Abstract
Wolbachia is an extremely widespread intracellular symbiont which causes reproductive manipulation on various arthropod hosts. Male progenies are killed in Wolbachia-infected lineages of the Japanese Ostrinia moth population. While the mechanism of male killing and the evolutionary interaction between host and symbiont are significant concerns for this system, the absence of Wolbachia genomic information has limited approaches to these issues. We determined the complete genome sequences of wFur and wSca, the male-killing Wolbachia of Ostrinia furnacalis and Ostrinia scapulalis. The two genomes shared an extremely high degree of homology, with over 95% of the predicted protein sequences being identical. A comparison of these two genomes revealed nearly minimal genome evolution, with a strong emphasis on the frequent genome rearrangements and the rapid evolution of ankyrin repeat-containing proteins. Additionally, we determined the mitochondrial genomes of both species’ infected lineages and performed phylogenetic analyses to deduce the evolutionary dynamics of Wolbachia infection in the Ostrinia clade. According to the inferred phylogenetic relationship, two possible scenarios were proposed: (1) Wolbachia infection was established in the Ostrinia clade prior to the speciation of related species such as O. furnacalis and O. scapulalis, or (2) Wolbachia infection in these species was introgressively transferred from a currently unidentified relative. Simultaneously, the relatively high homology of mitochondrial genomes suggested recent Wolbachia introgression between infected Ostrinia species. The findings of this study collectively shed light on the host-symbiont interaction from an evolutionary standpoint.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Endosymbionts have a variety of effects on host biology. While mutualistic symbionts are necessary for the growth of their hosts, there are numerous instances in which symbionts negatively affect their hosts in a parasitic manner [1]. Wolbachia is an intracellular bacterial symbiont, notable for its widespread distribution among insects [2]. Wolbachia infection has been reported in a wide variety of insect lineages, as well as some other terrestrial arthropods and filarial nematodes. Wolbachia strains are classified into at least 17 supergroups, in which Supergroups A and B contain a majority of strains found in arthropod taxa [3]. In many associations between Wolbachia and its hosts, Wolbachia induces reproductive manipulation on the hosts, which typically results in infected females having a higher fitness level than uninfected females [1, 3]. Due to the vertical transmission of Wolbachia from mothers to their offspring, such manipulation is thought to spread Wolbachia infection throughout the host population. Wolbachia has four modes of reproductive manipulation: cytoplasmic incompatibility, feminization of genetic males, induction of parthenogenesis, and male killing [3]. Male killing is the process by which male offspring of infected mothers are killed during embryonic development or at a later stage, and is observed in a variety of bacterial parasites, including at least four different groups of bacteria [1]. Wolbachia-induced male killing has been reported for relatively limited groups, including Drosophila flies [4,5,6], ladybugs [7, 8], and some lepidopteran insects such as Acraea species [8,9,10], Hypolimnas bolina [11], Homona magnanima [12], and Ostrinia species [13, 14].
The genus Ostrinia is a member of the Crambidae family of Lepidoptera and contains several important agricultural pests. For example, Ostrinia furnacalis (Asian corn borer) and Ostrinia nubilalis (European corn borer) are serious maize pests. While O. furnacalis occurs in East and Southeast Asia and Oceania, O. nubilalis is found from Western Europe to Central Asia and in Northern America [15, 16]. Also, Ostrinia scapulalis (Adzuki bean borer), which is distributed sympatrically with the above two species in the Palearctic region, attacks some crops such as hops, hemp, and legumes [15, 16]. Thus, the understanding of reproductive manipulation caused by Wolbachia possibly leads to the development of biological control strategies for these pests. Historically, the genus Ostrinia was largely classified into three groups and are denoted by the letters I, II, and III, according to the morphology of male genitalia [17]. However, a recent phylogenomic study confirmed the monophyly of only group III and proposed a new classification system in which the genus is divided into three new species groups (Clade I, II, and III; summarized in Supplementary Fig. S1) [18]. All species of group III [17] were incorporated into Clade III, the Ostrinia nubilalis species group, along with some previously classified species in group II. Clade III species, particularly former group III members including O. furnacalis, O. nubilalis, and O. scapulalis, are morphologically very similar and genetically less diversified [15, 18, 19]. Although genetic differentiation was generally observed, several studies have described incomplete reproductive isolation among these species: no apparent postzygotic isolation was observed in any pair between O. furnacalis, O. nubilalis, and O. scapulalis in laboratory settings [20]. In addition, potential gene flow was detected between O. furnacalis–O. scapulalis and O. furnacalis–O. nubilalis pairs in natural populations [20, 22]. Five Clade III species (O. furnacalis, O. scapulalis, Ostrinia orientalis, Ostrinia zaguliaevi, and Ostrinia zealis) are distributed in Japan, which are characterized by host plants, female sex pheromone components, and/or male mid-tibia morphology (Supplementary Fig. S1; note that O. orientalis may be synonymized with O. scapulalis according to Frolov et al., 2007) [15, 16]. As for Japanese Ostrinia populations, the level of reproductive isolation among species in natural settings is largely untested.
