagomorph systematics
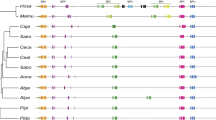
The order Lagomorpha was proposed by Gridley (1912) to distinguish the lagomorphs from the rodents. After some controversy, it is now accepted that the lagomorphs are phylogenetically more related to rodents and classified within the superorder Glires, considered a sister group of the superorder Euarchonta that includes primates (Horner et al. 2007; Murphy et al. 2001). The order Lagomorpha is divided into two families: Ochotonidae and Leporidae. The divergence time between these two families is not agreed upon. According to fossil data, the leporids separated from ochotonids during the Oligocene or Upper Eocene [30–40 million years ago (Mya)] (Dawson 1981). A study of 19 nuclear and three mitochondrial genes from 42 representative species of all extant orders of placental mammals estimated that the Ochotona diverged from the rabbit 50–55 Mya (Springer et al. 2003). The family Ochotonidae is restricted to the genus Ochotona (pikas) that includes 25 extant species divided in four subgenera: Pika, Logotona, Conothoa, and Ochotona (Lanier and Olson 2009; Lissovsky 2014; Melo-Ferreira et al. 2015). Using different mutation rates, the divergence time of these four subgenera was estimated to be 4–7 Mya (Melo-Ferreira et al. 2015). The family Leporidae can be divided into two groups: hares and rabbits. The hare group encompasses a single genus, Lepus, comprising 29 species. However, a recurrent introgression of mitochondrial DNA among the hare species (Melo-Ferreira et al. 2012) may have some implications in the species status. The rabbit group includes ten genera (Brachylagus, Bunolagus, Caprolagus, Nesolagus, Oryctolagus, Pentalagus, Poelagus, Pronolagus, Romerolagus, and Sylvilagus) and 25 species. According to fossil data, the family Leporidae developed and differentiated in North America and Asia around 20 Mya (Dawson 1981; Lopez-Martinez 1989) with a posterior radiation. Mathee et al. (2004) using molecular data suggested that the great Leporid radiation happened 14 Mya. The three best-studied Leporid genera are the European rabbit (Oryctolagus), the Jackrabbit (Lepus), and the Cottontail (Sylvilagus) that are estimated to have diverged 12 Mya (Matthee et al. 2004) (Fig. 1). The genus Oryctolagus is monospecific (O. cuniculus), while for genus Sylvilagus, more than 16 species are described.
The genus Ochotona is present in Northern Asia and West North America, while Oryctolagus is present in Continental Europe, North Africa (Morocco and Algeria), England, Australia, New Zealand, South America, and islands over the world. The genus Lepus is distributed in Holarctic, Indomalayan, and Afro-tropical regions (except tropical forests), and the genus Sylvilagus is restricted to the American continent (Corbet 1994).
European rabbit natural history
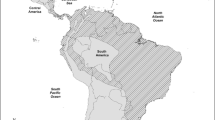
The European rabbit (O. cuniculus) has been used as a laboratory animal model in immunology (see “Introduction and historical perspective” section). This species originated in the Iberian Peninsula where two morphologically differentiated subspecies coexist: O ryctolagus cuniculus algirus and Oryctolagus cuniculus cuniculus (Cabrera 1914, Lopez-Martinez 1989; Corbet 1994). Studies of the mtDNA and X and Y chromosomal and nuclear markers indicate that these two subspecies diverged in the Pleistocene (about 2 Mya) (Biju-Duval et al. 1991; Monnerot et al. 1994; Branco et al. 2000, 2002; Carneiro et al. 2011; Geraldes et al. 2005; reviewed in Ferrand and Branco 2007). The subspecies O. c. algirus is present in wild populations of the Southwestern Iberian Peninsula, in the Portuguese archipelago islands Azores and Madeira, and in the Spanish Canary Islands, while O. c. cuniculus is present in the northeastern of Iberian Peninsula, in all domestic breeds, and in all wild populations of Continental Europe (north of Pyrenees), Great Britain, and overseas. In the Iberian Peninsula, a contact zone exists where the two subspecies can hybridize, suggesting a post-glacial expansion from the southwestern refuge to the north and from the northeastern refuge to the south or west (Branco et al. 2000, 2002; Carneiro et al. 2011; Geraldes et al. 2005; reviewed in Ferrand and Branco 2007) (Fig. 2). Characterization of French and other non-Iberian European rabbit populations showed a significant loss of genetic diversity when compared with the Iberian populations evidencing a strong bottleneck effect which suggests that the Pyrenees were an important natural barrier for the rabbit dispersion (Abrantes et al. 2013a; Alves et al. 2015; Branco et al. 2000, 2002; Carneiro et al. 2011, 2014; Esteves et al. 2004; Geraldes et al. 2005; Pinheiro et al. 2015; Surridge et al. 2008; van der Loo et al. 1999a). Rabbit domestication probably happened in France in one single event as population genetic studies in domestic breeds showed these breeds represent a subset of the genetic variability found in the French O. c. cuniculus wild populations (Alves et al. 2015; Carneiro et al. 2011, 2014; reviewed in Ferrand and Branco 2007; Geraldes et al. 2005; Queney et al. 2001).
Geographic distribution of European rabbit (Oryctolagus cuniculus). The subspecies O. c. algirus occupies the southwestern part of Iberia. European rabbit of the northwestern areas of Iberia and of the rest of Europe belong to the subspecies O. c. cuniculus. The contact zone between the two subspecies is indicated in dark gray
The European rabbit was introduced by man in more than 800 islands all over the world (Flux and Fullagar 1992). The reasons for this success are particular features such as high growth and reproduction rate and efficient food utilization (associated with coprophagy and the ability to shift between growth strategies: r selection—early age of maturity, large number of young, a large reproductive effort or K selection—delayed reproduction; small number of young; and a smaller reproductive effort according to the carrying capacity of the environment) (Parry 1981). Other aspects are also fundamental in its success in islands, such as the absence of competitors, few predators, and nonexistence of diseases. Some of these introductions constitute natural laboratories showing the consequences of an extreme bottleneck event on a population, which can be very useful for conservation-genetic studies. Indeed, the study of founder effects has become increasingly important in population genetics, speciation theory, and conservation biology. Founder effects are particularly important because they can increase the demographic stochasticity, the rate of inbreeding, and the fixation of deleterious alleles, therefore reducing adaptive potential and increasing the probability of population extinction (e.g., Lande 1994; O'Brien et al. 1994; Newman and Pilson 1997; Saccheri et al. 1998).
The status of the European rabbit presents a “conservation paradox” (Lees and Bell 2008). On the one hand, in its natural Mediterranean ecosystem, the European rabbit is a key species whose natural populations have suffered great declines. On the other hand, in places like Australia and New Zealand where the European rabbit was introduced, its populations have grown to an extent that it is considered a plague.
Viral diseases
Most of the host genes encoding cellular viral restriction factors are engaged in evolutionary arm race dynamics as a result of the genetic conflict between the host and the virus. This back-and-forth process between the interacting virus and host, leading to the rapid evolution of both, is an example of the “Red Queen hypothesis,” the evolutionary law proposed by Van Valen (1973). In a co-evolutionary process, the host restriction factor exerts selective pressure on the viral antagonist, creating a disadvantageous condition for the virus. In response, new mutations that allow the virus to evade host restriction factor action are selected in the pathogen. The European rabbit is a perfect model to study these co-evolutionary processes since two very important viral diseases, rabbit hemorrhagic disease and myxomatosis, have been dramatically affecting both the wild European rabbit populations and domestic breeds.
Rabbit hemorrhagic disease virus (RHDV)
RHDV is responsible for the rabbit hemorrhagic disease (RHD) that affects both subspecies of European rabbit (O. c. cuniculus and O. c. algirus) (Abrantes et al. 2012a; Alda et al. 2010; Muller et al. 2009). The disease was first reported in the early 1980s in China following the importation of Angora rabbits from Germany and quickly spread worldwide (Abrantes et al. 2012b). RHD causes an acute fulminant hepatitis, and high mortalities are observed in wild and domestic rabbits accounting for a strong ecological and economic burden (Delibes-Mateos et al. 2008; 2014; Marchandeau et al. 1998, 2000; Mitro and Krauss 1993; Villafuerte et al. 1995). RHDV belongs to the genus Lagovirus, family Caliciviridae, and has a single-stranded positive-sense RNA genome of ∼7.5 kb that encodes structural and non-structural proteins. Non-structural proteins include the replicative machinery, while the capsid protein constitutes the major structural protein.
Emergence of RHDV as a pathogenic virus for the European rabbit still remains controversial. The most widely accepted hypotheses propose the emergence either from a non-pathogenic form of the virus or through a species jump (Kerr et al. 2009; Esteves et al. 2015), but as yet, there has not been entirely conclusive evidence to support these hypotheses. In addition, the recent emergence of a genetically distinct variant of RHDV in France in 2010 (Le Gall-Recule et al. 2011) that also affects both subspecies of European rabbit (Abrantes et al. 2013b; Almeida et al. 2015; Dalton et al. 2014, 2015; Lopes et al. 2014) showed the importance of non-pathogenic strains in RHDV evolution (Lopes et al. 2015) but also provided evidence for a role of other leporid species in RHDV epidemiology (Camarda et al. 2014; Puggioni et al. 2013).
Host resistance mechanisms to RHDV are also poorly documented due to the lack of a viable cell culture system. Nonetheless, indirect systems were able to demonstrate the ability of the RHDV pathogenic strains to attach to histo-blood group antigens (HGBA) present on the surface of host respiratory and gut epithelial cells that act as co-factors for the virus to start the infectious process (Ruvoen-Clouet et al. 1995; Ruvoen-Clouet et al. 2000). Animals with low expression of histo-blood group antigen (HBGA) were shown to be less susceptible to infection by older RHDV strains (Nyström et al. 2011). In addition, in rabbits under 2 months of age, the absence of such structures might account for their natural resistance to RHDV (Ruvoen-Clouet et al. 2000). However, as hepatocytes do not to express this antigen and infection in young rabbits is accompanied by hepatic lesions due to virus replication (Mikami et al. 1999), another hepatic cellular receptor must exist. Experimental inoculation trials also showed the importance of the immune responses in the course of RHDV infection. Indeed, young rabbits present an early and strong inflammatory response that is accompanied by a systemic B cell response that precedes a specific antiviral antibody response (Ferreira et al. 2008; Marques et al. 2012). In contrast, in adult rabbits, an early depletion of the B and T leukocyte populations is observed that irremediably compromises the immune response (Ferreira et al. 2006; Marques et al. 2012).
Myxomatosis
Wild and domestic populations of European rabbit (O. cuniculus) have been suffering an alarming decline in the last decades. One of the greatest forces behind such decrease has been the highly lethal disease myxomatosis, caused by the rabbit-specific myxoma virus (MYXV) (Aragão 1927; Ratcliffe et al. 1952). The course of infection, pathogenesis, and classical diagnostic symptoms for different MYXV strains are described and reviewed elsewhere (Best et al. 2000; Best and Kerr 2000; Kerr and McFadden 2002; Zuniga 2002); nevertheless, infection with highly virulent MYXV strains culminate with death, normally between days 9 and 12 post-infection. On the other hand, MYXV is naturally found in two American leporid species, the tapeti (Sylvilagus brasiliensis) and the brush rabbit (Sylvilagus bachmani), in which it causes an innocuous localized cutaneous fibroma (Fenner and Ratcliffe 1965; Marshall and Regnery 1960a, b).
MYXV, a poxvirus of the genus Leporipoxvirus, is a classic example of a pathogen adapted to its native host (Sylvilagus) that can infect a new susceptible host (Oryctolagus) yet adapt to some populations of the new species. This scenario was observed in Australia and Europe, where certain European rabbit populations became resistant to MYXV infection in a co-evolutionary process between host and pathogen (Kerr 2012; Kerr et al. 2015). However, the genetic basis of resistance in these wild European rabbit populations is still unknown. The origin of differential pathogenicity of MYXV infection in native Sylvilagus hosts and in the susceptible Oryctolagus host is also uncertain.
The complete genomic DNA sequencing of the MYXV Lausanne (Lu) strain, the most studied and most often used strain both in in vivo and in vitro studies, revealed a 161.8-kb genome with a total of 171 open reading frames (ORFs) (Cameron et al. 1999). Besides the genes associated with the viral replicative machinery and structure, MYXV genome contains genes encoding host-interactive immunomodulatory and host range proteins, primarily located within the 15–25 kb at both ends of the genome (Cameron et al. 1999). The function of these modulators has been assessed in vivo and/or in vitro, by infecting laboratory European rabbits and specific cell lines, respectively, with mutant viruses. The role of 26 of these modulators is described elsewhere (Kerr and McFadden 2002; Kerr 2012; Liu et al. 2010; Spiesschaert et al. 2011; Stanford et al. 2007); however, these proteins can be generally classified as viroreceptors, virokines, anti-apoptotic factors, immune modulators, and host range factors.
Innate immune system
Cytokines are a group of small proteins with critical roles in the immune response that include interleukins, chemokines, and other signaling proteins named according to the cells that produce them or their main activities.
Interleukins
Interleukins (ILs) are secreted by leukocytes and possess the ability to act on other cells. The ILs are involved in multiple biological activities which may explain why they have sites evolving under positive selection (Neves et al. 2014b). During the last years, these proteins have been characterized in European rabbit. Perkins et al. (2000) reported the cDNA sequences of European rabbit IL2, IL4, IL6, and IL10 identifying some truncated forms for IL2, IL4, and IL10. The genomic region of European rabbit chromosome 3 encoding IL5, IL4, and IL13 was sequenced and assembled as part of the whole genome OryCun 2.0 and independently from a second sequence of a normal NZW rabbit by the NIH Sequencing Center (NISC) as part of the ENCODE project to compare coding and noncoding regions in mammalian species with corresponding human sequences (Margulies et al. 2007). Rabbit ENm002 (Accession: PRJNA13686 ID: 13686) from the 1 % of rabbit genome sequenced by ENCODE was assembled from sequences of overlapping BAC clones. Gertz et al. (2013) published a detailed comparison of the genomic sequences from the two assemblies with those of nine other mammalian species including promoter regions, DNAse hypersensitive sites, and transcription factor binding sites. A resequencing study of DNA from the OryCun 2.0 DNA donor and two other rabbits (Mage and Mage 2012) did not confirm the IL4 genomic sequence in OryCun 2.0 where there appeared to be a frameshift in exon 2 of IL4 that could have led to exon skipping and expression of a described alternative product IL4δ2 (Perkins et al. 2000).
IL6 is a particularly relevant cytokine for rabbit because of its involvement in the immune response against RHDV (Marques et al. 2012). A mutation in the usual stop codon in the European rabbit IL6 results in an additional 27 amino acids, which are absent in the Cottontail rabbit and Jackrabbit (Perkins et al. 2000). Subsequent studies of IL6 in other genera (riverine rabbit, Amami rabbit, pygmy rabbit, volcano rabbit, and American pika) showed that the Amami rabbit has also an additional extension of 17 amino acids due to a deletion in the stop codon. Two independent events may have occurred: one between 2 and 8 Mya in the ancestor of the European rabbit and the other in the Amami rabbit ancestor at a maximum of 9 Mya. The absence of this IL6 extension in riverine rabbit, sister genus of European rabbit, suggests this evolutionary event happened by convergence and could play some functional role (Neves et al. 2014a).
IL7 is expressed in the European rabbit bone marrow and, in vitro, it is required for differentiation of lymphoid progenitors to B and T lineage cells. It was reported that a novel European rabbit IL7 isoform (IL7II) is generated by alternative splicing, with an 11 amino acid insertion encoded by a separate exon, exon 2b, which is conserved in other lagomorphs including American pika, cottontail, and jackrabbit. Analysis of sequence downstream and upstream of IL7 exon 2b revealed high conservation between European rabbit, cottontail, jackrabbit, and American pika, supporting the possibility that IL7 exon 2b may be functional in all Lagomorpha (Siewe et al. 2010). More recently, a study characterized genetically the ILs implicated in the immune response against inflammatory processes, rabbit hemorrhagic disease virus and myxoma virus (IL1α, IL1β, IL2, IL4, IL8, IL10, IL12A, IL12B, IL15, and IL18) in the European rabbit, pygmy rabbit, cottontail rabbit, European brown hare, and American pika. Among lagomorphs, the interleukins are generally well-conserved, the major differences occurring between leporids and the American pika. This study also describes another transcript for American pika IL8 with an insertion of four amino acids in the 5′ region of exon 2. One of the hypotheses that this study highlights is that the European rabbit could be a more appropriate animal model than mouse for human innate immunity studies (Neves et al. 2015a). Indeed, for the ILs studied, the nucleotide and amino acid genetic distances between human and European rabbit sequences were lower than those obtained between human and mouse or rat.
Chemokines and chemokine receptors
Chemokines and their receptors play crucial roles in immune and inflammatory responses (Charo and Ransohoff 2006). Some genetic variants of these cytokines have been characterized in lagomorphs. The CXC motif chemokine receptor 4 (CXCR4) is highly conserved among five genera (Ochotona, Lepus, Sylvilagus, Bunolagus, and Oryctolagus) (Abrantes et al. 2008). Studies of the leporid C–C motif chemokine receptor 5 (CCR5) genes revealed the first striking difference between genera. European rabbit shares with riverine rabbit (Bunolagus monticularis) and Amami rabbit (Pentalagus furnessi) a very dramatic change at the second extracellular loop of CCR5 resulting from a gene conversion event with the paralogous CCR2, where the CCR5 sequence motif QTLKMT was replaced by the CCR2 motif HTIMRN (Abrantes et al. 2011). Such a remarkable alteration was not observed in the eastern cottontail (Sylvilagus floridanus) or in European and Iberian hares (Lepus europeaus and Lepus granatensis) (Carmo et al. 2006). The most likely evolutionary scenario that could explain this pattern is that the recombination event happened in the ancestor of Oryctolagus, Bunolagus, and Pentalagus genera at ∼8 Mya, probably conferring some selective advantage and thus became fixed in the ancestral population (Fig. 3).