Wolbachia causes male killing in Japanese populations of both O. furnacalis and O. scapulalis [13, 14]. The infection rate of Wolbachia in both species is comparatively low: a 5-year survey showed that 13 individuals out of 79 O. furnacalis females (16.5%) were infected in a particular area [21], whereas another study reported that 17 individuals out of 372 O. scapulalis females (4.6%) sampled from six locations were infected [23]. In addition, male-killing Wolbachia infection was found in other two “Clade III” species, O. orientalis and O. zaguliaevi in Japan [24]. The striking feature of infected strains of Ostrinia moths is that when the infection is cured with antibiotics, cured females produce all-male broods due to female-specific death [14]. This is in contrast to the situation with other male-killing Wolbachia, where antibiotic treatment restores a balanced sex ratio encompassing both sexes, as observed in Acraea butterflies [9, 10], Hypolimnas bolina [25], and Homona magnanima [12]. When Wolbachia is present in O. furnacalis, the expression of Ostrinia furnacalis Masculinizer gene (OfMasc) is downregulated at the embryonic stage [13]. OfMasc is an ortholog of Masculinizer gene, which was first discovered in the silkworm Bombyx mori and demonstrated to be required for both masculinization and dosage compensation [26]. OfMasc is thought to have masculinizing activity in O. furnacalis because knockdown of OfMasc results in the expression of female-type splicing variants of Ostrinia furnacalis doublesex (Ofdsx) in male embryos [27]. Males infected with Wolbachia were rescued from male killing by the injection of artificially synthesized OfMasc cRNA, and the expression of Z-linked genes was increased in Wolbachia-infected embryos [13]. Therefore, the depletion of OfMasc, which probably results in the failure of dosage compensation, may be the underlying mechanism of male-specific death in Wolbachia infection. Female-specific death in antibiotic treatment is also attributed to the aberrant expression of sex chromosome-linked genes [28]. These studies highlight the Wolbachia’s ingenious strategy for reproductive manipulation of Ostrinia hosts, generating increased interests in the evolutionary context of their association.
Genomic information is a critical resource for characterizing host-symbiont interactions. Recent advances in sequencing technology have resulted in the accumulation of Wolbachia genome sequences, which have aided in the identification of factors responsible for reproductive manipulation [29, 30] as well as the characterization of host-symbiont association dynamics at an ecological scale [31,32,33]. In order to obtain reliable insights, high-quality genome sequences from a diverse set of Wolbachia strains representing both phenotypic and phylogenetic diversity are required. However, there are a few genomes available for male-killing Wolbachia [34,35,36]. One is the complete genome sequence of wInn strain (Supergroup A), a male-killer of Drosophila innubila [35]. The other two are draft genome assemblies. wRec strain (Supergroup A), which causes cytoplasmic incompatibility in its native host Drosophila recens and male killing in an introgressed host Drosophila subquinaria, has a genome with reduced prophage regions compared to its close relative wMel strain [36]. The only Lepidoptera-associated male-killing strain with a draft genome is wBol1-b, a Supergroup B strain that infects H. bolina butterfly. The wBol1-b genome assembly comprised 144 contigs, 91 of which were organized into 13 scaffolds [34]. Along with the scarcity of genomic data on male-killing Wolbachia, the number of Wolbachia genome assemblies infecting Lepidopteran hosts is also limited. Indeed, no complete genomes of Wolbachia associated with Lepidoptera have been published, although a few are available in public databases. Despite the low number of Lepidoptera-associated Wolbachia genomes, a significant number of Wolbachia strains are present in Lepidoptera, presumably at a higher rate than in other arthropods [37, 38].
Mitochondrial genomes are also useful for deciphering host-symbiont associations. Since both inherited symbionts and mitochondria are transmitted maternally in infected lineages, examining mitochondrial haplotypes and Wolbachia infection status enables us to infer the mode of Wolbachia acquisition among allied species [31, 39,40,41]. Specifically, there are three distinct ways in which Wolbachia can be acquired by host species: cladogenic acquisition, introgression, and horizontal transmission [31, 32, 42]. Cladogenic acquisition occurs when related species codiverged (along cladogenesis) with Wolbachia derived from a common ancestor that was previously infected. Alternatively, Wolbachia infection can be introduced into a species via hybridization with infected sibling species and subsequent backcrossing. This is referred to as introgression. The relatively low level of genetic divergence of mitochondria and Wolbachia between host species can serve as an indicator in this case [42, 43]. The third mode is horizontal transmission, which typically results in discordance of phylogenetic relationship between Wolbachia and the host’s mitochondria.
Here, we report the complete genome sequences of male-killing Wolbachia infecting O. furnacalis and O. scapulalis (referred to as wFur and wSca, respectively), as well as the mitochondrial genomes of both hosts of infected lines. The comparison of these male-killing Wolbachia genomes revealed some genome rearrangements and gene repertoire differences between them, despite extremely high similarity, both of which would represent a trend in Wolbachia genome evolution. Along with mitochondrial genome analysis, the obtained genomic data enabled us to infer the evolutionary history of the Wolbachia infection in Ostrinia moths.
Materials and Methods
Insects
wFur-infected O. furnacalis and wSca-infected O. scapulalis were used in this study. An O. furnacalis line infected with wFur was previously described [13] and is maintained in our laboratory. wSca-infected O. scapulalis line was newly established by collecting an infected female moth in Matsudo, Japan (35.8° N, 139.9° E) in September 2020. Sex pheromone analysis was used to determine the species of collected Ostrinia females. Pheromone glands (PG) of three females from each matriline were immersed in hexane to extract sex pheromone. The PG extracts were analyzed using a GC–MS unit (QP2010 SE GC–MS, Shimadzu) equipped with a capillary column (DB-Wax, 0.25 mm i.d. × 30 m; Agilent Technologies, Santa Clara, CA, USA). The initial column oven temperature of 80 ℃ was held for 2 min and then increased at 8 ℃/min to 240 ℃. The final temperature was maintained for 2 min. The flow rate of the helium carrier gas was 1.0 mL/min. The diagnostic ion of the acetate ion at 61 (m/z) and its eliminated formation at 194 (m/z) were used for GC–MS analysis, in addition to the retention time. Both O. furnacalis and O. scapulalis lines were reared on an artificial diet (Insecta-LFS, Nosan Corp.) at 25 ℃ under a photoperiod of 16L8D and maintained by crossing with males of Wolbachia-uninfected counterpart lines derived from females collected in Nishi-Tokyo, Japan (35.7° N, 139.5° E). The infection was treated by adding 0.3 g tetracycline-HCl to a 500 g artificial diet.
Reverse Transcription-Polymerase Chain Reaction
Reverse transcription-polymerase chain reaction (RT-PCR) was used to examine the sex-specific splicing patterns of O. scapulalis doublesex (Osdsx). Egg masses (96 h post oviposition; hpo) from infected, uninfected, and cured lines were sampled. Total RNA and genomic DNA were extracted simultaneously from the egg masses using TRI Reagent (Molecular Research Center, Inc.) according to the manufacturer’s protocol. Total RNA was subjected to reverse transcription using AMV-RTase and an oligo-dT primer (TaKaRa). The resultant cDNA was used to perform PCR to examine the sex-specific splicing patterns of Osdsx. Genomic PCR of Wolbachia-specific wsp gene was used to examine the infection status. PCR was performed with KOD FX Neo DNA polymerase (TOYOBO) for Osdsx and KOD One PCR Master Mix (TOYOBO) for wsp. Primer sequences were reported previously [44].
Molecular Sexing
Ostrinia larvae were sexed using a previously reported method for O. furnacalis [13]. Briefly, genomic DNA was purified using DNeasy Blood & Tissue Kit (QIAGEN), and the dosage of Z-linked genes (Tpi and kettin) normalized by the autosome-linked gene (EF-1α) was evaluated by genomic qPCR. Individual larvae were photographed prior to DNA extraction, and body lengths were measured using ImageJ software.
Genome Sequencing and Assembly
Genomic DNA was purified from the ovaries of wFur-infected O. furnacalis and wSca-infected O. scapulalis females using Genomic-tip 100/G (QIAGEN) according to the manufacturer’s protocol. Prepared DNA samples were sequenced on NovaSeq 6000 (Illumina) and Sequel (PacBio) platforms. For O. furnacalis, two DNA samples were prepared each from an infected female. One of them was used for Illumina sequencing, and subsequently the remainder was combined with the other sample for PacBio (Sequel system) sequencing in continuous long read (CLR) mode. For O. scapulalis, one and four ovaries were used to prepare DNA samples for Illumina and PacBio (Sequel II system, CLR mode) sequencing, respectively. Illumina sequencing generated 54.48 and 31.79 Gb (150 bp, paired-end), and PacBio sequencing generated 45.17 and 166.46 Gb of raw data for O. furnacalis and O. scapulalis, respectively. Long-reads obtained by PacBio sequencing were assembled using Canu v2.1 assembler [45]. An estimated genome size of 438.5 Mb was given as a parameter with reference to the deposited genome assembly of O. furnacalis (accession number: GCA_004193835.1).