With these findings, it became imperative to study the genetic aspects of CCR5 ligands in leporid genera. The first assessed were the C–C motif chemokines CCL1 to CCL14, which includes the genes of macrophage inflammatory proteins (MIP) and the monocyte chemotactic proteins (MCP) and RANTES. Of these 14 genes, all were found to be functional in European rabbit with the notable exception of the CCR5 disease-related MPC2/CCL8. By extending this study to other leporids, CCL8/MP2 was found also pseudogenized in the African riverine rabbit (B. monticularis), while functional and transcribed in Lepus and Sylvilagus species (van der Loo et al. 2012). Inter-species comparisons of CCL3, CCL4, and CCL5 between the genera Oryctolagus, Sylvilagus, and Lepus revealed evidence of strong purifying selection at the sites of receptor/CCR binding (de Matos et al. 2014). Studies of CCL14 revealed that this ligand was functional in each of five leporid genera tested (including Pentalagus, Brachylagus, and Romerolagus) but underwent a pseudogenization process in Ochotonidae family (Neves et al. 2015b). CCL16 was described as being a pseudogene in the European rabbit (Shibata et al. 2013). The functional outcomes of such differences between species remain to be evaluated.
Toll-like receptors
Toll-like receptors (TLRs) are important components of the innate immune system as they are a class of pattern recognition receptors (PRRs) that sense conserved molecular structures of pathogens, the pathogen-associated molecular patterns (PAMPs), and elicit an immune response (Medzhitov 2001). TLRs are type I transmembrane glycoproteins with an extracellular leucine-rich repeat (LRR) domain involved in ligand binding and recognition and a cytoplasmic Toll/IL-1 receptor-like (TIR) domain that is essential for intracellular signaling (Kang and Lee 2011). In mammals, 13 TLR genes have been identified, TLR1 to TLR13, but the number of genes can vary between species. For example, in humans, ten functional TLRs exist, TLR1 to TLR10, while TLR11 is a pseudogene and TLR12 and TLR13 are absent (Roach et al. 2005; Zhang et al. 2004); in mouse, 12 TLRs are found as TLR1-9 and TLR11-13 genes are functional, but TLR10 is disrupted (Roach et al. 2005).
TLRs can be subdivided into two groups according to their cellular location and type of ligand recognized. TLRs 1, 2, 4, 5, and 6 are usually expressed on the plasma membrane and interact with extracellular ligands from bacteria, parasites, and fungi (Barton and Kagan 2009). TLRs 3, 7, 8, and 9 are mainly expressed intracellularly within endosomes and/or lysosomes and detect nucleic acids with a microbial origin: TLR3 detects double-stranded RNA, TLR7, and TLR8 sense single-stranded RNA, and TLR9 recognizes bacterial single and double-stranded DNA and unmethylated CpG motifs (Arpaia and Barton 2011). TLR10 was considered an orphan TLR with no known ligands, but recent studies pinpointed a role in sensing viral and bacterial pathogens (Lee et al. 2014; Regan et al. 2013) but also as an inhibitor of the immune response (Oosting et al. 2014). TLR11 and TLR12 are located in the endosomes and recognize profilin from apicomplexan parasites (Andrade et al. 2013; Koblansky et al. 2013; Yarovinsky et al. 2005). TLR13 is an endosomal TLR that detects the 23S ribosomal RNA of gram-positive and gram-negative bacteria (Li and Chen 2012; Oldenburg et al. 2012).
In the European rabbit, orthologues of TLR1-6 and TLR8-10 have been found (Abrantes et al. 2013a; Areal et al. 2011; Astakhova et al. 2009; Kajikawa et al. 2005; Liu et al. 2012), and the expression profiles of TLR2-6 and TLR8-TLR10 have been determined (Chen et al. 2014; Liu et al. 2012). Although only a few studies have addressed their ligand specificity, it seems that rabbit TLRs act similarly to TLRs from other mammalian species. Indeed, rabbit TLR2 and TLR4 have been shown to recognize LPS from gram-negative bacteria (Chen et al. 2014; Kajikawa et al. 2005), while TLR9 senses synthetic CpG that mimics unmethylated CpG present in microbial DNA (Chuang et al. 2014; Liu et al. 2012).
Considering the impact of myxoma and rabbit hemorrhagic disease viruses in the European rabbit populations, viral-sensing TLRs might be crucial for resistance as they comprise the host first line of defense against such pathogens. The genetic characterization of the viral TLR3 from several European rabbit populations showed a pattern of diversity in agreement with the species evolutionary history with rabbits from the Iberian Peninsula showing higher levels of diversity than non-Iberian rabbits and domestic breeds (Abrantes et al. 2013a; Ferrand and Branco 2007). Determination of the expression levels of TLR3 in different rabbit organs also revealed a normal receptor activity (Chen et al. 2014). The first study on rabbit TLR7 and TLR8 reported these genes to be absent and pseudogenized, respectively (Astakhova et al. 2009). In more recent studies, TLR8 was found to be functional, although with a low activity that was attributed to a longer undefined region, but no traces of TLR7 in the rabbit genome or expression were detected (Chen et al. 2014; Lai et al. 2014). In vertebrates, TLR7 and TLR8 genes are located within a highly conserved syntenic region that surprisingly is not observed for rabbit (Abrantes et al., unpublished observations). Although TLR7 and TLR8 share a high degree of similarity and both receptors are activated by similar ligands, they appear to be maintained in most mammalian species as they present some degree of functional independence and different cellular expression (Heil et al. 2004; Jurk et al. 2002; Ohto et al. 2014). Thus, at this point, the mechanisms behind the deletion of TLR7 from the rabbit genome, the implications of effects of this deletion on the innate immune response against RNA viruses such as RHDV, and how rabbit TLR8 (or other TLRs) might compensate for this gene loss are unknown.
Antiviral proteins (RIG-I, TRIM5)
The antiviral “task force” in the Lagomorpha order is still poorly understood, but two major groups of factors have been characterized, including the restriction factor tripartite motif-containing protein 5 alpha (TRIM5α) and the pattern recognition receptors in the RIG-I-like receptor (RLR) family (retinoic acid-inducible gene-I (RIG-I/DDX58), melanoma differentiation-associated factor 5 (MDA5/IFIH1), and laboratory of genetics and physiology 2 (LGP2/DHX58)).
TRIM5α, the largest isoform encoded by the TRIM5 gene, is a cytoplasmic factor that restricts retroviral infection in a species-specific fashion (Hatziioannou et al. 2004; Nakayama et al. 2005; Perron et al. 2004; Song et al. 2005; Yap et al. 2004). Structurally, TRIM5α is constituted by an N-terminal RING domain, two B-Box domains, a long Coiled-Coil domain, and a C-terminal PRYSPRY domain, particularly composed of four “variable loops” (Nisole et al. 2005; Ozato et al. 2008). For the human immunodeficiency virus-1 (HIV-1), restriction specificity has been mapped to the PRYSPRY domain: human TRIM5α is not effective against HIV-1, while rhesus monkey TRIM5α exhibits restriction activity against the virus; however, a single amino acid alteration (R332P) in the human TRIM5α PRSYPRY domain results in a restriction activity similar to rhesus TRIM5α (Li et al. 2006; Stremlau et al. 2005; Yap et al. 2005). European rabbit and European brown hare (Lepus europaeus) TRIM5α proteins were the first to be identified in Lagomorpha and, importantly, were shown to be active against diverse retroviruses (Fletcher et al. 2010; Schaller et al. 2007). The study of Lagomorpha TRIM5α was then extended to Sylvilagus and Ochotona genera (de Matos et al. 2011). As in primates, the specificity of Lagomorpha TRIM5α restriction has been assigned to the variable loops of the C-terminal PRYSPRY domain, once striking diversity between sequences and strong evidence of positive selection were described (de Matos et al. 2011; Fletcher et al. 2010). Endoviruses like the rabbit endogenous lentivirus type K (RELIK) could have been involved in shaping the evolution of Lagomorpha TRIM5α (de Matos et al. 2011), a hypothesis recently supported by the ability of Lagomorpha TRIM5α to restrict a wide range of retroviruses, and particularly viral vectors containing RELIK capsid (Yap and Stoye 2013).
The RLRs, a family of DExD/H helicases, constitute one of the classes of PRRs able to recognize “nonself” RNA from actively replicating viruses in the cytoplasm of infected cells (Dixit and Kagan 2013; Kato et al. 2011; Kawai and Akira 2006; Yoneyama et al. 2005). As host proteins involved in pathogen detection, it is expected that RLRs would be subjected to long-term selective pressures to avoid or reduce potential infections. Positive evolutionary selection was detected for the RIG-I, MDA5, and LGP2 genes when analyzing mammalian sequences available in open databases (Lemos de Matos et al. 2013). RLR evolution was studied in the leporid genera Oryctolagus, Sylvilagus, and Lepus (Lemos de Matos et al. 2014), which include species with different susceptibility to MYXV. Particularly, RIG-I was previously described as playing a protective role in sensing MYXV in non-permissive human myeloid cells (Wang et al. 2008). The amino acid differences of positively selected sites in RIG-I between the MYXV susceptible host European rabbit and the virus natural host brush rabbit were located in the repressor domain. This protein region is responsible for viral RNA recognition and binding (Cui et al. 2008; Dixit and Kagan 2013), and the observed differences might play a determinant role in how MYXV is sensed. Two other major observations resulted from this study: first, a putative alternative splicing case in Oryctolagus and Lepus MDA5 isoforms and, second, evidence of episodic selection on MDA5 and LGP2 of the eastern cottontail (S. floridanus) (Lemos de Matos et al. 2014).
HBGAs ABO(H)
The ability of RHDV to bind to HBGA has been maintained throughout virus evolution but with the acquisition of different binding patterns to allow the infection of previously less susceptible hosts (Nyström et al. 2011). In addition, it has been shown that in rabbit populations that suffered RHDV outbreaks, there is an increase in the frequency of weakly binding phenotypes. This suggests that the virus “selected” animals with weak binding HBGA phenotypes. While understanding the genetic bases of the ABO/H polymorphism responsible for such phenotypes in rabbits has proven difficult (at least six ABO genes arranged in tandem exist in the rabbit genome; Nystrom et al., unpublished observations), it is clear that the polymorphic ABH expression generates genetic resistance to RHDV at the population level, and such resistance leads to the evolution of novel RHDV binding patterns in a textbook example of host–pathogen co-evolution (Le Pendu et al. 2014; Nyström et al. 2011)
The histo-blood group antigens ABO(H) present on the surface of red cells were the first alloantigen system to be defined in mammals (Landsteiner 1900). These antigens are complex glycans attached to proteins or lipids present on the surface of red cells and epithelial cells but might also occur as free oligosaccharides in biological fluids. The A, B, and H antigens are synthesized by the sequential action of glycosyltransferase enzymes encoded by three loci, ABO, H, and Secretor. Synthesis of these antigens begins with the addition of a fucose by α1,2 fucosyltransferases in α1,2 linkage to an oligosaccharide precursor chain forming the H antigen. In human, these transferases are the product of two genes, H or FUT1 and Secretor (Se) or FUT2, that encode enzymes with different precursor affinity. In rabbits, a third α1,2-fucosyltransferase is encoded by the Sec1 gene (Hitoshi et al. 1996) that in other mammal species is a pseudogene (Apoil et al. 2000; Borges et al. 2008; Bureau et al. 2001; Iwamori and Domino 2004).
Interesting results were observed for rabbit Sec1 as an association between a Sec1 allele and resistance, and survival during an RHDV outbreak could be established (Guillon et al. 2009). However, this Sec1 allele coded for a functional α1,2-fucosyltransferase and was always associated with active FUT2 enzymes leading the authors to hypothesize that this Sec1 allele was in linkage disequilibrium with a polymorphism in the promotor region affecting Fut2 expression. This hypothesis was proven wrong, but a novel mechanism of resistance involving the neofunctionalization of Sec1 was unraveled (Nystrom et al. 2015). Indeed, rabbit Sec1 acts as an inhibitor of Fut1, the main factor responsible for the synthesis of RHDV-binding glycans. When cells were co-transfected with Fut1 and Sec1, synthesis of RHDV binding sites was impaired, whereas this was not observed when Fut1 was transfected alone. An increase in the cellular Sec1:Fut1 ratio further decreased the abundance of binding sites. By confocal microscopy, the authors showed that Sec1 acted by displacing Fut1 to the cytoplasm from the Golgi, where it is normally located. This novel mechanism of resistance is not observed in species where Sec1 is a pseudogene. It is also associated with extensive gene conversion between Fut1, Fut2, and Sec1 in rabbits (and other leporids) which is consistent with the neofunctionalization of Sec1 in rabbits (Abrantes et al. 2009; Nystrom et al. 2015).
Adaptive immune system
Immunoglobulins
Heavy chain
Variable region
Despite having more than 200 V H genes (Ros et al. 2004; Gertz et al. 2013), the European rabbit expresses just one IGHV, the most D-proximal VH1 gene, to generate 80–90 % of its antibody repertoire (Knight and Becker 1990; Knight 1992) (Fig. 4). These VHa allotypic markers are found on all isotypes of rabbit heavy chains, are highly divergent in sequence (Kindt 1975; Margolies et al. 1977; Mage et al. 1984; 2006), and behave as Mendelian alleles (Oudin 1956, 1960; Dubiski et al. 1959; Dray et al. 1962; Kim and Dray 1972). The remaining 10–20 % of Ig molecules are encoded by the VHn genes, VHx, VHy, and VHz (Kim and Dray 1973; Horng et al. 1976; Roux 1981), that map at least 100 kb upstream of VH1 (Mage et al. 2006) and do not express VHa allotype-specific determinants. Normal rabbits that mainly rearrange the VH1a gene utilize both somatic hypermutation and a gene conversion-like mechanism to diversify rearranged heavy chain VHDH and light chain VJ genes both in the appendix and other gut-associated lymphoid tissues of the young during the development of a primary pre-immune repertoire (reviewed in Mage et al. 2006) and throughout life in germinal centers of secondary lymphoid tissues such as spleen and lymph nodes. There, during specific immune responses to infection or immunization, rabbits develop the high affinity highly specific antibodies they are known to produce (reviewed in Mage et al. 2006).
Diagram of part of the rabbit IGHV and IGHC regions from a VH1a2 rabbit described by Ros et al. (2004) adapted to annotate positions in OryCun 2.0. The OryCun 2.0 donor rabbit was heterozygous for the VH1a2 and VH1a1 allotypes. Arrows point to the VH1a2-encoding gene in overlapping clones 225P18 (AY386697) and 219D23 (AY386695) and the locations of IGHM, IGHG, and IGHE on OryCun 2.0 unplaced scaffold chrUn0439 (NW_003159763.1). Partial matches are found in unplaced scaffold chrUn1855 (NW_003161178.1) with Cα13, hs1, 2, and hs4. Note that BAC 38A2 (AY386694) with 16 VH genes does not overlap 225P18. Reprinted from Ros et al. (2004), with permission from Elsevier
Alicia was a mutant rabbit strain described by Kelus and Weiss (1986). In contrast to normal individuals of the a2 lineage, the young homozygous ali/ali mutant Alicia rabbits produced no detectable amounts of a2 molecules as newborns, and their serum contained mostly Ig resulting from rearranged genes not encoding VHa allotype-associated epitopes (VHa negative or VHn genes) (Kelus and Weiss 1986; DiPietro et al. 1990; Chen et al. 1993). It was shown that a VH1a2 gene in a parental rabbit had a deletion of 10 kb of genomic DNA containing the VH1 gene (Knight and Becker 1990; Allegrucci et al. 1990; Ros et al. 2004). As the Alicia rabbits aged, the a2 specificities appeared again in their serum (Kelus and Weiss 1977; DiPietro et al. 1990; Allegrucci et al. 1990; Chen et al. 1993; Pospisil et al. 1995). This was found to result from rearrangements of either the functional VH4 or VH7 genes localized upstream of VH1. The VH9 or a VH9-like germline gene may have served as a gene conversion donor to recreate a2 positive sequences by somatic gene conversion (Sehgal et al. 1998; Zhu et al. 1999).
The maintenance of the VHn gene usage at low frequency in VDJ rearrangements has been suggested to represent a relic of an ancestral immunological response to pathogens (Pinheiro et al. 2011). This question was addressed by sequencing VDJ genes for another leporid, genus Lepus, which separated from European rabbit 12 Mya. The results obtained revealed that Lepus also uses the VHn genes in 5–10 % of its VDJ rearrangements, showing that the VHn genes are a conserved ancestral polymorphism that has been maintained in the leporid genome and is being used for the generation of VDJ rearrangements by both Lepus and Oryctolagus genera (Pinheiro et al. 2013a).
VH region diversity is well characterized in European rabbit domestic breeds. Three serologically defined allotypic lineages exist, the so-called VHa allotypes al, a2, and a3 (±20 % amino acid sequence differences). Serological surveys in Iberian Peninsula animals revealed that sera from the subspecies algirus, when tested with VHa locus-specific alloantisera, showed no reaction at all (“a-blank”). The sequencing of rearranged VH genes expressed in these rabbits showed a new major allotypic lineage, designated a4, endemic to the Iberian Peninsula, clearly related to the known VHa genes and also showing high amino acid divergence from the a1, a2, and a3 (Esteves et al. 2004).
Su and Nei (1999) considered the three serological types distinguished in snowshoe hare (Lepus americanus) as evidence for the trans-specific nature of the VH1 polymorphism. These types had been identified by specific patterns of cross-reaction with rabbit anti a-locus allo-antisera (De Poorter 1984). The most frequent allele (am1) was characterized by anti-a2 antisera raised in a3 rabbits, the other two (am2 and am3) by two different anti-a3 sera. They were named as alleles of a hypothetical am-locus, where am stands for americanus and numbered according to their mean population frequencies. The am frequency distributions showed some similarity with those reported for the rabbit a-locus (van der Loo 1993, 1987): indeed, a systematic frequency hierarchy was observed, but the predominant hare allele am1 was most closely related with the least frequent rabbit allele a2. Su and Nei (1999) further compared the extent of sequence divergence between genes encoding the rabbit allotypes VH1al, VH1 a2, and VH1a3 with that between human and mouse VH gene sequences and concluded that, assuming a “normal” mutation rate, the VH1 polymorphism had persisted for about 50 Mya. To evaluate this hypothesis, the nucleotide sequences of VH genes expressed in specimens of Lepus species were determined. The results obtained showed that a class of Lepus sequences called a2L formed a monophyletic cluster with the VH1 sequences of the rabbit a2 allotype. It confirmed earlier observations of VHa2-related determinants in serum of Lepus timidus (van der Loo et al. 1977). The fact that this “trans-species a2 cluster” did not include genes of other rabbit VHa allotypes (a1, a3, and a4) suggests that the divergence of the VHa lineages preceded the Lepus versus Oryctolagus split (Esteves et al. 2005).
Sequencing of the VH genes in European Lepus further revealed an additional and ancient VH lineage, the sL lineage (Esteves et al. 2005; Pinheiro et al. 2013a). The finding of this ancient lineage raised the hypothesis that the VH specificities could be associated with different environments, and so VDJ genes from a third leporid genus, Sylvilagus, native to North America, were sequenced. A fifth and equally divergent VHa lineage, the a5, and an ancient lineage, the sS, related to the hares’ sL were found, but no VHn genes were observed (Fig. 5). These results show that the studied leporids employ different VH lineages in the generation of the antibody repertoire, suggesting that the leporid VH genes are subject to strong selective pressure likely imposed by specific pathogens (Pinheiro et al. 2014a).
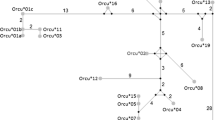
Phylogenetic tree of leporid VH genes. Human VH clan 3 genes were used to root the tree. Groups were collapsed for simplification; the number of sequences in each group is indicated. The a1, a2, a3, and a4 groups include only European rabbit sequences, the a2L and sL groups include only hares sequences, and the a5.1, a5.2, and sS groups include only S. floridanus sequences. The VHn group includes European rabbit and hare sequences. BI posterior probabilities are depicted in front of each node. Reprinted from Pinheiro et al. (2014), with permission from Springer
Constant region
The European rabbit has no gene encoding IgD (Lanning et al. 2003) and single genes encoding IgE and IgM (reviewed in Gertz et al. 2013) (Fig. 4). Differences in gene numbers of IgG, IgE, and IgA as well as their molecular structures have been noted in mammals, and the European rabbit is unique among them.
Most mammals have one IGHA gene. Humans, chimpanzees, gorillas, and gibbons have two (Kawamura et al. 1992). European rabbit has the most complex IgA system observed, with 13 IGHA genes encoding 13 IgA subclasses (Burnett et al. 1989); of these 13 subclasses, 11 are expressed and are differentially distributed among the mucosal tissues (Spieker-Polet et al. 1993) (Fig. 4). Southern analysis of genomic DNA samples from other lagomorphs (Lepus, Sylvilagus, and Ochotona) showed multiple C alpha hybridizing fragments. Thus, it is likely that all lagomorphs have multiple IgA isotypes and hence complex secretory immune systems (Burnett et al. 1989). This indicates that the IGHA gene expansion happened in the Lagomorpha ancestor and is probably an ongoing system with high variability between the different lagomorph species. Using maximum-likelihood analyses, an evolutionary study of the IgA constant regions in mammals showed 18 positively selected sites. Interestingly, two of the changes at the positively selected sites in the Calpha 1 domain, codons 45.2 and 116 (IMGT numbering, Lefranc et al. 2005a, b), can generate putative N-glycosylation sites, which can affect protein function. Residue 45.2 is a glycosylation site of IgA in several species as well as rabbit IgA7, IgA8, IgA11, and IgA13. In contrast, the putative N-glycosylation site at residue 116 appears only in rabbit IgA7, IgA8, IgA11, and IgA13 (Pinheiro et al. 2013b).
In contrast to what is seen with IgA, the European rabbit is unique in having only one IgG gene (Knight et al. 1985), for which allelic variation has been well characterized (Bernstein et al. 1983a; Martens et al. 1984; Ros et al. 2004). The vast majority of mammalian species have multiple subclasses of IgG, with horse having seven IgG subclasses (Wagner et al. 2004). The d (d11/d12) and e (e14/e15) allotypes distinguished by serology correspond to amino acid substitutions in the hinge and CH2 domains, respectively (Hamers and Hamers-Casterman 1965).
Sequencing of wild and domestic European rabbits’ IgG revealed new polymorphisms in all constant domains mostly among wild Iberian rabbits (Esteves et al 2002a; Pinheiro et al. 2014b, 2015). Serologic studies found the e15 allotype in species of the genera Oryctolagus, Lepus, Sylvilagus, Romerolagus, and Ochotona (Cazenave et al. 1987; van der Loo and Hamers-Casterman 1979) but failed to identify the d11 and d12 allotypes in Lepus and Sylvilagus genera (Hamers-Casterman et al. 1979). The sequencing of the IGHG CH2 domain for 12 species belonging to genera Oryctolagus, Sylvilagus, and Lepus showed a hotspot of variation at position 92 (e locus) with four different amino acids in Lepus species (Esteves et al. 2002b). Sequencing of the IGHG hinge domain in six species of the same genera showed no variation in amino acid length and that the variability was restricted to positions 8 and 9, with four different residues that can occur at position 9 (d locus). This variation concerns sites of potential O-glycosylation and/or proteolytic cleavage, suggesting that the underlying genetic diversity could be the outcome of selection (Esteves et al. 2006). Later studies on the evolution of the complete IGHG gene in six leporid genera, Bunolagus, Brachylagus, Lepus, Pentalagus, Romerolagus, and Sylvilagus, confirmed the two hotspots of variation in the hinge and CH2 domains and that the same length of the hinge region between CH1 and CH2 is shared by all leporids (Pinheiro et al. 2014b). In particular, 11 studied Lepus species share exactly the same hinge motif, suggesting its maintenance is a result of an advantageous structure or conformation. Pinheiro et al. (2014b) further found evidence for positive selection acting on all IgG constant domains but with a greater incidence in the CH3 domain, suggesting that the C-terminal CH3 has some functional relevance in the leporids and could be related to complement-mediated protective mechanisms against specific pathogens.
Light chain
Lambda
Early studies revealed that some rabbits have more than one copy of genes encoding constant regions of lambda light chains (IGL) at some then unknown autosomal locus (Dray et al. 1962; Mage et al. 1968; Gilman-Sachs et al. 1969). Some rabbit strains were c7 negative (e.g., the ACEP strain then at Jackson Laboratory) and only expressed c21, and some were c21 negative and only expressed c7. Mating and genetic studies showed that c7 and c21 are not alleles (Gilman-Sachs et al. 1969). Although originally thought to recognize allelic forms of lambda chains, anti-c7 and anti-c21 antisera recognize two distinct isotypic forms. The genes encoding known functional lambda light chain constant region sequences c7 (Hazyer et al. 1990) and c21 (Duvoisin et al. 1988) are found on chromosome 21 in the OryCun2 genome assembly; some lambda light chain constant region sequences are also found in OryCun2 in unplaced scaffolds (Gertz et al. 2013). The serologically distinguishable types c7 and c21 are encoded by IGLC6 and IGLC5. The amino acid sequences of IGLC6 and IGLC5 on chromosome 21 in OryCun2 differ at four codon positions confirming lambda constant region genes sequences previously reported by Hayzer et al. (1990).
In the IMGT database, positions shown for OryCun 2.0 chromosome 21 are numbered as found in contig NW_003159316.1. Approximately 3 Mb at the telomeric end is absent from the OryCun 2.0 assembly; therefore, positions 1–1922929 on contig NW_003159316.1.correspond to 3000001–4922929 on chromosome 21. The IMGT tabulation of IGLC genes found on chromosome 21 (NC_013689.1) is at http://imgt.org/IMGTrepertoire/index.php?section=LocusGenes&repertoire=genetable&species=rabbit&group=IGLC. In addition, four IGLJ genes are described, including the two that are functional and associated with IGLC6 and IGLC5, at http://imgt.org/IMGTrepertoire/index.php?section=LocusGenes&repertoire=genetable&species=rabbit&group=IGLJ, and 43 IGLV genes, of which ∼20 are functional and two potentially functional are described at http://imgt.org/IMGTrepertoire/index.php?section=LocusGenes&repertoire=genetable&species=rabbit&group=IGLV.
Interestingly, only two of the six functional IGLC genes described have been found to be expressed in the domestic rabbits analyzed. The sequences of rabbit IGCL5 and IGCL6 have been reported to be functional in association with IGLJ5 or IGLJ6. It will be important to study other leporids and wild rabbits to compare their lambda light chain gene organization and expression patterns with those observed in domestic European rabbit breeds.
Kappa
In rabbit, the IGKC- and IGKJ-containing gene fragment of the kappa light chain locus underwent a duplication leading to the existence of two different kappa light chains (IGK1 and IGK2) (Kelus and Weiss 1977). The allelic b-locus allotypes of IGKC1 may be the first immunological polymorphisms documented (reviewed in Kelus and Gell 1967). The different allotypes were initially identified by serology and were later found to reflect multiple amino acid sequence differences encoded by the IGKC1 gene fragment. At the time the nomenclature was established by Dray et al. (1962), the allotypes b4, b5, and b6 were already known. Later, a less-frequently occurring b9 type was independently discovered by Dubiski and Muller (1967) and by Carbonara and Mancini (1968).
The genetic variation of kappa light chains is particularly interesting, because the IGKC1 locus is the site of a major molecular change: the formation of a supplementary disulfide bond linking an extra Cys in the constant region domain of kappa light chains to one in the variable or the joining region (Poulsen et al. 1972; Strosberg et al. 1975; McCartney-Francis et al. 1984). Figure 6 shows the additional disulfide bond between IGKV position 80 (96 in IMGT numbering system) and IGKC1 position 171 found in b4, b5, and b6 rabbits and the alternative bond between position 108 of the J region and 171 found in some but not all kappa light chains of b9 rabbits. Phylogenetic analyses place the b9 constant region sequence closest to that of the second IGKC2 gene sequence and to those of other species (Fig. 7).
Three-dimensional structure of rabbit IGCK. An additional disulfide bond between IGKV position 80 (96 in IMGT numbering system) and IGKC1 position 171 found in b4, b5, and b6 rabbits and the alternative bond between position 108 of the J region and 171 found in some but not all kappa light chains of b9 rabbits are marked
Duplication of IGKC loci is rarely observed in vertebrates. Its existence in rabbit was confirmed by genomic data obtained by the OryCun 2.0 assembly on chromosome 2 (Gertz et al. 2013). The DNA donor rabbit appeared to be homozygous for the IGKC1 gene encoding b5 allotype, although 9 years earlier, sera typed by the Mage laboratory included both kappa allotypes b4 and b5 as well as IGHV-encoded allotypes a1 and a2 (haplotypes with VH1a1, d11, e15 and VH1a2, d12, e15). Gertz et al (2013) found that heterozygosity remained for the heavy chain allotypes even after several more years of inbreeding. This sequenced rabbit’s DNA encoded both VH1a1 and VH1a2.
Detailed analyses and tabulation of the positions of IGKC2 and its three associated JΚ genes and IGKC1 with its five associated JΚ genes and their 3′ enhancers are available in Gertz et al. (2013) where Table 4 and online resource Table 5 list further details as well as putative VΚ genes and pseudogenes found in three unplaced scaffolds. Gaps of more than 250 bp within the assembly of VK genes are also listed in Table 4 of Gertz et al. (2013). The transcriptional orientations of 86 VK genes and pseudogenes between IGKC1 and IGKC2 vary implying that only some rearrangements of VK to JΚ lead to deletion. Whether pseudogenes or functional, the retained VK remain available to contribute to diversification of rearranged VJ by gene conversion. IGKC1 and its 3′ enhancer are at position 98.93 and 98.92 Mb, respectively. Their transcriptional orientations and those of the associated IGKJ genes are on the forward strand, whereas IGKC2 at 98.22 Mb with its associated JΚ and 3′ enhancer are on the reverse strand (Gertz et al. 2013). Eight VK genes on the forward strand are found beyond the 3′ enhancer of IGKC2. Only two of the 16 VK surrounding IGKC2 encode Cys 80. The others encode Ala or Pro. It is this group of IGKV that lacks Cys 80 that are able to generate functionally rearranged light chains with the IGKC2 constant region. The remainder of the VK genes, IGKC1 and IGKC2, may all encode Cys 80. Documented gaps in the assembly plus the unknown status of 21 VK genes or pseudogenes in unplaced scaffolds make it likely that the IGK locus contains additional VK genes not placed on chromosome 2 in the current assembly. Moreover, copy number variations (CNV) were already reported within the region containing VK genes (Fontanesi et al. 2012; Carneiro et al. 2014). CNV may well reflect differences between rabbits of different strains and allotypes. VK gene sequences although similar are not identical in rabbits of different allotypes.
Population genetic evidence for adaptive molecular change at the rabbit IGKC1 locus
The rabbit expresses antibodies by preferentially assembling those gene segments that lead to a maximal amount of allo-antigenicity at both the variable domain of the heavy chain and the constant domain of the light chain: i.e., at parts that are shared by all antibody classes. This is remarkable as more plesiomorphic functional gene fragments (phylogenetically less derived) are available. The plesiomorphic IGKC2 or bas locus (Kelus and Weiss 1977) has developed low expression (Emorine and Max 1983; Bernstein et al. 1984; Sperry et al. 1989) but is upregulated in expression in the Basilea strain (bas) where IGK1 expression has become impossible due to a mutation in the acceptor site for splicing IGKJ to IGKC1 (Lamoyi and Mage 1985).
The degree of inter-allelic diversity of the rabbit IGKC1 gene fragment has been documented in numerous studies, revealing pairwise amino acid differences of up to 40 % between the four allotypes occurring in domestic breeds and their feral descendants (Bernstein et al. 1983b). The discovery of more “wild-type” b-alleles in the original distribution range of the genus (Iberia and Maghreb; reviewed in Cazenave et al. 1987; van der Loo et al. 1991, 1999a) did not significantly affect this mean distance. Such divergence is similar or even surpasses what is currently observed at vertebrate MHC loci, where average pairwise amino acid distances between alleles are generally 10–12 % and rarely exceed 25 % at exons encoding the antigen-binding clefts (Gutierrez-Espeleta et al. 2001).
The extreme allelic diversity at the IGCK1 light chain has so far been only found in rabbit and led to speculations of a possible causal link with the emergence of the inter-domain disulfide bond (McCartney-Francis et al. 1984). For reaching some understanding of the underlying evolutionary pathways, it is paramount to know the genomic basis of the polymorphism, which had been confused in the past by what had been called “latent allotypes.” The debate was settled by genomic studies, first by Northern blotting (Emorine et al. 1984; Matthyssens et al. 1985) and more recently by the rabbit whole genome sequence OryCun 2.0 (Gertz et al. 2013). These studies provide no support for a genomic base of the “latent allotype” phenomenon which has so far only been observed in blood samples. This subject was reviewed extensively by McCartney-Francis (1987) who concluded that “…the current molecular data suggests that latent allotypy is not a generalized phenomenon in the rabbit.” Clearly, the b locus alleles are products of a single IGKC1 gene.
Whereas innumerable papers and monographs question the deterministic mechanisms shaping MHC diversity, evidence of diversity enhancement selection at the rabbit IGKC1-locus is comparatively well supported by a number of evolutionary studies, both at the population genetic and molecular levels. At the molecular level, among the more salient features are the highly significant dn/ds ratios (i.e., excess of nonsynonymous versus synonymous nucleotide substitutions), culminating in the unprecedented situation where inter-allelic variation at the JK-CK1 gene region is essentially limited to the coding regions (Bernstein et al. 1983a, b). For detailed comparisons of evolutionary patterns at the IGKC1-locus with those at MHC loci, see van der Loo and Verdoodt (1992); van der Loo et al. (1999b).
Studies at the population level revealed significant and systematic non-random allele correlations such as heterozygous excess and digenic disequilibria (van der Loo et al. 1987, 1993; van der Loo et al. 1987, 1996). In a worldwide study of feral rabbit populations, allele frequencies at the b-locus were more similar among continents than among populations within continents (i.e., F st about zero, which is a strong indication of balancing selection; van der Loo 1993). What is more, these frequencies matched the allelic imbalance in gene expression. Indeed, in the context of allelic exclusion, heterozygous rabbits constitutively use one allele more than the other (“pecking order” b4 > b5/b6 > > b9; Lummus et al. 1967; Kerremans 1982). Whatever may be the reason why heterozygous b4/b9 rabbits use the b4 gene about three times more often than the b9 gene, the fact that this b9 allele appears to be maintained at low frequency in all areas studied provides explicit evidence of frequency-dependent selection.
As with MHC, the selection forces underlying the observed departures of expectations under neutrality are such that it seems unlikely that they are due to “survival of the fittest” but rather to some mechanism which favors the generation of heterozygotes (“selection-anticipating” mechanism; van der Loo and Verdoodt 1992). Studies on feral rabbits indicate that heterozygous excess is similar among age groups (van der Loo, unpublished). The absence of differential survival rates is also supported by the fact mentioned above that the much higher level of gene diversity in the native range of the species does not lead to increased disease resistance and is part of the “conservation paradox” discussed in Lees and Bell (2008). Disassortative mating system and maternal–fetal effects are among models which in rabbit could favor the selective absorption of embryos sharing maternal genotypes (Potts et al. 1991) or sperm competition [reviewed in Bernatchez and Landry (2003)]. Although the richness of well-documented genetic markers makes rabbit suitable for studies on genetically based mating preferences, we are not aware of such studies.