Construction of Wolbachia Genomes
Both wFur and wSca genomes were constructed through a following procedure. To identify contigs derived from Wolbachia in the Canu assembly, the generated contigs were subjected to blastn search as a query against RefSeq representative prokaryotic genomes database (ref_prok_rep_genomes) with the following parameters: “evalue 1e-20,” “max_target_seqs 60,” and “max_hsps 10.” Illumina short-reads were mapped to the whole Canu assembly containing draft Wolbachia contigs with BWA-MEM v.0.7.17 [46], and the resultant SAM file was converted to BAM format using samtools v.1.9 [47]. Pilon v.1.23 [48] was utilized to polish the draft Wolbachia contig. To confirm the unlikeliness of polishing errors due to misalignments of host-derived reads, the overall uniformity of the coverage of mapped short-reads was examined (Supplementary Fig. S2). In order to locate the overlap region at the beginning and end of the sequence, a blastn search was performed using the same polished contig as both a subject and a query. The overlap region was manually modified to construct a single-coverage genome (see Results). The oriC regions of wFur and wSca were identified by performing a homology search against the oriC sequence of wMel strain. The oriC sequence of wMel strain (DoriC ID: ORI10030016) was retrieved from the DoriC 5.0 database [49]. The binding sites in the oriC sequence were searched using the same criteria as previously described [50]. For genome annotation, stand-alone PGAP v5.2 (2021–05-19.build5429) [51] was utilized.
Construction of Mitochondrial Genomes
By performing a blastn search on mitochondrial gene sequences in the Canu assembly of wFur-infected O. furnacalis, a contig corresponding to a mitochondrial genome (mitogenome) was identified. Publicly available COI (Gene ID 65331444), COII (65331445), and ND5 (65331454) sequences from an O. furnacalis mitogenome (accession number: NC_056248.1) were used as queries. Illumina short-reads were mapped to the identified mitogenome contig with BWA-MEM v.0.7.17 [46]. Visual inspection of mapped reads was performed using IGV v.2.8.7 [52], and overlaps were manually modified to construct a single-coverage mitogenome. Subsequently, Illumina short-reads from wSca-infected O. scapulalis were mapped to the constructed mitogenome of wFur-infected O. furnacalis with BWA-MEM v.0.7.17 and visualized using IGV v.2.8.7. Detected mutations were manually modified to construct a mitogenome of wSca-infected O. scapulalis.
Genome Analysis
Some Wolbachia genes were searched in the wFur and wSca genomes using tblastn. The deduced amino acid sequences of wMel strain cifA (AAS14330.1), cifB (AAS14331.1), wmk (AAS14326.1), and TomO (AAS14922.1) were downloaded from GenBank and used as queries. To calculate the average nucleotide identity (ANI) between Wolbachia or mitochondrial genomes, FastANI v1.33 [53] was used. In order to conduct syntenic analysis, the wFur and wSca genomes were aligned using MUMmer v4.0.0rc1 [54] in the “nucmer” mode, and a dot plot was generated with the packaged program, mummerplot. A comparison of the gene repertoires of wFur and wSca was performed. To obtain non-redundant sequences for each strain’s annotated protein sequences, duplicated identical sequences were removed using SeqKit [55] in “rmdup” mode. Reciprocal blastp searches were used to identify orthologous gene pairs between the non-redundant wFur and wSca protein sequences. The hmmscan program was used in conjunction with HmmerWeb v.2.41.2 [56] to conduct a domain search against the Pfam 35.0 database using an E-value threshold of 0.05.
Phylogenetic Analysis
For Wolbachia, annotated protein sequences derived from 136 Wolbachia genome assemblies found in NCBI RefSeq as of December 2021 were downloaded. Additionally, two complete genome assemblies included in the analysis were not registered in RefSeq but were found in GenBank (GCA_000953315 and GCA_020995475). BUSCO v.5.2.2 [57] was used to determine the completeness of each genome assembly using rickettsiales_odb10. Two assemblies (GCF_000174095.1 and GCF_000167475.1) with low complete BUSCO percentage were excluded from the subsequent analysis. Using OrthoFinder v2.5.4 [58] with the following options: “-M msa” and “-os,” single-copy orthologs were identified across the aforementioned assemblies and the wFur and wSca genomes (140 in total; Supplementary table S1). Each of the 63 predicted single-copy orthologous groups was aligned using MAFFT v.7.490 [59] with default parameters, and the resultant alignments were trimmed using trimAl v.1.4.1 [60] in the “automated1” mode. The trimmed alignments for each strain were concatenated using SeqKit v2.1.0 [55], and were used for phylogenetic analysis.
A maximum likelihood (ML) tree was constructed using IQ-TREE v.1.6.12 [61], and node support was estimated with 1000 ultrafast bootstrap replicates [62]. ModelFinder [63] was used to determine the best-fitting substitution model, and the HIVw + F + R4 model was chosen. Additionally, we conducted phylogenetic analysis using only Wolbachia strains belonging to Supergroup B. Wolbachia genomes belonging to Supergroup B were chosen based on the constructed ML tree of all genome assemblies, and subjected to the following analysis. wMel (GCF_000008025.1) and wRi (GCF_000022285.1) genomes (Supergroup A) were also included as outgroups. The phylogenetic analysis was performed in the manner described previously. A total of 339 orthologs were used, with the HIVw + F + R3 model being the best-fitting.
In order to conduct phylogenetic analysis on mitochondrial genomes, we used publicly available sequences from GenBank (accession numbers are shown in Fig. 5). Mitogenomes were annotated de novo using MitoZ v2.4 [64] in the “annotate” mode with the following parameters: “–genetic_code 5” and “–clade Arthropoda.” Phylogenetic trees were constructed using the amino acid sequences of 13 protein-coding genes (PCGs) and the nucleotide sequences of 13 PCGs, rRNAs, and tRNAs. In both cases, ML trees were constructed using a partitioned model with partitioning scheme selection (“-m MFP + MERGE” option) in IQ-TREE v.1.6.12 [65]. Alignments of each deduced amino acid sequence were generated using MAFFT v.7.471, trimmed with trimAl v1.4.rev22, and subsequently concatenated. Each protein was assigned a partition. For nucleotide sequences, mitogenomes were aligned directly using MAFFT v.7.471, and partitions for each gene were defined based on annotated positions in a single strain (MN793323.1). The optimal partition scheme and substitution model for each partition were selected based on Bayesian information criterion.
Results
Characterization of wSca-Induced Male Killing in O. scapulalis
We used wFur-infected O. furnacalis [13] and wSca-infected O. scapulalis lines for genome sequencing. A wSca-infected O. scapulalis line was newly established from a field-collected female moth in this study. We used GC–MS to identify species using hexane extract of pheromone glands. The extract of the collected line contained (E)-11- and (Z)-11-tetradecenyl acetates (retention times were 16.87 min and 16.99 min, respectively), confirming that it is O. scapulalis [16]. This wSca-infected line exhibited a nearly entirely female-biased sex ratio through at least 11 generations, indicating the presence of male killing (Fig. 1 a and b). Antibiotics treatment of infection resulted in the occurrence of only male progeny. In order to ascertain the period during which males are killed in wSca-infected O. scapulalis, molecular sexing using qPCR was performed on 4 and 14 days post-hatch (dph) larvae. As a result, both females and males were detected in 4 dph larvae, but only females were observed in 14 dph larvae (Fig. 1c). In addition, wSca-infected males were significantly smaller than females in 4 dph larvae, unlike uninfected larvae, which showed no difference in body length between sexes (Fig. 1d). The splicing pattern of Osdsx, which represents the phenotype of sexual differentiation, was female in the infected line, male in the cured progeny, and both female and male in the uninfected line (Fig. 1e). These observations demonstrate that wSca disrupts the host’s sex determination cascade from embryogenesis, and that wSca-infected males die during the larval stage of O. scapulalis.