The trans-species nature, another benchmark of MHC evolution (Figueroa et al. 1988), was also documented for the rabbit b-locus allotypes (Bouton and van der Loo 1997). In contrast with MHC, where allelic divergence is believed to be uniquely due to very long allele persistence times and rates of mutant-acceptance are “normal” and uniform (Klein et al. 1998), at the b-locus, evolutionary rates can differ greatly, i.e., rate monotony was rejected at P < 0.0001 by the likelihood rate test (Felsenstein 1981; van der Loo et al. 1999a) (Fig. 8). An important difference between both loci is the number of alleles (about 20 for IGKC1 versus more than 1,000 for human MHC class II). This difference could in part be explained by Klein’s hypothesis that MHC allelic lineages can be maintained over tens of millions of years, whereas in the b-locus, allelic turnover rates appear much shorter, at least for the predominant lineages. Indeed, the predominant alleles of the subspecies O. c. cuniculus were replaced by related but different alleles in subspecies O. c. algirus (van der Loo et al. 1999a).
Comparing allelic divergence with inter-species divergence at IGKC loci: Distance trees were obtained using MEGA software (Tamura et al. 2013) applied to the CDS regions of mammalian IGKC genes (312 base pairs). Whereas synonymous distances (Ks) between leporid IGKC1 are similar to those of between IGKC genes of higher primates and consistent with 20 Mya of separation, rates at amino acid replacement sites (Ka) can differ significantly among rabbit alleles, where distances can approach those between placental mammals separated for 80 My. Mammalian IGKC’s are represented by mouse (mm), rat (rr), sheep (oa), swine (ss), horse (ec), monkey (mf), chimp (pt), and human (hs). Leporid IGKC1 genes are bla1 (snowshoe hare) and rabbit allotypes (b9, b4, b4wc, b4wd, b5, b5wf, b95, b6, b6w2, and b9). Rabbit IGKC2 (bas1). The inter-species distances Ks and Ks between rabbit b4wc and hare bla1 are similar to those between human and monkey, whereas inter-allele distances between IGKC1 alleles can be much larger and Ka/Ks ratios significantly higher (not shown)
Taken together, the actual “state of the art” suggests the existence of an intrinsic relationship between the unusual evolutionary patterns at the b locus and a process of molecular optimization related to the supplementary disulfide bond. The optimization through structural stabilization of the molecules by the extra interdomain disulfide bond (Traxlmayr and Obinger 2012) may contribute to the well-known stability of rabbit antibodies and maintenance of this stability during development of high specificity via the processes of somatic hypermutation and gene conversion.
MHC or RLA
A standardized nomenclature for the rabbit MHC (RLA) class I and class II genes and alleles has not yet been established (reviewed in Mage and Rogel-Gaillard 2016). The HLA complex in man between myelin oligodendrocyte glycoprotein (MOG) and kinesin family member C1 (KIFC1) spans 3.6 Mb. Mapping of RLA to chromosome 12 at position q1.1 (Rogel-Gaillard et al. 2001) is confirmed in the OryCun 2.0 assembly. However, MOG and KIFC1 are mapped less than 2 Mb apart. It is probable that the OryCun 2.0 assembly in the RLA region is incomplete. Recent sequences of BACs from a New Zealand White rabbit homozygous for MHC that do not completely cover the RLA region already have a total length of 2 Mb (Rogel Gaillard, C. personal communication and reviewed in Mage and Rogel-Gaillard 2016). In NCBI annotation release 101, rabbit beta-2-microglobulin (B2M) is mapped to chromosome 17 NC_013685.1 (27607481-27614097) on the reverse strand.
Early studies of a rabbit RLA class Ia gene reported similar organization to HLA (Marche et al. 1985). Characterization and expression of the rabbit class II beta genes DQ, DR, and DP were reported by the laboratory of Katherine Knight between 1988 and 1993 (reviewed in Mage and Rogel-Gaillard 2016). Rabbit DM genes (DMA and DMB) were characterized, and the relative order of class II genes in the rabbit MHC was reported by Hermel et al. (1999). Although Fain et al. (2001) only found a single copy of the MHC class II DQA gene in O. cuniculus, they identified six alleles. A population genetic study analyzed the exon 2 of the DQA gene in European rabbit populations from Portugal, Spain, and France and in domestic breeds identified 28 alleles. The Iberian rabbit populations were well differentiated from the French population and domestic breeds and had the higher allelic diversity. However, the DQA nucleotide diversity was higher in the French population (Magalhães et al. 2015). A study that compared allelic diversity and evolutionary history of the DQA gene in wild and domesticated breeds of Oryctolagus, with Pronolagus and several species of Lepus and Sylvilagus (Surridge et al. 2008), showed persistence of some alleles present before divergence of the genera. The authors suggest that trans-species evolution and maintenance of DQA alleles over millions of years of leporid evolution probably result from balancing selection driven by exposure to pathogens and that alleles may have increased in diversity by recombination and possibly by gene conversion. Two papers about the role of MHC diversity studied in European rabbits (O. cuniculus) (Oppelt et al. 2010) and European brown hares (L. europeaus) (Smith et al. 2010) were summarized in a perspective by Oliver and Piertney (2010). In their overview, they wrote:
“… MHC diversity may be defined by an amalgam of effects, incorporating reproductive success, mating systems, antagonistic coevolution with parasites, fluctuating selection pressures, heterozygote advantage or copy number optimality, population demography and extrinsic environmental factors, all of which are likely to be context dependent.”
T cells
T cells express a unique antigen-binding receptor designated T cell receptor (TR) that binds specific antigens and triggers an immune response. T cells are divided into αβ and γδ according to the heterodimeric TR chains expressed and while αβ TRs recognize antigenic fragments presented by class I or class II molecules of the major histocompatibility complex, γδ TRs can also recognize free antigens (Hayday 2000). The TR chains are encoded by four distinct gene families, α, β, γ, and δ, located within three chromosomal loci, TRA/TRD, TRB, and TRG. Like immunoglobulins, complete T cell antigen receptor chains are assembled through a somatic rearrangement process that combines variable (V), diversity (D; present only in the β and δ chains), joining (J), and constant (C) gene segments. This process generates an array of receptor specificities from a limited set of germline elements. Rabbits, in contrast to most species where αβ T cells are the predominant T cell population, are considered a “γδ high” species as γδ TRs account for 20–50 % of the T cells (Porcelli and Modlin 1999).
In rabbits, only the genomic organization of the γ and β chains loci, TRG and TRB, respectively, has been disclosed. The rabbit TRG locus is located in chromosome 10, flanked by the genes steroidogenic acute regulatory protein D3-N-terminal like (STARD3NL) and amphysin (AMPH), and spans ∼70 kb. Although the genomic organization of the TRG locus is highly variable between species, the rabbit TRG locus appears to be the smallest and simplest among the mammalian TRG loci described so far. Indeed, the locus is composed of ten TRGV, two TRGJ, and one TRGC genes (Massari et al. 2012). The TRGV genes belong to four different subgroups, TRGV1-4, with seven genes belonging to the TRGV1 subgroup, while the remaining subgroups are composed by a single gene. Evolutionary analyses suggest that the TRGV1 genes arose by several rounds of duplication (Cho et al. 2005; Massari et al. 2012). The two TRGJ genes, TRGJ1 and TRGJ2, have different length, but the TRGJ characteristic FGXG amino acid motif is present in both genes. Two variant forms of TRGC can be produced by alternative splicing of the connecting region of exon 2 (Isono et al. 1995). Much of the diversity of the TRG locus results from the variability at the VJ junctions due to the addition of a few nucleotides (N regions) and the deletion of some nucleotides from the V and J segments (Isono et al. 1995).
The rabbit TRB locus has not yet been assigned to a chromosome, but it is contained within the conserved mammalian syntenic block that is flanked by the DBH-like monooxygenase protein 2 (MOXD2) and ephrin type-B receptor 6 (EPHB6) genes at the 5′ and 3′ end, respectively. The rabbit TRB locus spans ∼600 kb and, as in most other species, the general structural organization is maintained with a pool of 74 TRBV genes located upstream of two D-J-C gene clusters positioned in tandem, followed by one TBRC gene and one TRBV gene with an inverted transcriptional orientation at the 3′ end (Antonacci et al. 2014). The 74 TRBV genes are distributed in 24 subgroups with six of the subgroups presenting more than one gene; 16 of these genes are pseudogenes. The high number of rabbit TRBV genes seems to be associated with massive duplications within the TRBV subgroups, followed by diversification, rather than to the appearance of novel subgroups. Interestingly, the emergence of some of the subgroups occurred at more than 90 million years ago, before divergence of rabbit from dog, mouse, and human (Antonacci et al. 2014; Lamoyi and Mage 1987). Each J-D-C cluster is composed of one TRBD, six TRBJ, and one TRBC genes. The rabbit TRBC genes exhibit great similarity at the nucleotide and amino acid level and are probably under strong concerted evolutionary pressures, but their 3′ UTR is highly divergent (Antonacci et al. 2014).
In contrast to the TRG and TRB loci, the genomic organization of the rabbit TRA/TRD locus has not yet been reported. The TRA/TRD locus is unique as it contains elements for distinct TR chains, and some V genes are expressed in both α and δ chains. Comparative genomics showed that mammals, including marsupials, share a similar gene organization at this locus, with most of the V elements located at the 5′ end of the locus and a single TRDV gene with inverted orientation located at the 3′ end (Bosc and Lefranc 2003; Connelley et al. 2014; Glusman et al. 2001; Parra et al. 2008). This suggests that this genomic region is highly conserved, despite the variable number of TRAV/TRDV genes found in mammalian genomes. In mammals, the TRA/TRD locus is part of a syntenic block flanked by olfactory receptor (OR) genes and by the defender against cell death gene 1 (DAD1) gene at the 5′ and 3′ ends, respectively (Bosc and Lefranc 2003; Connelley et al. 2014; Glusman et al. 2001; Parra et al. 2008). A scaffold located in the rabbit chromosome 17 and flanked by these same genes is available at ENSEMBL (ensembl.org; scaffold GL018676). Further analysis of this scaffold confirmed the co-localization of the TRA and TRD genes as a few TRAV and TRDV genes annotated, as well as TRAC, TRDC, and a “bifunctional” TRAVDV gene, but J segments are not annotated. The scaffold also suggests that the rabbit TRA/TRD locus spans ∼1 Mb. As in other mammals, rabbit 5′ TRAV genes seem to be interspersed with the OR loci, and a TRDV gene at the 3′ end also appears in an inverted orientation (Bosc and Lefranc 2003; Connelley et al. 2014; Glusman et al. 2001; Parra et al. 2008). Although incompletely annotated, the rabbit TRA/TRD locus presents the overall organization of the mammalian TRA/TRD locus and immediate flanking regions.
Endogenous viruses
The endogenous retroviruses are genetic fossils that can give us useful information about the evolutionary history of this virus family (Feschotte and Gilbert 2012; Stoye 2012). In retroviral infections (family Retroviridae), integration of the viral DNA into the host genome, after reversing from RNA, is used as a mechanism of replication. Occasionally throughout evolution, the host infection by exogenous retroviruses resulted in stably integrated endogenous retroviruses (ERVs) that can be transmitted vertically from one host generation to the next (Gifford and Tristem 2003).
The first endogenous retrovirus detected in the lagomorphs was an endogenous gamma-retrovirus, rabbit endogenous retrovirus-H (RERV-H), which was originally thought to be a human retrovirus. It was found in the European rabbit genome, encoding a functional viral protease (Griffiths et al. 2002; Voisset et al. 2003). Interestingly, the searching of this endogenous gamma-retrovirus in other lagomorphs showed that is apparently restricted to European rabbit.
The first described endogenous lentivirus was found in the European rabbit genome and was named the RELIK (Katzourakis et al. 2007). The search for conserved regions of this lentivirus in other lagomorphs showed that the RELIK was also present in the genome of other leporids like B. monticularis, L. granatensis, L. europaeus, and S. brasiliensis but absent in Ochotona, suggesting that the insertion of RELIK happened after the split of Ochotonidae and Leporidae family in the leporids’ ancestor that happened at 12 Mya (Keckesova et al. 2009; van der Loo et al. 2009).
Recently, a genomic mining of endogenous viral sequences of mouse mammary tumor virus (MMTV) in the European rabbit and Ochotona species was performed. No significant MMTV viral elements were found in the European rabbit genome. On the other hand, significant matches of MMTV viral elements were found in American pika (Ochotona principes) genome, denominated pika-BERV. This new pika-BERV was found in other pika species. The finding of different insertions of endogenous beta-retro virus (EBVs) in Ochotona genomes might support the possible existence of infecting exogenous beta-retroviruses, after the divergence of Ochotonidae and Leporidae families (Lemos de Matos et al. 2015)
Future perspectives
Infectious diseases, therapeutic development, and autoimmune disease models
Infectious diseases
-
1.
Tuberculosis: As early as 1865, Jean-Antoine Villemin transmitted tuberculosis to rabbits (Daniel 2015; Villemin 2015). In the 1920s, Max B. Lurie had compared “the fate of human and bovine tubercle bacilli in various organs of the rabbit” (Lurie 1928) and embarked on years of studies in the rabbit. Some colonies of rabbits are more susceptible to bovine strains (Mycobacterium bovis) and relatively resistant to human-derived strains of Mycobacterium. Lurie bred and selected lines of rabbits with different degrees of susceptibility to infection. Although these rabbit lines were not maintained, a revisiting of Lurie’s genetic experiments (Werneck-Barroso 1999) makes a strong case for the conclusion that “this monumental study of natural contagion may be one of the strongest available pieces of in vivo evidence for the role of innate resistance in natural tuberculous infection.” A book published in 2006, also “updates and revises the ground-breaking work of Max B. Lurie”…and “summarizes the combined 80 years of research by Drs. Lurie and Dannenberg” (Dannenberg 2006). With Dannenberg, Manabe et al. (2003) reported that the M. tuberculosis strain genotype correlated with “the relative pathogenicity, including disease severity, in the rabbit model” and suggested genotyping could provide a means to correlate “stage-specific M. tuberculosis genes important in human disease.” The OryCun 2.0 rabbit genome assembly used DNA from a single female rabbit of the Thorbecke partially inbred strain that was more susceptible to tuberculosis than outbred New Zealand White rabbits (Dorman et al. 2004; Mendez et al. 2008). Unfortunately, the genetic basis for the differences was not yet determined when the entire strain was lost in a fire shortly after the sequencing project was initiated. Nevertheless, studies of tuberculosis using the rabbit model continue to make major contributions (Subbian et al. 2013a; 2013b; Tsenova et al. 2014).
-
2.
Syphilis: Robert Koch used the rabbit as an animal model for his pioneering work, and rabbit remains the best model for studies of syphilis. A recent study with extensive references can be found, for example, in Salazar et al. (2007) who used quantitative real-time PCR to analyze kinetics of distribution of Treponema pallidum DNA into tissues and organs of rabbits after intra-testicular infection.
-
3.
HIV: HIV vaccine studies in rabbits include reports of Tervo and Keppler (2010), Pan et al. (2013), and Chen et al. (2013). Heyndrickx et al. (2013) described producing gp140 trimers derived from selected patient isolates as immunogens and concluded that “the rabbit is a valuable model for HIV vaccine studies.” Sanders et al. (2015) made extensive use of rabbits for studies of native-like trimers as immunogens for development of HIV-1 vaccines to induce broadly neutralizing antibodies, and Qin et al. (2015) characterized a large panel of rabbit monoclonal antibodies elicited to the V3 loop of gp120 that included “the most potent and broadly neutralizing anti-V3 loop mAb isolated from a vaccinated animal or human to date.”
-
4.
Papillomavirus: Human papillomavirus (HPV) is highly species specific. The S. floridanus (eastern cottontail rabbit) papillomavirus (CRPV) and a rabbit model contributed to the development of two prophylactic vaccines (Gardasil and Cervarix). However, therapeutics including vaccines for established infection are still needed, and the CRPV rabbit model continues to contribute to such investigations (Hu et al. 2006, 2007, 2014).
Select agents, vaccines, and therapeutics
Rabbits have been valuable models for testing and development of vaccines and therapeutics including those against select agents.
-
1.
Tularemia: Tularemia is a threat to humans via natural infections and the causative agent of tularemia, Francisella tularensis, is also listed as a select agent. Rabbits and hares are natural hosts for Francisella tularensis (reviewed in Foley and Nieto 2010; Thomas and Schaffner 2010; Sjöstedt 2011).
-
2.
Anthrax: The rabbit serves as a valuable model for studies of the select agent Bacillus anthracis and contributed to development of the therapeutic monoclonal antibody Raxibacumab for use in treatment of inhalation anthrax. In 2012, it was the first therapeutic approved by the United States Food and Drug Agency (FDA) under the animal efficacy rule after using only rabbits and monkeys (reviewed in Kummerfeldt 2014).
-
3.
Pox viruses: Therapeutics for orthopox infections are needed for treatment of natural infections with monkeypox virus, secondary accidental exposures, or complications of Vaccinia immunizations. A review of animal models used in investigations of therapeutics refers to several studies that used rabbit pox models Smee (2013). Smee (2013) also notes that although less likely, “potential deliberate release of smallpox (Variola) and monkeypox viruses” are of concern.
-
4.
Therapeutics: Rabbit anti-thymocyte globulin-Genzyme™ (Thymoglobulin) is a widely used FDA-approved therapeutic produced by immunization of rabbits with human thymocytes (reviewed in Deeks and Keating 2009). It is used for immunosuppression regimens including in transplant patients during acute rejection of kidney transplants and to prevent graft-versus-host disease after allogeneic hematopoietic stem cell transplantation. This therapeutic can only be used in immunosuppressed patients because it is a fully rabbit product that would be recognized as a foreign antigen. A second similar product, ATG-Fresenius S®, is produced by immunization of rabbits with human Jurkat T cells. Gharekhani et al. (2013) reviewed the two therapeutics used in solid organ transplantation, and Storeck et al. (2015) reviewed their use in allogeneic hematopoietic cell transplantation.
Humanized rabbit monoclonal and polyclonal antibodies are under development as therapeutics. Rabbits with human immunoglobulin genes are currently being developed to provide a source for polyclonal therapeutics produced in rabbits (Flisikowska et al. 2011). Additional information can be found in patents, most recently US Patent 08652842 concerning “Humanized immunoglobulin loci.”
Partially humanized rabbit mAbs produced in b9 rabbits and selected by phage display (Popkov, et al. 2003; Rader 2009) are particularly useful for preclinical investigations in mouse models because they may react with human and mouse target antigens (Popkov et al. 2004). A high affinity chimeric rabbit/human Fab selected by phage display co-crystallized with HIV-1 Rev (Stahl et al. 2010) leading to determination of the crystal structure of this protein that plays an essential role in viral replication (DiMattia et al. 2010). Its high affinity (∼40 pM) and ability to depolymerize self-associated Rev suggest therapeutic potential. Zhuang et al. (2014) demonstrated that the Fab with an attached Tat cell-penetrating peptide has high anti-HIV-1 activity and low cytotoxicity.
Autoimmune disease models
Models in rabbit include those for systemic lupus erythematous (SLE) (James et al. 1995), neuropsychiatric SLE (reviewed in Mage and Rai 2012), and experimental autoimmune encephalitis (reviewed in Baxter 2007).
Future sequencing of lagomorph genomes
The Lagomorph Genomics Consortium (LaGomiCs) arose as a collaboration between the World Lagomorph Society (WLS) (www.worldlagomorphsociety.org) and the European COST Action TD1101, “A Collaborative European Network on Rabbit Genome Biology—RGB-Net 2011-2015.” The stated objective of LaGomiCs is to “sequence the genomes of all extant and select extinct lagomorph species” in the coming years. Some of the efforts are under way, and a publication that summarizes a lengthy detailed “white paper” drafted during meetings of consortium members has been (submitted) for publication.
With the advent of “next-generation sequencing,” it has become feasible to obtain more extensive information about genomic sequences of lagomorphs. However, obtaining complete and accurate assemblies of complex loci such as those of the immunoglobulin genes, MHC, and TCR regions remains challenging. New approaches, such as via deep sequencing, better assembly strategies including assembly of sequences of overlapping BAC clones, and third generation long reads will be needed before such complex regions are fully mapped and annotated.
References
Abrantes J, Esteves PJ, Carmo CR, Muller A, Thompson G, van der Loo W (2008) Genetic characterization of the chemokine receptor CXCR4 gene in lagomorphs: comparison between the families Ochotonidae and Leporidae. Int J Immunogenet 35:111–117. doi:10.1111/j.1744-313X.2007.00735.x
Abrantes J, Posada D, Guillon P, Esteves PJ, Le Pendu J (2009) Widespread gene conversion of alpha-2-fucosyltransferase genes in mammals. J Mol Evol 69:22–31
Abrantes J, Carmo CR, Matthee CA, Yamada F, Loo W, Esteves PJ (2011) A shared unusual genetic change at the chemokine receptor type 5 between Oryctolagus, Bunolagus and Pentalagus. Conserv Genet 12:325–330. doi:10.1007/s10592-0099990-1
Abrantes J, Lopes AM, Esteves PJ (2012a) Complete genomic sequences of rabbit hemorrhagic disease virus G1 strains isolated in the European rabbit original range. J Virol 86:13886. doi:10.1128/JVI.02683-12
Abrantes J, van der Loo W, Le Pendu J, Esteves PJ (2012b) Rabbit haemorrhagic disease (RHD) and rabbit haemorrhagic disease virus (RHDV): a review. Vet Res 43:12. doi:10.1186/1297-9716-43-12
Abrantes J, Areal H, Esteves PJ (2013a) Insights into the European rabbit (Oryctolagus cuniculus) innate immune system: genetic diversity of the toll-like receptor 3 (TLR3) in wild populations and domestic breeds. BMC Genet 14:73. doi:10.1186/1471-2156-14-73
Abrantes J, Lopes AM, Dalton KP, Melo P, Correia JJ, Ramada M, Alves PC, Parra F, Esteves PJ (2013b) New variant of rabbit hemorrhagic disease virus, Portugal, 2012-2013. Emerg Infect Dis 19:1900–1902. doi:10.3201/eid1911.130908
Alda F, Gaitero T, Suárez M, Merchán T, Rocha G, Doadrio I (2010) Evolutionary history and molecular epidemiology of rabbit haemorrhagic disease virus in the Iberian Peninsula and Western Europe. BMC Evol Biol 10:347. doi:10.1186/1471-2148-10-347
Allegrucci M, Newman BA, Young-Cooper GO, Alexander CB, Meier D, Kelus AS, Mage RG (1990) Altered phenotypic expression of immunoglobulin heavy-chain variable-region (VH) genes in Alicia rabbits probably reflects a small deletion in the VH genes closest to the joining region. Proc Natl Acad Sci U S A 87:5444–5448
Almeida T, Lopes AM, Magalhães MJ, Neves F, Pinheiro A, Gonçalves D, Leitão M, Esteves PJ, Abrantes J (2015) Tracking the evolution of the G1/RHDVb recombinant strains introduced from the Iberian Peninsula to the Azores islands, Portugal. Infect Genet Evol. doi:10.1016/j.meegid.2015.07.010
Alves JM, Carneiro M, Afonso S, Lopes S, Garreau H, Boucher S, Allain D, Queney G, Esteves PJ, Bolet G, Ferrand N (2015) Levels and patterns of genetic diversity and population structure in domestic rabbits. Plos One (in press)
Andrade WA et al (2013) Combined action of nucleic acid-sensing Toll-like receptors and TLR11/TLR12 heterodimers imparts resistance to Toxoplasma gondii in mice. Cell Host Microbe 13:42–53. doi:10.1016/j.chom.2012.12.003
Antonacci R, Giannico F, Ciccarese S, Massari S (2014) Genomic characteristics of the T cell receptor (TRB) locus in the rabbit (Oryctolagus cuniculus) revealed by comparative and phylogenetic analyses. Immunogenetics 66:255–266. doi:10.1007/s00251-013-0754-1
Apoil PA, Roubinet F, Despiau S, Mollicone R, Oriol R, Blancher A (2000) Evolution of alpha 2-fucosyltransferase genes in primates: relation between an intronic Alu-Y element and red cell expression of ABH antigens. Mol Biol Evol 17:337–351
Aragão HDB (1927) Myxoma dos coelhos. Memórias do Instituto Oswaldo Cruz 20:237–247
Areal H, Abrantes J, Esteves PJ (2011) Signatures of positive selection in Toll-like receptor (TLR) genes in mammals. BMC Evol Biol 11:368. doi:10.1186/1471-2148-11-368
Arpaia N, Barton GM (2011) Toll-like receptors: key players in antiviral immunity. Curr Opin Virol 1:447–454. doi:10.1016/j.coviro.2011.10.006
Astakhova NM, Perelygin AA, Zharkikh AA, Lear TL, Coleman SJ, MacLeod JN, Brinton MA (2009) Characterization of equine and other vertebrate TLR3, TLR7, and TLR8 genes. Immunogenetics 61:529–539. doi:10.1007/s00251-009-0381-z
Barton GM, Kagan JC (2009) A cell biological view of Toll-like receptor function: regulation through compartmentalization. Nat Rev Immunol 9:535–542. doi:10.1038/nri2587
Baxter AG (2007) The origin and application of experimental autoimmune encephalomyelitis. Nat Rev Immunol 7:904–912
Bernatchez L, Landry C (2003) MHC studies in nonmodel vertebrates: what have we learned about natural selection in 15 years? J Evol Biol 16:363–377
Bernstein KE, Alexander CB, Mage RG (1983a) Nucleotide sequence of a rabbit IgG heavy chain from the recombinant F-I haplotype. Immunogenetics 18:387–397
Bernstein KE, Skurla RM, Mage RG (1983b) The sequences of rabbit κ light chains of b4 and b5 allotypes differ more in their constant regions than in their 3' untranslated regions. Nucl Acids Res 11:7205–7214
Bernstein KE, Lamoyi E, McCartney-Francis N, Mage RG (1984) Sequence of a cDNA encoding Basilea kappa light chains (K2 isotype) suggests a possible relationship of protein structure to limited expression. J Exp Med 159:635–640
Best SM, Kerr PJ (2000) Coevolution of host and virus: the pathogenesis of virulent and attenuated strains of myxoma virus in resistant and susceptible European rabbits. Virology 267:36–48
Best SM, Collins SV, Kerr PJ (2000) Coevolution of host and virus: cellular localization of virus in myxoma virus infection of resistant and susceptible European rabbits. Virology 277:76–91
Biju-Duval C, Ennafaa H, Dennebouy N, Monnerot M, Mignotte F, Soriguer RC, Gaaied A, Hili A, Mounolou JC (1991) Mitochondrial DNA evolution in lagomorphs: origin of systematic heteroplasmy and organization of diversity in European rabbits. J Mol Evol 33:92–102
Borges BN, Paiva TS, Harada ML (2008) Evolution of the SEC1 gene in New World monkey lineages (Primates, Platyrrhini). Genet Mol Res 7:663–678
Bosc N, Lefranc MP (2003) The mouse (Mus musculus) T cell receptor alpha (TRA) and delta (TRD) variable genes. Dev Comp Immunol 27:465–497
Bouton C, van der Loo W (1997) The trans-species nature of rabbit b locus polymorphism is supported by studies on the snow-shoe hare. Immunogenetics 45:444–446
Branco M, Ferrand N, Monnerot M (2000) Phylogeography of the European rabbit (Oryctolagus cuniculus) in the Iberian Peninsula inferred from RFLP analysis of the cytochrome b gene. Heredity 4:307
Branco M, Monnerot M, Ferrand N, Templeton AR (2002) Postglacial dispersal of the European rabbit (Oryctolagus cuniculus) on the Iberian Peninsula reconstructed from nested clade and mismatch analyses of mitochondrial DNA genetic variation. Evolution 56:792
Bureau V, Marionneau S, Cailleau-Thomas A, Le Moullac-Vaidye B, Liehr T, Le Pendu J (2001) Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC. Eur J Biochem 268:1006–1019
Burnett RC, Hanly WC, Zhai SK, Knight KL (1989) The IgA heavy-chain gene family in rabbit: cloning and sequence analysis of 13 C alpha genes. EMBO J 8:4041–4047
Cabrera (1914) Fauna Ibérica. Museo Nacional de Ciências Naturales de Madrid, Madrid
Camarda A et al (2014) Detection of the new emerging rabbit haemorrhagic disease type 2 virus (RHDV2) in Sicily from rabbit (Oryctolagus cuniculus) and Italian hare (Lepus corsicanus). Res Vet Sci 97:642–645
Cameron C, Hota-Mitchell S, Chen L, Barrett J, Cao JX, Macaulay C, Willer D, Evans D, McFadden G (1999) The complete DNA sequence of myxoma virus. Virology 264:298–318
Carbonara A, Mancini G (1968) Ati Ass Genet, Ital. 13:229
Carmo CR, Esteves PJ, Ferrand N, van der Loo W (2006) Genetic variation at chemokine receptor CCR5 in leporids: alteration at the 2nd extracellular domain by gene conversion with CCR2 in Oryctolagus, but not in Sylvilagus and Lepus species. Immunogenetics 58:494–501
Carneiro M, Afonso S, Geraldes A, Garreau H, Bolet G, Boucher S, Tircazes A, Queney G, Nachman MW, Ferrand N (2011) The genetic structure of domestic rabbits. Mol Biol Evol 28:1801–1816
Carneiro M, Rubin CJ, Di Palma F, Albert FW, Alföldi J, Barrio AM, Pielberg G, Rafati N, Sayyab S, Turner-Maier J, Younis S, Afonso S, Aken B, Alves JM, Barrell D, Bolet G, Boucher S, Burbano HA, Campos R, Chang JL, Duranthon V, Fontanesi L, Garreau H, Heiman D, Johnson J, Mage RG, Peng Z, Queney G, Rogel-Gaillard C, Ruffier M, Searle S, Villafuerte R, Xiong A, Young S, Forsberg-Nilsson K, Good JM, Lander ES, Ferrand N, Lindblad-Toh K, Andersson L (2014) Rabbit genome analysis reveals a polygenic basis for phenotypic change during domestication. Science 345:1074–1079
Cazenave PA, Bennamar A, Sogn JA, Kindt TJ (1987) Immunoglobulin genes in feral population. In: Dubiski, S., editor. The rabbit in contemporary immunological research. Longman Scientific & Technical; p. 148
Cedar H, Bergman Y (2008) Choreography of Ig allelic exclusion. Curr Opin Immunol 20:308–317
Charo IF, Ransohoff RM (2006) The many roles of chemokines and chemokine receptors in inflammation. N Engl J Med 354:610–621. doi:10.1056/NEJMra052723
Chen HT, Alexander CB, Young-Cooper GO, Mage RG (1993) VH gene expression and regulation in the mutant Alicia rabbit. Rescue of VHa2 allotype expression. J Immunol 150:2783–2793
Chen Y, Vaine M, Wallace A, Han D, Wan S, Seaman MS, Montefiori D, Wang S, Lu S (2013) A novel rabbit monoclonal antibody platform to dissect the diverse repertoire of antibody epitopes for HIV-1 Env immunogen design. J Virol 87:10232–10243
Chen C, Zibiao H, Ming Z, Shiyi C, Ruixia L, Jie W, SongJia L (2014) Expression pattern of Toll-like receptors (TLRs) in different organs and effects of lipopolysaccharide on the expression of TLR 2 and 4 in reproductive organs of female rabbit. Dev Comp Immunol 46:341–348. doi:10.1016/j.dci.2014.05.008
Cho KS, Zhai SK, Esteves PJ, Knight KL (2005) Characterization of the T-cell receptor gamma locus and analysis of the variable gene segment expression in rabbit. Immunogenetics 57:352–363. doi:10.1007/s00251-005-0795-1
Chuang TH, Lai CY, Tseng PH, Yuan CJ, Hsu LC (2014) Development of CpG-oligodeoxynucleotides for effective activation of rabbit TLR9 mediated immune responses. PLoS One 9, e108808. doi:10.1371/journal.pone.0108808
Cohen C (1987) Erythrocyte antigens. In: Dubiski S (ed) The rabbit in contemporary immunological research. Longman Scientific &Technical and John Wiley & Sons, New York, pp pp 1–pp 12
Connelley TK, Degnan K, Longhi CW, Morrison WI (2014) Genomic analysis offers insights into the evolution of the bovine TRA/TRD locus. BMC Genomics 15:994. doi:10.1186/1471-2164-15-994
Corbet GB (1994) Taxonomy and origins. In: Tompson HV, King CM (eds) The European rabbit. Oxford Science, Oxford, pp 1–6
Cui S, Eisenacher K, Kirchhofer A, Brzozka K, Lammens A, Lammens K, Fujita T, Conzelmann KK, Krug A, Hopfner KP (2008) The C-terminal regulatory domain is the RNA 5'-triphosphate sensor of RIG-I. Mol Cell 29:169–179
Dalton KP, Nicieza I, Abrantes J, Esteves PJ, Parra F (2014) Spread of new variant RHDV in domestic rabbits on the Iberian Peninsula. Vet Microbiol 169:67–73. doi:10.1016/j.vetmic.2013.12.015
Dalton KP, Abrantes J, Lopes AM, Nicieza I, Álvarez ÁL, Esteves PJ, Parra F (2015) Complete genome sequence of two rabbit hemorrhagic disease virus variant b isolates detected on the Iberian Peninsula. Arch Virol 160:877–881. doi:10.1007/s00705-014-2329-3
Daniel TM (2015) Jean-Antoine Villemin and the infectious nature of tuberculosis. Int J Tuberc Lung Dis: 19:267–268
Dannenberg A M Jr (2006) Pathogenesis of human pulmonary tuberculosis: insights from the rabbit model. ASM Press, ISBN 1555813739, 9781555813734
Dawson M (1981). Evolution of the modern lagomorphs. In: Proceeding of the World Lagomorph Conference. K Myers and CD Mclnnes (Eds.), Univ. Guelph, Ontario, pp 1-16
de Matos AL, van der Loo W, Areal H, Lanning DK, Esteves PJ (2011) Study of Sylvilagus rabbit TRIM5alpha species-specific domain: how ancient endoviruses could have shaped the antiviral repertoire in Lagomorpha. BMC Evol Biol 11:294
de Matos AL, Lanning DK, Esteves PJ (2014) Genetic characterization of CCL3, CCL4 and CCL5 in leporid genera Oryctolagus. Sylvilagus Lepus Int J Immunogen 41:154–158. doi:10.1111/iji.12095
De Poorter M (1984) An experimental test of predictions from different hypothesis of self-regulation in the snowshoe hare (Lepus americanus Erxleben, 1777). Vrije University; Brussels.