Characterization of male killing in wSca-infected O. scapulalis. a Brood sex ratios in the wSca-infected matriline. The female:male ratio of each mating is shown. Tetracycline treatment was conducted to remove wSca from a subpopulation of the second generation. Sexing was conducted based on the morphology of the pupal abdominal tips. b An adult female moth infected with wSca. Bar, 10 mm. c The sex ratios of wSca-infected larvae at 4 and 14 dph. The number indicates the sample size of each group. d The body length of wSca-infected and uninfected larvae at 4 dph. Data presented are mean ± standard error of the mean. Dot plots show all data points individually. An asterisk denotes statistical significance (P < 0.001; N.S., not significant, P > 0.1; Wilcoxon rank-sum test). e Splicing patterns of Osdsx in uninfected, wSca-infected, and infection-cured embryos at 96 hpo. The results of technical duplicates of the RT-PCR assay are presented for each sample. The numbers indicate individual samples (biological duplicates for each condition). Adult female and male moths were used as positive controls. The letters F and M indicate female- and male-type splicing variants, respectively. AF: adult female; AM: adult male; NT: no template
The Construction of the wFur and wSca Genomes
We conducted both long-read (PacBio) and short-read (Illumina) sequencing. First, long-reads were assembled by Canu assembler [45] with an estimated genome size of 438.5 Mb according to the only available genome assembly of O. furnacalis in Genbank. The resultant assembly of O. furnacalis and O. scapulalis contained 4890 and 3535 contigs (1.25 Gb and 1.44 Gb), respectively. For each assembly of O. furnacalis and O. scapulalis, the blastn search using each contig as a query resulted in only one contig that frequently aligned to publicly available Wolbachia sequences. Given that both of these two contigs were 1.3–1.4 Mbp in length, which is within the range of typical Wolbachia genome sizes [3], and their coverage was among the highest in each assembly (Supplementary Fig. S3), these two contigs were most likely the genomes of wFur and wSca. The polishing process using Illumina reads by Pilon [48] identified a few errors in initial contigs (two changes for wFur and no changes for wSca; all of them were homopolymer errors, e.g., AAAAA to AAAAAA). Since the polished genomes had an overlap region of approximately 50 kb (wFur) and 80 kb (wSca) at the head and tail of the contigs, duplicated sequences were removed to make the genomes single coverage. When there was inconsistency between the head and tail overlapping regions, we consulted the mapped short-reads to determine which one should be adopted. Due to the circular nature of the Wolbachia genome, we manually set the starting points of the sequences to the estimated oriC regions. The oriC region was identified using a blastn search with the wMel oriC sequence as a query. wFur and wSca had identical estimated oriC sequences (404 bp). It contained three DnaA-, four CtrA-, and two IHF-binding sites, and was flanked by hemE and CBS domain protein genes, all of which are characteristic of Wolbachia [50].
To check the quality of constructed genome sequences, we mapped Illumina paired-end short-reads against wFur and wSca genomes and examined the coverage of all mapped reads as well as that of properly paired reads (Supplementary Fig. S4). An aberrant peak was detected exclusively in the coverage of all mapped reads, which resulted from misalignments of host-derived partial rDNA sequences. No other regions exhibited discrepancy, suggesting little effect of host-derived reads in the genome construction process. Next, BUSCO program [57] was used to assess the completeness of the genomes. This program was routinely used to determine the presence/absence of benchmarking single-copy orthologs expected to exist among specific taxa. The BUSCO profiles of the wFur and wSca genomes against the Rickettsiales database were identical, with 362/364 single-copy orthologs detected, comparable to the profiles of complete Wolbachia genome assemblies (Table 1 and Supplementary Fig. S5). Thus, we ascertained that our assemblies are of sufficient quality to conduct further analyses.
The Properties of the wFur and wSca Genomes
The genomes of wFur and wSca are 1,321,828 and 1,320,340 bp in length, respectively (Fig. 2). The wFur genome contained 1040 protein-coding genes, 158 pseudogenes, 34 tRNAs, 3 rRNAs, and 4 ncRNAs, while the wSca genome contained 1050 protein-coding genes, 153 pseudogenes, 34 tRNAs, 3 rRNAs, and 4 ncRNAs. Some of host-manipulating Wolbachia genes were screened individually (Supplementary Table S2). Each of the two genomes encodes a putatively pseudogenized copy of cifA, one of the two factors responsible for cytoplasmic incompatibility [29]. Regarding cifB, there are no full-length copies, but one gene shares partial homology with the wMel cifB sequence. This gene, named Oscar, was recently demonstrated to have male-killing activity in lepidopteran insects [66]. Two intact copies of wmk, a candidate gene for male killing [30], were found in each of the two genomes. Three genes (including two putative pseudogenes) share homology with TomO gene, a growth-inhibitory and RNA-interacting factor of wMel [67]. However, two of them might be a single ORF misinferred during the automated annotation process.
Circular map of the wFur and wSca genomes. Circles arranged in order from outer to inner indicate the following: the locations of annotated coding sequences (CDS) on the positive (outermost) and the negative (second outermost) strands, the position of annotated RNAs, GC content, and GC skew. The GC content and GC skew are calculated in 10 kb windows and expressed as deviations from an average of the whole sequence
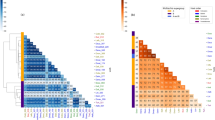
Subsequently, phylogenetic analysis using 138 Wolbachia genome assemblies was performed to ascertain the phylogenetic relationship of wFur and wSca within Wolbachia (Fig. 3 and Supplementary Fig. S6). As a result, wFur and wSca were assigned to Supergroup B collectively. The most closely related strain was that which infects the Noctuid moth, Spodoptera picta. Notably, the second closest strain was wTpre, which is found in a parasitoid wasp Trichogramma pretiosum and is responsible for parthenogenesis [68]. Intriguingly, the wFur and wSca strains did not cluster phylogenetically with the wBol1-b strain, which induces male killing in the H. bolina butterfly [34].