Deeks ED, Keating DM (2009) Rabbit antithymocyte globulin (thymoglobulin): a review of its use in the prevention and treatment of acute renal allograft rejection. Drugs 69:1483–1512
Delibes-Mateos M, Delibes M, Ferreras P, Villafuerte R (2008) Key role of European rabbits in the conservation of the Western Mediterranean basin hotspot. Conserv Biol 22:1106–1117. doi:10.1111/j.1523-1739.2008.00993.x
Delibes-Mateos M, Ferreira C, Carro F, Escudero MA, Gortázar C (2014) Ecosystem effects of variant rabbit hemorrhagic disease virus, Iberian Peninsula. Emerg Infect Dis 20:2166–2168. doi:10.3201/eid2012.140517
DiMattia MA, Watts NR, Stahl SJ, Rader C, Wingfield PT, Stuart DI, Steven AC, Grimes JM (2010) Implications of the HIV-1 Rev dimer structure at 3.2 Å resolution for multimeric binding to the Rev response element. Proc Natl Acad Sci U S A 107:5810–5814
DiPietro LA, Short JA, Zhai SK, Kelus AS, Meier D, Knight KL (1990) Limited number of immunoglobulin VH regions expressed in the mutant rabbit “Alicia”. Eur J Immunol 20:1401–1404
Dixit E, Kagan JC (2013) Intracellular pathogen detection by RIG-I-like receptors, chapter 4. In Frederick WA (ed) Advances in immunology. Academic Press, p 99–118
Dorman SE, Hatem CL, Tyagi S, Aird K, Lopez-Molina J, Pitt ML, Zook BC, Dannenberg AM Jr, Bishai WR, Manabe YC (2004) Susceptibility to tuberculosis: clues from studies with inbred and outbred New Zealand White rabbits. Infect Immun 72:1700–1705
Dray S, Young GO (1959) Two antigenically different gamma-globulins in domestic rabbits revealed by isoprecipitins. Science 129:1023–1025
Dray S, Dubiski S, Kelus A, Lennox ES, Oudin J (1962) A notation for allotypy. Nature 195:785–7866
Dubiski S, Muller PJ (1967) A "new" allotypic specificity (A9) of rabbit immunoglobulin. Nature 214:696–697
Dubiski S, Dudziak Z, Skalba D, Dubiski A (1959) Serum groups in rabbits. Immunology 2:84
Duvoisin RM, Hayzer DJ, Belin D, Jaton J-C (1988) A rabbit Ig λ L chain C region gene encoding c21 allotypes. J Immunol 141:1596–1601
Emorine L, Max EE (1983) Structural analysis of a rabbit immunoglobulin κ 2 J-C locus reveals multiple deletions. Nucleic Acids Res 11:8888–8890
Emorine L, Sogn JA, Trinh D, Kindt TJ, Max EE (1984) A genomic gene encoding the b5 rabbit immunoglobulin kappa constant region: implications for latent allotype phenomenon.Proc Natl Acad Sci U S A 81:1789–1793
Esteves PJ, Alves PC, Ferrand N, van der Loo W (2002a) Restriction fragment alleles of the rabbit IGHG genes with reference to the rabbit IGHGCH2 or e locus polymorphism. Anim Genet 33:309–311
Esteves PJ, Alves PC, Ferrand N, van der Loo W (2002b) Hotspot variation at the CH2-CH3 interface of leporid IgG antibodies (Oryctolagus, Sylvilagus and Lepus). Eur J Immunogenet 29:529–535
Esteves PJ, Lanning D, Ferrand N, Knight KL, Zhai SK, van der Loo W (2004) Allelic variation at the VHa locus in natural populations of rabbit (Oryctolagus cuniculus, L.). J Immunol 172:1044–1053
Esteves PJ, Lanning D, Ferrand N, Knight KL, Zhai SK, van der Loo W (2005) The evolution of the immunoglobulin heavy chain variable region (IgV (H)) in Leporids: an unusual case of trans-species polymorphism. Immunogenetics 57:874–882
Esteves PJ, Carmo C, Godinho R, van der Loo W (2006) Genetic diversity at the hinge region of the unique immunoglobulin heavy gamma (IGHG) gene in leporids (Oryctolagus, Sylvilagus and Lepus). Int J Immunogenet 33:171–177
Esteves PJ, Abrantes J, Bertagnoli S, Cavadini P, Gavier-Widén D, Guitton J-S, Lavazza A, Lemaitre E, Letty J, Lopes AM, Neimanis AS, Ruvoën-Clouet N, Le Pendu J, Marchandeau S, Le Gall-Reculé G (2015) Emergence of pathogenicity in lagoviruses: evolution from pre-existing non-pathogenic strains or through a species jump? Plos pathogens (in press)
Fain MA, Zhao T, Kindt TJ (2001) Improved typing procedure for the polymorphic single-copy RLA-DQA gene of the rabbit reveals a new allele. Tissue Antigens 49:332–338
Felsenstein J (1981) Evolutionary trees and DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fenner F, Ratcliffe F (1965) Myxomatosis. Cambridge University Press, Cambridge, UK
Ferrand N, Branco M (2007) The evolutionary history of the European rabbit (Oryctolagus cuniculus): major patterns of population differentiation and geographic expansion inferred from protein polymorphism. In: Weiss S, Ferrand N (eds) Phylogeography of southern European refugia. Springer, Berlin Heidelberg, pp 207–235
Ferreira PG, Costa-e-Silva A, Oliveira MJ, Monteiro E, Cunha EM, Aguas AP (2006) Severe leukopenia and liver biochemistry changes in adult rabbits after calicivirus infection. Res Vet Sci 80:218–225
Ferreira PG, Dinis M, Costa ESA, Aguas AP (2008) Adult rabbits acquire resistance to lethal calicivirus infection by adoptive transfer of sera from infected young rabbits. Vet Immunol Immunopathol 121:364–369
Feschotte C, Gilbert C (2012) Endogenous viruses: insights into viral evolution and impact on host biology. Nat Rev Genet 13:283–296. doi:10.1038/nrg3199
Figueroa F, Gúnther E, Klein J (1988) MHC polymorphism pre-dating speciation. Nature 335:265–267
Fletcher AJ, Hue S, Schaller T, Pillay D, Towers GJ (2010) Hare TRIM5alpha restricts divergent retroviruses and exhibits significant sequence variation from closely related Lagomorpha TRIM5 genes. J Virol 84:12463–12468
Flisikowska T, Thorey IS, Offner S, Ros F, Lifke V, Zeitler B, Rottmann O, Vincent A, Zhang L, Jenkins S, Niersbach H, Kind AJ, Gregory PD, Schnieke AE, Platzer J (2011) Efficient immunoglobulin gene disruption and targeted replacement in rabbit using zinc finger nucleases. PLoS One 6, e21045. doi:10.1371/journal.pone.0021045
Flux JEC, Fullagar PJ (1992) World distribution of the Rabbit (Oryctolagus cuniculus) on islands. Mammal Rev 22:151–202
Foley JE, Nieto NC (2010) Tularemia. Vet Microbiol 140:332–338
Fontanesi L, Martelli PL, Scotti E, Russo V, Rogel-Gaillard C, Casadio R, Vernesi C (2012) Exploring copy number variation in the rabbit (Oryctolagus cuniculus) genome by array comparative genome hybridization. Genomics 100:245–251
Geraldes A, Rogel-Gaillard C, Ferrand N (2005) High levels of nucleotide diversity in the European rabbit (Oryctolagus cuniculus) SRY gene. Anim Genet 36:349–351
Gertz EM, Schäffer AA, Agarwala R, Bonnet-Garnier A, Rogel-Gaillard C, Hayes H, Mage RG (2013) Accuracy and coverage assessment of Oryctolagus cuniculus (rabbit) genes encoding immunoglobulins in the whole genome sequence assembly (OryCun 2.0) and localization of the IGH locus to chromosome 20. Immunogenetics 65:749–762
Gharekhani A, Entezari-Maleki T, Dashti-Khavidaki S, Khalili H (2013) A review on comparing two commonly used rabbit anti-thymocyte globulins as induction therapy in solid organ transplantation. Expert Opin Biol Ther 13:1299–1313
Gifford R, Tristem M (2003) The evolution, distribution and diversity of endogenous retroviruses. Virus Genes 26:291–315
Gilman-Sachs A, Mage RG, Young GO, Alexander C, Dray S (1969) Identification and genetic control of two rabbit immunoglobulin allotypes at a second light chain locus, the c locus. J Immunol 103:1159–1167
Glusman G et al (2001) Comparative genomics of the human and mouse T cell receptor loci. Immunity 15:337–349
Gridley J (1912) The lagomorphs and independent order. Science 36:285–286
Griffiths DJ, Voisset C, Venables PJ, Weiss RA (2002) Novel endogenous retrovirus in rabbits previously reported as human retrovirus 5. J Virol 76:7094–7102
Guillon P, Ruvoen-Clouet N, Le Moullac-Vaidye B, Marchandeau S, Le Pendu J (2009) Association between expression of the H histo-blood group antigen, alpha1,2fucosyltransferases polymorphism of wild rabbits, and sensitivity to rabbit hemorrhagic disease virus. Glycobiology 19:21–28
Gutierrez-Espeleta GA, Hedrick PW, Kalinowski ST, Garrigan D, Boyce WM (2001) Is the decline of desert bighorn sheep from infectious disease the result of low MHC variation? Heredity 86:439–450
Hamers R, Hamers-Casterman C (1965) Molecular localization of A chain allotypic specificities in rabbit IgG (7S gamma-globulin). J Mol Biol 14:288–289
Hamers-Casterman C, Wittouck E, van der Loo W, Hamers R (1979) Phylogeny of the rabbit gamma-chain determinants: a d12-like antigenic determinant in Pronolagus rupestris. J Immunogenet 6:373–381
Hatziioannou T, Perez-Caballero D, Yang A, Cowan S, Bieniasz PD (2004) Retrovirus resistance factors Ref1 and Lv1 are species-specific variants of TRIM5alpha. Proc Natl Acad Sci U S A 101:10774–10779
Hayday AC (2000) [gamma][delta] cells: a right time and a right place for a conserved third way of protection. Annu Rev Immunol 18:975–1026
Hayzer DJ, Young-Cooper GO, Mage RG, Jaton J-C (1990) cDNA clones encoding immunoglobulin lambda chains from rabbit expressing the phenotype c7. Eur J Immunol 20:2707–2712
Heil F et al (2004) Species-specific recognition of single-stranded RNA via toll-like receptor 7 and 8. Science 303:1526–1529
Hermel E, Han M, Hague B, Kindt T, Monaco JJ (1999) Isolation and mapping of the rabbit DM genes. Immunogenetics 49:295–302
Heyndrickx L, Stewart-Jones G, Jansson M, Schuitemaker H, Bowles E, Buonaguro L, Grevstad B, Vinner L, Vereecken K, Parker J, Ramaswamy M, Biswas P, Vanham G, Scarlatti G, Fomsgaard A, NGIN Consortium (2013) Selected HIV-1 Env trimeric formulations act as potent immunogens in a rabbit vaccination model. PLoS One 8, e74552. doi:10.1371/journal.pone.0074552
Hitoshi S, Kusunoki S, Kanazawa I, Tsuji S (1996) Molecular cloning and expression of a third type of rabbit GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferase. J Biol Chem 271:16975–16981
Horner DS, Lefkimmiatis K, Reyes A, Gissi C, Saccone C, Pesole G (2007) Phylogenetic analyses of complete mitochondrial genome sequences suggest a basal divergence of the enigmatic rodent Anomalurus. BMC Evol Biol 8:7–16
Horng WJ, Knight KL, Dray S (1976) Heavy chain variable region allotypic sub-specificities of rabbit immunoglobulins. I. Identification of three subpopulations of a1 IgG molecules. J Immunol 116:117–125
Hu J, Peng X, Schell TD, Budgeon LR, Cladel NM, Christensen ND (2006) An HLA-A2.1-transgenic rabbit model to study immunity to papillomavirus infection. J Immunol 177:8037–8045
Hu J, Peng X, Budgeon LR, Cladel NM, Balogh KK, Christensen ND (2007) Establishment of a cottontail/rabbit papillomavirus/HLA-A2.1 transgenic rabbit model. J Virol 81:7171–7177
Hu J, Budgeon LR, Balogh KK, Peng X, Cladel NM, Christensen ND (2014) Long-peptide therapeutic vaccination against CRPV-induced papillomas in HLA-A2.1 transgenic rabbit. Trials Vaccinol 3:134–142
Isono T, Kim CJ, Seto A (1995) Sequence and diversity of rabbit T-cell receptor gamma chain genes. Immunogenetics 41:295–300
Iwamori M, Domino SE (2004) Tissue-specific loss of fucosylated glycolipids in mice with targeted deletion of alpha(1,2)fucosyltransferase genes. Biochem J 380:75–81. doi:10.1042/BJ20031668
James JA, Gross T, Scofield RH, Harley JB (1995) Immunoglobulin epitope spreading and autoimmune disease after peptide immunization: Sm B/B'-derived PPPGMRPP and PPPGIRGP induce spliceosome autoimmunity. J Exp Med 181:453–461
Jurk M et al (2002) Human TLR7 or TLR8 independently confer responsiveness to the antiviral compound R-848. Nat Immunol 3:499. doi:10.1038/ni0602-499
Kajikawa O et al (2005) Gene expression of Toll-like receptor-2, Toll-like receptor-4, and MD2 is differentially regulated in rabbits with Escherichia coli pneumonia. Gene 344:193–202. doi:10.1016/j.gene.2004.09.032
Kang JY, Lee JO (2011) Structural biology of the Toll-like receptor family. Annu Rev Biochem 80:917–941. doi:10.1146/annurev-biochem-052909-141507
Kato H, Takahasi K, Fujita T (2011) RIG-I-like receptors: cytoplasmic sensors for non-self RNA. Immunol Rev 243:91–98
Katzourakis A, Tristem M, Pybus OG, Gifford RJ (2007) Discovery and analysis of the first endogenous lentivirus. Proc Natl Acad Sci U S A 104(15):6261–6265
Kawai T, Akira S (2006) Innate immune recognition of viral infection. Nat Immunol 7:131–137
Kawamura S, Saitou N, Ueda S (1992) Concerted evolution of the primate immunoglobulin alpha-gene through gene conversion. J Biol Chem 267:7359–7367
Keckesova Z, Ylinen LM, Towers GJ, Gifford RJ, Katzourakis A (2009) Identification of a RELIK orthologue in the European hare (Lepus europaeus) reveals a minimum age of 12 million years for the lagomorph lentiviruses. Virology 384:7–11. doi:10.1016/j.virol.2008.10.045
Kelus AS, Gell PG (1967) Immunoglobulin allotypes of experimental animals. Prog Allergy 11:141–184
Kelus AS, Weiss S (1977) Variant strain of rabbits lacking immunoglobulin κ polypeptide chain. Nature 265:156–158
Kelus AS, S Weiss (1986) Mutation affecting the expression of immunoglobulin variable regions in the rabbit. Proc Natl Acad Sci USA 83:4883–4886
Kerr PJ (2012) Myxomatosis in Australia and Europe: a model for emerging infectious diseases. Antiviral Res 93:387–415
Kerr P, McFadden G (2002) Immune responses to myxoma virus. Viral Immunol 15:229–246
Kerr PJ, Kitchen A, Holmes EC (2009) Origin and phylodynamics of rabbit hemorrhagic disease virus. J Virol 83:12129–12138
Kerr P, Liu J, Cattadori I, Ghedin E, Read A, Holmes E (2015) Myxoma virus and the Leporipoxviruses: an evolutionary paradigm. Viruses 7:1020–1061
Kerremans Ph (1982) Studie van interlokaliteits varianties in pecking order van de uitdrukking van immunoglobuline allotypen. Licentiaats Thesis, Vrije Universiteit Brussel, Brussels
Kim BS, Dray S (1972) Identification and genetic control of allotypic specificities on two variable region subgroups of rabbit immunoglobulin heavy chains. Eur J Immunol 2:509–514
Kim BS, Dray S (1973) Expression of the a, x, and y variable region genes of heavy chains among IgG, IgM, and IgA molecules of normal and a locus allotype-suppressed rabbits. J Immunol 111:750–760
Kindt TJ (1975) Rabbit immunoglobulin allotypes: structure, immunology, and genetics. Adv Immunol 21:35–86
Klein J, Sato A, Nagl S, O'hUigín C (1998) Molecular trans-species polymorphism. Annu Rev Ecol Syst 29:1–21
Knight KL (1992) Restricted VH gene usage and generation of antibody diversity in rabbit. Annu Rev Immunol 10:593–616
Knight KL, Becker RS (1990) Molecular basis of the allelic inheritance of rabbit immunoglobulin VH allotypes: implications for the generation of antibody diversity. Cell 60:963–970
Knight KL, Burnett RC, McNicholas JM (1985) Organization and polymorphism of rabbit immunoglobulin heavy chain genes. J Immunol 134:1245–1250
Koblansky AA et al (2013) Recognition of profilin by Toll-like receptor 12 is critical for host resistance to Toxoplasma gondii immunity. J Immunol 38:119–130. doi:10.1016/j.immuni.2012.09.016
Komatsu M (1987) Polymorphisms and deficiencies of complement components. In: Dubiski S (ed) The rabbit in contemporary immunological research. Longman Scientific Technical and John Wiley& Son, New York, pp 191–206
Kummerfeldt CE (2014) Raxibacumab: potential role in the treatment of inhalational anthrax. Infect Drug Resist 7:101–109
Lai CY et al (2014) TLR7/8 agonists activate a mild immune response in rabbits through TLR8 but not TLR7. Vaccine 32:5593–5599. doi:10.1016/j.vaccine.2014.07.104
Lamoyi E, Mage RG (1985) Lack of K1b9 light chains in Basilea rabbits is probably due to a mutation in an acceptor site for mRNA splicing. J Exp Med 162:1149–1160
Lamoyi E, Mage R (1987) A cluster of rabbit T-cell beta-chain variable region genes. Immunogenetics 25:55–62
Lande R (1994) Risk of population extinction from fixation of new deleterious mutations. Evolution 48:1460–1469
Landsteiner K (1900) Zur Kenntnis der antifermentativen, lytischen und agglutinierenden Wirkungen des Blutserums und der Lymphe. Zentralbl Bakteriol 27:357–362
Landucci Tosi S, Mage RG, Tosi R (1976) The homologues of rabbit b-allotypes on hare immunoglobulin light chains: an evolutionary puzzle. J Immunol 117:679–685
Landucci Tosi S, Mage RG, Tosi RM (l976b) A homologue of the rabbit κ-chain allotype b9 on IgG from a cottontail rabbit. J Immunol 117:686–689
Lanier HC, Olson LE (2009) Inferring divergence times within pikas (Ochotona spp.) using mtDNA and relaxed molecular dating techniques. Mol Phylogenet Evol 53:1–12
Lanning DK, Zhai SK, Knight KL (2003) Analysis of the 3' Cmu region of the rabbit Ig heavy chain locus. Gene 309:135–144
Le Gall-Recule G et al (2011) Detection of a new variant of rabbit haemorrhagic disease virus in France. Vet Rec 168:137–138. doi:10.1136/vr.d697
Le Pendu J, Nystrom K, Ruvoen-Clouet N (2014) Host-pathogen co-evolution and glycan interactions. Curr Opin Virol 7:88–94. doi:10.1016/j.coviro.2014.06.001
Lee SM et al (2014) Toll-like receptor 10 is involved in induction of innate immune responses to influenza virus infection. Proc Natl Acad Sci U S A 111:3793–3798. doi:10.1073/pnas.1324266111
Lees AC, Bell DJ (2008) A conservation paradox for the 21st century: the European wild rabbit Oryctolagus cuniculus, an invasive alien and an endangered native species. Mammal Rev 38:304–320
Lefranc MP, Pommie C, Kaas Q, Duprat E, Bosc N, Guiraudou D, Jean C, Ruiz M, Da Piedade I, Rouard M, Foulquier E, Thouvenin V, Lefranc G (2005) IMGT unique numbering for immunoglobulin and T cell receptor constant domains and Ig superfamily C-like domains. Dev Comp Immunol 29:185–203
Lemos de Matos A, Sousa-Pereira P, Lissovsky AA, van der Loo W, Melo-Ferreira J, Cui J, Esteves PJ (2015) Endogenization of mouse mammary tumor virus (MMTV)-like elements in genomes of pikas (Ochotona sp.). Virus Res 210:22–26. doi:10.1016/j.virusres.2015.06.021
Lemos de Matos A, McFadden G, Esteves PJ (2013) Positive evolutionary selection on the RIG-I-like receptor genes in mammals. PLoS One 8, e81864
Lemos de Matos A, McFadden G, Esteves PJ (2014) Evolution of viral sensing RIG-I-like receptor genes in Leporidae genera Oryctolagus, Sylvilagus, and Lepus. Immunogenetics 66:43–52
Li XD, Chen ZJ (2012) Sequence specific detection of bacterial 23S ribosomal RNA by TLR13. eLife 1:e00102. doi:10.7554/eLife.00102
Li Y, Li X, Stremlau M, Lee M, Sodroski J (2006) Removal of arginine 332 allows human TRIM5alpha to bind human immunodeficiency virus capsids and to restrict infection. J Virol 80:6738–6744
Lissovsky A (2014) Taxonomic revision of pikas Ochotona (Lagomorpha, Mammalia) at the species level. Mammalia 78:199–216
Liu J, Wennier S, McFadden G (2010) The immunoregulatory properties of oncolytic myxoma virus and their implications in therapeutics. Microbes Infect 12:1144–1152
Liu J et al (2012) Activation of rabbit TLR9 by different CpG-ODN optimized for mouse and human TLR9. Comp Immunol Microbiol Infect Dis 35:443–451. doi:10.1016/j.cimid.2012.03.008
Lopes AM, Correia J, Abrantes J, Melo P, Ramada M, Magalhães MJ, Alves PC, Esteves PJ (2014) Is the new variant RHDV replacing genogroup 1 in Portuguese wild rabbit populations? Viruses 7:27–36. doi:10.3390/v7010027
Lopes AM, Dalton KP, Magalhaes MJ, Parra F, Esteves PJ, Holmes EC, Abrantes J (2015) Full genomic analysis of new variant Rabbit Hemorrhagic Disease Virus (RHDVb) revealed multiple recombination events. J Gen Virol 96:1309–1319. doi:10.1099/vir.0.000070
Lopez-Martinez N (1989) Revision sistemática y biostratigrafca de los lagomorphos (Mammalia) del Terciário y Cuaternario de Espana. Memorias del Museo Paleontologico de la Universidad de Zaragoza, n°3. Diputacion General de Aragon.
Lummus Z, Cebra JJ, Mage R (1967) Correspondence of the relative cellular distribution and serum concentration of allelic allotypic markers in normal and “allotype-suppressed” heterozygous rabbits. J Immunol 99:737–743
Lurie MB (1928) The fate of human and bovine tubercle bacilli in various organs of the rabbit. J Exp Med 48:155–182
Magalhães V, Abrantes J, Munõz-Pajares AJ, Esteves PJ (2015) Genetic diversity comparison of the DQA gene in European rabbit (Oryctolagus cuniculus) populations. Immunogenetics doi:10.1007/s00251-015-0866-x
Mage RG, Mage MG (2012) Sequence of rabbit (Oryctolagus cuniculus) DNA from the OryCun 2.0 donor does not confirm a frameshift in exon 2 of IL4. Immunol Immunogenet Insights 2012:1–5. doi:10.4137/III.S9557
Mage RG, Rai G (2012) A rabbit model of systemic lupus erythematosus, useful for studies of neuropsychiatric SLE. In: Systemic lupus erythematosus. InTech-Rijeka, Croatia. ISBN: 978-953-307-868-7
Mage RG, Rogel-Gaillard (2016) Immunogenetics in the rabbit. In: Fontanesi F (ed) The genetics and genomics of the rabbit. CABI, Wallingford, Oxfordshire, UK, Chapter 8
Mage RG, Young GO, Reisfeld RA (1968) The association of the c7 allotype of rabbits with some light polypeptide chains which lack b locus allotype. J Immunol l0l:6l7–6l620
Mage RG, Young-Cooper GO, Alexander C (1971) Genetic control of variable and constant regions of immunoglobulin H chains. Nat New Biol 230:63–64
Mage RG, Bernstein KE, McCartney-Francis N, Alexander CB, Young-Cooper GO, Padlan EA, Cohen GH (1984) The structural and genetic basis for expression of normal and latent VHa allotypes of the rabbit. Mol Immunol 21:1067–1081
Mage RG, Lanning D, Knight KL (2006) B cell and antibody repertoire development in rabbits: the requirement of gut-associated lymphoid tissues. Dev Comp Immunol 30:137–153
Manabe YC, Dannenberg AM Jr, Tyagi SK, Hatem CL, Yoder M, Woolwine SC, Zook BC, Pitt ML, Bishai WR (2003) Different strains of Mycobacterium tuberculosis cause various spectrums of disease in the rabbit model of tuberculosis. Infect Immun 71:6004–6011
Marchandeau S, Chantal J, Portejoie Y, Barraud S, Chaval Y (1998) Impact of viral haemorrhagic disease on a wild population of European rabbits in France. J Wildlife Dis 34:429–435
Marchandeau S, Chaval Y, Le Goff E (2000) Prolonged decline in the abundance of wild European rabbits Oryctolagus cuniculus and high immunity level over three years following the arrival of rabbit haemorrhagic disease. Wildlife Biol 6:141–147
Marche PN, Tykocinski ML, Max EE, Kindt TJ (1985) Structure of a functional rabbit class I MHC gene: similarity to human class I genes. Immunogenetics 21:71–82
Margolies MN, Cannon LE, Kindt TJ, Fraser B (1977) The structural basis of rabbit VH allotypes: serologic studies on a1 H chains with defined amino acid sequence. J Immunol 119:287–294
Margulies EH et al (2007) Analyses of deep mammalian sequence alignments and constraint predictions for 1% of the human genome. Genome Res 17:760–774. doi:10.1101/gr.6034307
Marques RM, Costa ESA, Aguas AP, Teixeira L, Ferreira PG (2012) Early inflammatory response of young rabbits attending natural resistance to calicivirus (RHDV) infection. Vet Immunol Immunopathol 150:181–188. doi:10.1016/j.vetimm.2012.09.038
Marshall ID, Regnery DC (1960a) Myxomatosis in a California brush rabbit (Sylyilagus bachmani). Nature 188:73–74
Marshall ID, Regnery DC (1960b) Myxomatosis in a California brush rabbit (Sylyilagus bachmani). Nature 188:73–74
Martens CL, Currier SJ, Knight KL (1984) Molecular genetic analysis of genes encoding the heavy chains of rabbit IgG. J Immunol 133:1022–1027
Massari S, Ciccarese S, Antonacci R (2012) Structural and comparative analysis of the T cell receptor gamma (TRG) locus in Oryctolagus cuniculus. Immunogenetics 64:773–779. doi:10.1007/s00251-012-0634-0
Matthee CA, van Vuuren BJ, Bell D, Robinson TJ (2004) A molecular supermatrix of the rabbits and hares (Leporidae) allows for the identification of five intercontinental exchanges during the Miocene. Syst Biol 53:433–447
Matthyssens G, Borremans B, Van Der Loo W & Hamers R (1985) Genomic evidence for the truly allelic nature of the b locus polymorphism of the rabbit immunoglobulin light chain. In: Protides of the biological fluids: Proceedings of the Thirty-Third Colloquium Vol. 33, p. 123-125 Elsevier
McCartney-Francis (1987) Lattent allotype expression in the rabbit. In Dubiski S (ed) The rabbit in contemporary immunological research. Longman Scientific & Technical, p 134
McCartney-Francis N, Skurla RM Jr, Mage RG, Bernstein KE (1984) Kappa-chain allotypes and isotypes in the rabbit: cDNA sequences of clones encoding b9 suggest an evolutionary pathway and possible role of the interdomain disulfide bond in quantitative allotype expression. Proc Natl Acad Sci U S A 81:1794–1798
Medzhitov R (2001) Toll-like receptors and innate immunity. Nat Rev Immunol 1:135–145. doi:10.1038/35100529
Melo-Ferreira J, Boursot P, Carneiro M, Esteves PJ, Farelo L, Alves PC (2012) Recurrent introgression of mitochondrial DNA among hares (Lepus spp.) revealed by species-tree inference and coalescent simulations. Syst Biol 61:367–381
Melo-Ferreira J, Lemos de Matos A, Areal H, Lissovsky AA, Carneiro M, Esteves PJ (2015) The phylogeny of pikas (Ochotona) inferred from a multilocus coalescent approach. Mol Phylogenet Evol 84:240–244
Mendez S, Hatem CL, Kesavan AK, Lopez-Molina J, Pitt ML, Dannenberg AM Jr, Manabe YC (2008) Susceptibility to tuberculosis: composition of tuberculous granulomas in Thorbecke and outbred New Zealand White rabbits. Vet Immunol Immunopathol 122:167–174
Mikami O, Park JH, Kimura T, Ochiai K, Itakura C (1999) Hepatic lesions in young rabbits experimentally infected with rabbit haemorrhagic disease virus. Res Vet Sci 66:237–242
Mitro S, Krauss H (1993) Rabbit hemorrhagic disease: a review with special reference to its epizootiology. Eur J Epidemiol 9:70–78
Monnerot M, Vigne JD, Biju-Duval C, Casane D, Cailou C, Hardy C, Mougel F, Soriguer R, Dennebouy N, Mounolou JC (1994) Rabbit and man: genetic and historic approach. Genet Sel Evol 26(suppl 1):167–182
Muller A et al (2009) Evolution of rabbit haemorrhagic disease virus (RHDV) in the European rabbit (Oryctolagus cuniculus) from the Iberian Peninsula. Vet Microbiol 135:368–373
Murphy W, Eizirik E, O'Brien SJ, Madsen O, Scally M, Douday CJ, Teeling E, Ryder OA, Stanhope MJ, de Jong WW, Springer MS (2001) Resolution of the early placental mammal radiation using Bayesian phylogenetics. Science 294:2348–2351
Nakayama EE, Miyoshi H, Nagai Y, Shioda T (2005) A specific region of 37 amino acid residues in the SPRY (B30.2) domain of African green monkey TRIM5alpha determines species-specific restriction of simian immunodeficiency virus SIVmac infection. J Virol 79:8870–8877
Neves F, Abrantes J, Pinheiro A, Almeida T, Costa PP, Esteves PJ (2014a) Convergent evolution of IL-6 in two leporids (Oryctolagus and Pentalagus) originated an extended protein. Immunogenetics 66:589–595. doi:10.1007/s00251-014-0787-0
Neves F, Abrantes J, Steinke JW, Esteves PJ (2014b) Maximum-likelihood approaches reveal signatures of positive selection in IL genes in mammals. Innate Immun 20:184–191
Neves F, Abrantes J, Almeida T, Lemos de Matos A, Costa PP, Esteves PJ (2015a) Genetic characterization of interleukins (IL1α, IL1β, IL2, IL4, IL8, IL10, IL12A, IL12B, IL15 and IL18) with relevant biological roles in lagomorphs. Innate Immunity (in press)
Neves F, Abrantes J, Lissovsky AA, Esteves PJ (2015b) Pseudogenization of CCL14 in the Ochotonidae (pika) family. Innate Immun 21:647–654. doi:10.1177/1753425915577455
Newman D, Pilson D (1997) Increased probability of extinction due to decreased genetic effective population size: experimental populations of Clarkia pulchella. Evolution 51:345–362
Nisole S, Stoye JP, Saib A (2005) TRIM family proteins: retroviral restriction and antiviral defence. Nat Rev Immunol 3:799–808
Nyström K et al (2011) Histo-blood group antigens act as attachment factors of rabbit hemorrhagic disease virus infection in a virus strain-dependent manner. PLoS Pathog 7, e100218
Nystrom K et al (2015) Neofunctionalization of the Sec1 alpha1,2fucosyltransferase paralogue in leporids contributes to glycan polymorphism and resistance to rabbit hemorrhagic disease virus. PLoS Pathog 11, e1004759. doi:10.1371/journal.ppat.1004759
O'Brien E, Kerber RA, Jorde LB, Rogers AR (1994) Founder effect: assessment of variation in genetic contributions among founders. Human Biol 66:185–204
Ohto U, Tanji H, Shimizu T (2014) Structure and function of toll-like receptor 8. Microbes Infect 16:273–282. doi:10.1016/j.micinf.2014.01.007
Oldenburg M et al (2012) TLR13 recognizes bacterial 23S rRNA devoid of erythromycin resistance-forming modification. Science 337:1111–1115. doi:10.1126/science.1220363
Oliver M, Piertney S (2010) Beyond splitting hares and rabbiting on about major histocompatibility complex complexity. Mol Ecol 19:4099–4101
Oosting M et al (2014) Human TLR10 is an anti-inflammatory pattern-recognition receptor. Proc Natl Acad Sci U S A 111:E4478–E4484. doi:10.1073/pnas.1410293111
Oppelt C, Starkloff A, Rausch P, VON Holst D, Rödel HG (2010) Major histocompatibility complex variation and age-specific endoparasite load in subadult European rabbits. Mol Ecol 19:4155–4167
Oudin J (1956) L'"alloiypie" de certains antigènes proteídique du sérum. Comp Rend Acad Sci Paris 242:2606–2608
Oudin J (1960) Allotypy of rabbit serum proteins I. Immunochemical analysis leading to the individualization of seven main allotypes. J Exp Med 112:107–124
Ozato K, Shin DM, Chang TH, Morse HC 3rd (2008) TRIM family proteins and their emerging roles in innate immunity. Nat Rev Immunol 8:849–860
Pan R, Sampson JM, Chen Y, Vaine M, Wang S, Lu S, Kong XP (2013) Rabbit anti-HIV-1 monoclonal antibodies raised by immunization can mimic the antigen-binding modes of antibodies derived from HIV-1-infected humans. J Virol 87:10221–10231
Parra ZE, Baker ML, Hathaway J, Lopez AM, Trujillo J, Sharp A, Miller RD (2008) Comparative genomic analysis and evolution of the T cell receptor loci in the opossum Monodelphis domestica. BMC Genomics 9:111. doi:10.1186/1471-2164-9-111
Parry GD (1981) The meanings of r- and K-selection. Oecologia (Berl) 48:260-264
Peng X, Roshwalb S, Cooper TK, Zimmerman H, Christensen ND (2015) High incidence of spontaneous cataracts in aging laboratory rabbits of an inbred strain. Vet Ophthalmol 18:186–190. doi:10.1111/vop.12203
Perkins HD, van Leeuwen BH, Hardy CM, Kerr PJ (2000) The complete cDNA sequences of IL-2, IL-4, IL-6 AND IL-10 from the European rabbit (Oryctolagus cuniculus). Cytokine+ 12:555–565. doi:10.1006/cyto.1999.0658
Perron MJ, Stremlau M, Song B, Ulm W, Mulligan RC, Sodroski J (2004) TRIM5alpha mediates the postentry block to N-tropic murine leukemia viruses in human cells. Proc Natl Acad Sci U S A 101:11827–11832
Pinheiro A, Lanning D, Alves PC, Mage RG, Knight KL, van der Loo W, Esteves PJ (2011) Molecular bases of genetic diversity and evolution of the immunoglobulin heavy chain variable region (IGHV) gene locus in leporids. Immunogenetics 63:397–408
Pinheiro A, de Mera IG, Alves PC, Gortázar C, de la Fuente J, Esteves PJ (2013a) Sequencing of modern Lepus VDJ genes shows that the usage of VHn genes has been retained in both Oryctolagus and Lepus that diverged 12 million years ago. Immunogenetics 65:777–784
Pinheiro A, Woof JM, Abi-Rached L, Parham P, Esteves PJ (2013b) Computational analyses of an evolutionary arms race between mammalian immunity mediated by immunoglobulin A and its subversion by bacterial pathogens. Plos One 8, e73934
Pinheiro A, Melo-Ferreira J, Abrantes J, Martinelli N, Lavazza A, Alves PC, Gortázar C, Esteves PJ (2014a) Sequencing of Sylvilagus VDJ genes reveals a new VHa allelic lineage and shows that ancient VH lineages were retained differently in leporids. Immunogenetics 66:719–726
Pinheiro A, Woof JM, Almeida T, Abrantes J, Alves PC, Gortázar C, Esteves PJ (2014b) Leporid immunoglobulin G shows evidence of strong selective pressure on the hinge and CH3 domains. Open Biol 4:140088
Pinheiro A, Almeida T, Esteves PJ (2015) Survey of genetic diversity in wild and domestic rabbits for IgG. Int J Immunogenet 42(5):364–36 doi:10.1111/iji.12222
Popkov M, Mage RG, Alexander CB, Thundivalappil S, Barbas CF 3rd, Rader C (2003) Rabbit immune repertoires as sources for therapeutic monoclonal antibodies: the impact of kappa allotype-correlated variation in cysteine content on antibody libraries selected by phage display. J Mol Biol 325:325–335
Popkov M, Jendreyko N, Gonzalez-Sapienza G, Mage RG, Rader C, Barbas CF 3rd (2004) Human/mouse cross-reactive anti-VEGF receptor 2 recombinant antibodies selected from an immune b9 allotype rabbit antibody library. J Immunol Methods 288:149–164
Porcelli SA, Modlin R (1999) THE CD1 SYSTEM: antigen-presenting molecules for T cell recognition of lipids and glycolipids. Annu Rev Immunol 17:297–329
Pospisil R, Young-Cooper GO, Mage RG (1995) Preferential expansion and survival of B lymphocytes based on VH framework 1 and framework 3 expression: “positive” selection in appendix of normal and VH-mutant rabbits. Proc Natl Acad Sci U S A 92:6961–6965
Potts WK, Manning CJ, Wakeland EK (1991) Mating patterns in seminatural populations of mice influenced by MHC genotype. Nature 352:619–621
Poulsen K, Fraser KJ, Haber E (1972) An active derivative of rabbit antibody light chain composed of the constant and the variable domains held together only by a native disulfide bond. Proc Natl Acad Sci U S A 69:2495–2409
Puggioni G et al (2013) The new French 2010 rabbit hemorrhagic disease virus causes an RHD-like disease in the Sardinian Cape hare (Lepus capensis mediterraneus). Vet Res 44:96. doi:10.1186/1297-9716-44-96
Qin Y, Banerjee S, Agrawal A, Shi H, Banasik M, Lin F, Rohl K, LaBranche C, Montefiori DC, Cho MW (2015) Characterization of a large panel of rabbit monoclonal antibodies against HIV-1 gp120 and isolation of novel neutralizing antibodies against the V3 loop. PLoS One 10(6), e0128823