Phylogenetic relationship of Supergroup B Wolbachia genomes. The maximum likelihood tree constructed from concatenated protein sequences of 339 single-copy orthologs is shown. wMel and wRi (Supergroup A) were used as outgroups. The strain names and their hosts are labeled. If no suitable strain name is available, it is denoted by “NA”. Clades composed of almost the same strains are collapsed, and the number of contained strains is labeled. Branch support calculated using 1000 replicates of ultrafast bootstrap is shown on the nodes. The sequences determined in this study are highlighted in red and bold, while strains associated with other lepidopterans are in blue
Comparative Analysis of the wFur and wSca Genomes
The ANI calculated by fastANI program [53] was 99.9755%, indicating extremely high similarity between these two Wolbachia. At the genome structure level, however, we found several large inversions between them (Fig. 4). To compare gene repertoires, we first removed duplicated, identical protein sequences from each genome. We then used blastp to compare the strains’ unique sequences (referred to as “non-redundant sequences”). As a result, 976 out of 982 non-redundant wFur sequences have reciprocal best hit counterparts in 993 non-redundant wSca sequences. Out of these 976 sequences, 952 were identical between two strains. The remaining 24 sequences differed between the strains to a varying degree, whereas 18 have a single amino acid substitution, one of the remaining six had 24 amino acid substitutions and seven gaps (Supplementary table S3). Most of the 6 wFur- and 17 wSca-specific sequences were hypothetical proteins or transposase variants (Supplementary table S3). However, wSca uniquely has three ankyrin repeat-containing proteins.
Analysis of the Mitochondrial Genome
The genome assemblies included information about both the hosts and Wolbachia. Mitochondria, in particular, are maternally inherited elements that are descended together with Wolbachia in infected lineages. Therefore, we analyzed mitochondrial genomes to gain insight into an evolutionary perspective of the host and symbiont. The mitochondrial genome of wFur-infected O. furnacalis was successfully extracted from the assembly using blastn, and constructed as a single-coverage complete contig. In contrast, since the mitochondrial genome of O. scapulalis was not found in the Canu assembly, it was reconstructed by mapping the short-reads to the mitochondrial genome of wFur-infected O. furnacalis. The resultant mitochondrial genomes are 15,266 bp for wFur-infected O. furnacalis and 15,249 bp for wSca-infected O. scapulalis.
The ANI between these two mitogenomes was 99.90%, which was greater than the ANI between normal O. furnacalis and O. scapulalis (98.77–98.78%) (Supplementary Fig. S7). Between the mitogenomes of wFur-infected O. furnacalis and wSca-infected O. scapulalis, we detected 14 SNPs (all of which were transitions; no transversions were detected) and several indels (1 base deletion, 2 base insertion, and 18 base deletion in the O. scapulalis mitochondrial genome compared to that of O. furnacalis, of which the last two were in AT-rich region). At the protein level, we found only two amino acid substitutions in the entire mitochondrial protein sequence. On the other hand, the mitochondrial haplotypes of infected Ostrinia differed substantially from those of uninfected counterparts. For example, 45 substitutions in whole protein sequences deduced from the mitochondrial genomes exist between normal (Genbank accession: MN793323) and wFur-infected O. furnacalis. These findings indicate that Wolbachia-infected Ostrinia species have a distinct mitochondrial haplotype from uninfected individuals, which likely reflects a long period of maternal lineage separation.
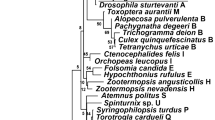
Phylogenetic analysis was performed using mitochondrial genomes of related species obtained from NCBI Genbank. Both infected lineages formed a cluster, which was located outside Clade III Ostrinia species including O. furnacalis, O. scapulalis, O. nubilalis, O. zealis, and Ostrinia kasmirica (Fig. 5). We conducted a phylogenetic analysis using either nucleotide or protein sequences, and found that the topology of resultant ML trees was identical except for the phylogenetic relationship between infected lineages and O. kasmirica and the detailed intraclade relationship within Clade III Ostrinia species (Supplementary Fig. S8). Essentially, the inferred trees were consistent with previous reports [69, 70]. According to these results, we conclude that the mitochondrial haplotype associated with Wolbachia infection originated in an ancestral population of Clade III Ostrinia species (probably excluding O. kasmirica based on phylogeny estimated by nucleotide sequences; Supplementary Fig. S8).