Queney G, Ferrand N, Weiss S, Mougel F, Monnerot M (2001) Stationary distributions of microsatellite loci between divergent population groups of the European rabbit (Oryctolagus cuniculus). Mol Biol Evol 18(12):2169–2178
Rader C (2009) Generation and selection of rabbit antibody libraries by phage display. Methods Mol Biol 525:101–128
Ratcliffe FN, Myers K, Fennessy BV, Calaby JH (1952) Myxomatosis in Australia: a step towards the biological control of the rabbit. Nature 170:7–11
Regan T, Nally K, Carmody R, Houston A, Shanahan F, Macsharry J, Brint E (2013) Identification of TLR10 as a key mediator of the inflammatory response to Listeria monocytogenes in intestinal epithelial cells and macrophages. J Immunol 191:6084–6092. doi:10.4049/jimmunol.1203245
Roach JC et al (2005) The evolution of vertebrate Toll-like receptors. Proc Natl Acad Sci U S A 102:9577–9582. doi:10.1073/pnas.0502272102
Rogel-Gaillard C, Piumi F, Billault A, Bourgeaux N, Save JC et al (2001) Construction of a rabbit bacterial artificial chromosome (BAC) library: application to the mapping of the major histocompatibility complex to position 12q1.1. Mamm Genome 12:253–255
Ros F, Puels J, Reichenberger N, van Schooten W, Buelow R, Platzer J (2004) Sequence analysis of 0.5 Mb of the rabbit germline immunoglobulin heavy chain locus. Gene 33:49–59
Roux KH (1981) A fourth heavy chain variable region subgroup, w, with 2 variants defined by an induced auto-antiserum in the rabbit. J Immunol 127:626–632
Ruvoen-Clouet N, Blanchard D, Andre-Fontaine G, Ganiere JP (1995) Partial characterization of the human erythrocyte receptor for rabbit haemorrhagic disease virus. Res Virol 146:33–41
Ruvoen-Clouet N, Ganiere JP, Andre-Fontaine G, Blanchard D, Le Pendu J (2000) Binding of rabbit hemorrhagic disease virus to antigens of the ABH histo-blood group family. J Virol 74:11950–11954
Saccheri I, Kuussaari M, Vikman W, Fortellius W, Hanski I (1998) Inbreeding and extinction in a metapopulation. Nature 392:491–494
Salazar JC, Rathi A, Michael NL, Radolf JD, Jagodzinski LL (2007) Assessment of the kinetics of Treponema pallidum dissemination into blood and tissues in experimental syphilis by real-time quantitative PCR. Infect Immun 75:2954–2958
Sanders RW, van Gils MJ, Derking R, Sok D, Ketas TJ, Burger JA, Ozorowski G, Cupo A, Simonich C, Goo L, Arendt H, Kim HJ, Lee JH, Pugach P, Williams M, Debnath G, Moldt B, van Breemen MJ, Isik G, Medina-Ramírez M, Back JW, Koff WC, Julien JP, Rakasz EG, Seaman MS, Guttman M, Lee KK, Klasse PJ, LaBranche C, Schief WR, Wilson IA, Overbaugh J, Burton DR, Ward AB, Montefiori DC, Dean H, Moore JP (2015) HIV-1 neutralizing antibodies induced by native-like envelope trimmers. Science 349(6244):aac4223. doi:10.1126/science.aac4223
Schaller T, Hue S, Towers GJ (2007) An active TRIM5 protein in rabbits indicates a common antiviral ancestor for mammalian TRIM5 proteins. J Virol 81:11713–11721
Sehgal D, Mage RG, Schiaffella E (1998) VH mutant rabbits lacking the VH1a2 gene develop a2+ B cells in the appendix by gene conversion-like alteration of a rearranged VH4 gene. J Immunol 160:1246–1255
Shibata K, Nomiyama H, Yoshie O, Tanase S (2013) Genome diversification mechanism of rodent and Lagomorpha chemokine genes. BioMed Res Int 2013:856265. doi:10.1155/2013/856265
Siewe BT, Kalis SL, Esteves PJ, Zhou T, Knight KL (2010) A novel functional rabbit IL-7 isoform. Dev Comp Immunol 34:828–836. doi:10.1016/j.dci.2010.03.003
Sjöstedt A (2011) Special topic on Francisella tularensis and tularemia. Front Microbiol 2:86. doi:10.3389/fmicb.2011.00086
Smee DF (2013) Orthopoxvirus inhibitors that are active in animal models: an update from 2008 to 2012. Future Virol 8:891–901
Smith S, Mang T, De Bellocq JG, Schaschl H, Zeitlhofer C, Hackländer K, Suchentrunk F (2010) Homozygosity at a class II MHC locus depresses female reproductive ability in European brown hares. Mol Ecol 19:4131–4143
Song B, Javanbakht H, Perron M, Park DH, Stremlau M, Sodroski J (2005) Retrovirus restriction by TRIM5alpha variants from Old World and New World primates. J Virol 79:3930–3937
Sperry AO, Blasquez VC, Garrard WT (1989) Dysfunction of chromosomal loop attachment sites: illegitimate recombination linked to matrix association regions and topoisomerase II. Proc Natl Acad Sci U S A 86:5497–5501
Spieker-Polet H, Yam PC, Knight KL (1993) Differential expression of 13 IgA heavy chain genes in rabbit lymphoid tissues. J Immunol 150:5457–5465
Spiesschaert B, McFadden G, Hermans K, Nauwynck H, Van de Walle GR (2011) The current status and future directions of myxoma virus, a master in immune evasion. Vet Res 42:76
Springer MS, Murphy WJ, Eizirik E, O'Brien SJ (2003) Placental mammal diversification and the Cretaceous-Tertiary boundary. Proc Natl Acad Sci U S A 100:1056–1061
Stahl SJ, Watts NR, Rader C, DiMattia MA, Mage RG, Palmer I, Kaufman JD, Grimes JM, Stuart DI, Steven AC, Wingfield PT (2010) Generation and characterization of a chimeric rabbit/human Fab for co-crystallization of HIV-1 Rev. J Mol Biol 397:697–708
Stanford MM, Werden SJ, McFadden G (2007) Myxoma virus in the European rabbit: interactions between the virus and its susceptible host. Vet Res 38:299–318
Storek J, Mohty M, Boelens JJ (2015) Rabbit anti-T cell globulin in allogeneic hematopoietic cell transplantation. Biol Blood Marrow Transplant 21:959–970
Stoye JP (2012) Studies of endogenous retroviruses reveal a continuing evolutionary saga. Nat Rev Microbiol 10:395–406. doi:10.1038/nrmicro2783
Stremlau M, Perron M, Welikala S, Sodroski J (2005) Species-specific variation in the B30.2 (SPRY) domain of TRIM5alpha determines the potency of human immunodeficiency virus restriction. J Virol 79:3139–3145
Su C, Nei M (1999) Fifty-million-year-old polymorphism at an immunoglobulin variable region gene locus in the rabbit evolutionary lineage. Proc Natl Acad Sci U S A 96:9710–9715
Subbian S, O'Brien P, Kushner NL, Yang G, Tsenova L, Peixoto B, Bandyopadhyay N, Bader JS, Karakousis PC, Fallows D, Kaplan G (2013a) Molecular immunologic correlates of spontaneous latency in a rabbit model of pulmonary tuberculosis. Cell Commun Signal 11:16. doi:10.1186/1478-811X-11-16
Subbian S, Bandyopadhyay N, Tsenova L, O'Brien P, Khetani V, Kushner NL, Peixoto B, Soteropoulos P, Bader JS, Karakousis PC, Fallows D, Kaplan G (2013b) Early innate immunity determines outcome of Mycobacterium tuberculosis pulmonary infection in rabbits. Cell Commun Signal 11:60. doi:10.1186/1478-811X-11-60
Surridge AK, van der Loo W, Abrantes J, Carneiro M, Hewitt GM, Esteves PJ (2008) Diversity and evolutionary history of the MHC DQA gene in leporids. Immunogenetics 60:515–525
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tervo HM, Keppler O (2010) High natural permissivity of primary rabbit cells for HIV-1, with a virion infectivity defect in macrophages as the final replication barrier. J Virol 84:12300–12314
Thomas LD, Schaffner W (2010) Tularemia pneumonia. Infect Dis Clin North Am 24:43–55
Traxlmayr MW, Obinger C (2012) Directed evolution of proteins for increased stability and expression using yeast display. Arch Biochem Biophys 526:174–180
Tsenova L, O'Brien P, Holloway J, Peixoto B, Soteropoulos P, Fallows D, Kaplan G, Subbian S (2014) Etanercept exacerbates inflammation and pathology in a rabbit model of active pulmonary tuberculosis. J Interferon Cytokine Res 34:716–726
van der Loo W (1993) Variance analysis of immunoglobulin alleles in natural populations of rabbit (Oryctolagus cuniculus): the extensive interallelic divergence at the b locus could be the outcome of overdominance-type selection. Genetics 135:171–187
van der Loo W, Hamers-Casterman C (1979) Phylogeny of the rabbit immunoglobulin allotypes: rabbit anti e-locus antisera detect two allelic forms of IgG molecules in Romerolagus diazi. Ann Inst Pasteur Immunol C 130:761
van der Loo W, Verdoodt B (1992) Patterns of interallelic divergence at the rabbit b-locus of the immunoglobulin light chain constant region are in agreement with population genetic evidence for overdominant selection. Genetics 132:1105–1117
van der Loo W, De Baetselier P, Hamers-Casterman C, Hamers R (1977) Evidence for quasi-silent germline genes coding for phylogenetically ancient determinants of the rabbit a locus allotypes. Eur J Immunol 7:15–22
van der Loo W, Arthur CP, Richardson BJ, Wallage-Drees M, Hamers R (1987) Nonrandom allele associations between unlinked protein loci: are the polymorphisms of the immunoglobulin constant regions adaptive? Proc Natl Acad Sci U S A 84:3075–3079
van der Loo W, Ferrand N, Soriguer RC (1991) Estimation of gene diversity at the b locus of the constant region of the immunoglobulin light chain in natural populations of European rabbit (Oryctolagus cuniculus) in Portugal, Andalusia and on the Azorean Islands. Genetics 127:789–799
Van der Loo W, Boussès P, Arthur CP, Chapuis JL (1996) Compensatory aspects of allele diversity at immunoglobulin loci: gene correlations in rabbit populations devoid of light chain diversity (Oryctolagus cuniculus L.; Kerguelen Islands). Genetics 144:1181–1194
van der Loo W, Mougel F, Sánchez MS, Bouton C, Castien E, Fonseca A, Ferrand N, Soriguer R, Monnerot M (1999a) Cytonuclear disequilibria in wild populations of rabbit (Oryctolagus cuniculus L.) suggest unequal allele turnover rates at the b locus (IGKC1). Immunogenetics 49:629–643
van der Loo W, Mougel F, Bouton C, Sanchez MS, Monnerot M (1999b) The allotypic patchwork pattern of the rabbit IGKC1 allele b5wf: genic exchange or common ancestry? Immunogenetics 49:7–14
van der Loo W, Abrantes J, Esteves PJ (2009) Sharing of endogenous lentiviral gene fragments among leporid lineages separated for more than 12 million years. J Virol 83:2386–2388. doi:10.1128/JVI.01116-08
van der Loo W, Afonso S, de Matos AL, Abrantes J, Esteves PJ (2012) Pseudogenization of the MCP-2/CCL8 chemokine gene in European rabbit (genus Oryctolagus), but not in species of Cottontail rabbit (Sylvilagus) and Hare (Lepus). BMC Genet 13:72. doi:10.1186/1471-2156-13-72
Van Valen L (1973) A new evolutionary law. Evol Theory 1:1–30
Villafuerte R, Calvete C, Blanco JC, Lucientes J (1995) Incidence of viral hemorrhagic disease in wild rabbit populations in Spain. Mammalia 59:651–660
Villemin JA (2015) On the virulence and specificity of tuberculosis. Int J Tuberc Lung Dis 19:256–266
Voisset C, Myers RE, Carne A, Kellam P, Griffiths DJ (2003) Rabbit endogenous retrovirus-H encodes a functional protease. J Gen Virol 84(Pt 1):215–225
Wagner B, Miller DC, Lear TL, Antczak DF (2004) The complete map of the Ig heavy chain constant gene region reveals evidence for seven IgG isotypes and for IgD in the horse. J Immunol 173:3230–3242
Wang F, Gao X, Barrett JW, Shao Q, Bartee E, Mohamed MR, Rahman M, Werden S, Irvine T, Cao J, Dekaban GA, McFadden G (2008) RIG-I mediates the co-induction of tumor necrosis factor and type I interferon elicited by myxoma virus in primary human macrophages. PLoS Pathog 4, e1000099
Werneck-Barroso E (1999) Innate resistance to tuberculosis: revisiting Max Lurie genetic experiments in rabbits. Int J Tuberc Lung Dis 3:166–168
Yap MW, Stoye JP (2013) Apparent effect of rabbit endogenous lentivirus type K acquisition on retrovirus restriction by lagomorph Trim5alphas. Philos Trans R Soc Lond B Biol Sci 368:20120498
Yap MW, Nisole S, Lynch C, Stoye JP (2004) Trim5alpha protein restricts both HIV-1 and murine leukemia virus. Proc Natl Acad Sci U S A 101:10786–10791
Yap MW, Nisole S, Stoye JP (2005) A single amino acid change in the SPRY domain of human Trim5alpha leads to HIV-1 restriction. Curr Biol 15:73–78
Yarovinsky F et al (2005) TLR11 activation of dendritic cells by a protozoan profilin-like protein. Science 308:1626–1629. doi:10.1126/science.1109893
Yoneyama M, Kikuchi M, Matsumoto K, Imaizumi T, Miyagishi M, Taira K, Foy E, Loo YM, Gale M Jr, Akira S, Yonehara S, Kato A, Fujita T (2005) Shared and unique functions of the DExD/H-box helicases RIG-I, MDA5, and LGP2 in antiviral innate immunity. J Immunol 175:2851–2858
Zhang D, Zhang G, Hayden MS, Greenblatt MB, Bussey C, Flavell RA, Ghosh S (2004) A Toll-like receptor that prevents infection by uropathogenic bacteria. Science 303:1522–1526. doi:10.1126/science.1094351
Zhu X, Boonthum A, Zhai SK, Knight KL (1999) B lymphocyte selection and age-related changes in VH gene usage in mutant Alicia rabbits. J Immunol 163:3313–3320
Zhuang X, Stahl SJ, Watts NR, DiMattia MA, Steven AC, Wingfield PT (2014) A cell-penetrating antibody fragment against HIV-1 Rev has high antiviral activity: characterization of the paratope. J Biol Chem 289:20222–20233
Zuniga MC (2002) A pox on thee! Manipulation of the host immune system by myxoma virus and implications for viral-host co-adaptation. Virus Res 88:17–33
Acknowledgments
This work is funded by national funds through FCT—Foundation for Science and Technology—under the project FCT-ANR/BIA-BIC/0043/2012. FCT also supported the doctoral fellowships of FN and AP (refs. SFRH/BD/81916/2011 and SFRH/BD/71252/2010) and the investigator grant of JA (ref. IF/01396/2013). This work is funded also by FEDER funds through the Operational Programme for Competitiveness Factors—COMPETE and by National Funds through FCT—Foundation for Science and Technology under the project PTDC/BIA-ANM/3963/2012 and FCOMP-01-0124-FEDER-028286 and “Genomics Applied To Genetic Resources” co-financed by North Portugal Regional Operational Programme 2007/2013 (ON.2-O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF). RGM was partially supported by the Intramural Research Program of the NIAID, NIH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pinheiro, A., Neves, F., Lemos de Matos, A. et al. An overview of the lagomorph immune system and its genetic diversity. Immunogenetics 68, 83–107 (2016). https://doi.org/10.1007/s00251-015-0868-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-015-0868-8