Phylogenetic relationship of mitochondrial genomes of Ostrinia and allied moth species. The maximum likelihood tree constructed from concatenated sequences of 13 proteins with partitions for each protein is shown. Two pyralid species (Lista haraldusalis and Ephestia kuehniella) were used as outgroups. Branch support calculated using 1000 replicates of ultrafast bootstrap is shown on the nodes. The sequence determined in this study are highlighted in red and bold
Discussion
We successfully sequenced the genomes of two closely related Wolbachia strains, wFur and wSca, which represent the first complete genomes of male-killing Wolbachia in lepidopteran hosts. The two genomes shared remarkably high homology, with over 95% of protein sequences identical. On the other hand, there are substantial differences between the genomes, most notably some large inversions (Fig. 4). It is well-established that Wolbachia genomes poorly retain synteny between distant strains, particularly parasitic strains found in arthropod hosts [71, 72]. We observed nearly minimal transition of genome structure in two male-killing Wolbachia from Ostrinia.
The wFur and wSca genomes encode at least 29 and 32 ankyrin repeat-containing proteins, respectively (confirmed by hmmscan). The ankyrin repeat motif is a 33 amino acid sequence that plays a role in protein–protein interaction. Although ankyrin repeat-containing proteins are found predominantly in eukaryotes, they are also used as effectors by various pathogenic and symbiotic microbes [73]. Certain genes encoding ankyrin repeat-containing proteins are found only in one of the two strains, implying that ankyrin repeat-containing proteins evolved rapidly in Wolbachia. The features identified by comparing closely related Wolbachia strains (wFur and wSca), namely rampant genome structure rearrangement and rapid evolution of ankyrin repeat-containing proteins, most likely represent a trend of minute genome evolution in Wolbachia.
The phylogenetic analysis identified Wolbachia of S. picta (the lily caterpillar) as the closest relative of wFur and wSca (Fig. 3). This implies the presence of Wolbachia host shifts between these lepidopteran hosts in evolutionary time scale. The second most closely related strain was wTpre, a Wolbachia strain found in the parasitoid wasp T. pretiosum. Given the close ecological relationship between parasitic wasps and their hosts, including moths, the high sequence homology detected among Wolbachia species possibly indicate Wolbachia transmission among lepidopteran insects and their parasites. Some studies showed that horizontal transmission of Wolbachia is possible with the involvement of parasitic wasps [74, 75]. Thus, wasps may be a major driver of macro-scale dynamics of Wolbachia, even though interspecies transmission is less well understood.
Mitochondrial analysis of Wolbachia-infected lineages revealed an evolutionary history of symbiotic association in the Ostrinia clade. Because mitochondria are inherited maternally, infected lineages’ mitochondrial haplotypes inevitably descended with Wolbachia. Thus, once infection is established in a population, this matriline (namely, the mitochondrial haplotype) becomes distinct from the rest of (uninfected) population from that point on, and genetic differentiation of mitochondria would proceed even in the same gene pool. Particularly in Ostrinia, the removal of Wolbachia results in female-specific death [14], which limits the emergence of Wolbachia-free matrilines that retain the original Wolbachia-associated mitochondrial haplotype. This unique feature likely underlies the distinctiveness of mitochondrial phylogeny in infected Ostrinia moths. Therefore, as a result of the phylogenetic tree, we can infer that the bifurcation of mitochondrial phylogeny between infected lineages and Clade III Ostrinia corresponds to the infection establishment, thus the infection was established before the speciation of Clade III Ostrinia, which includes O. furnacalis and O. scapulalis. Alternatively, however, another possibility exists: the bifurcation of mitochondrial phylogeny can correspond to the occurrence of a newly emerged Ostrinia lineage, which is currently undiscovered or perhaps extinct, and Wolbachia infection was established later in this clade. In this case, the subsequent introgressive transmission of infection to Clade III Ostrinia has to be assumed.
The ANI between mitochondrial haplotypes of wFur-infected O. furnacalis and wSca-infected O. scapulalis was greater than that between uninfected counterparts (Supplementary Fig. S7). This demonstrates that Wolbachia-infected O. furnacalis and O. scapulalis have unusually similar mitochondrial haplotypes compared to their divergence time. Among the possible modes of Wolbachia dynamics [31, 42], the situation is best explained by the introgressive transmission of Wolbachia. The cross between O. furnacalis and O. scapulalis produces fertile hybrids that exhibit both parents’ sex pheromone types in a laboratory setting [76]. Besides, in natural populations in China, presumed hybrid individuals and introgressed genotypes have been detected [20]. These reports demonstrate the incompleteness of reproductive isolation between these two species, which supports the possibility of Wolbachia introgression. In addition, the presence of male-killing Wolbachia in other Ostrinia species, namely O. orientalis (which could be synonymized with O. scapulalis [15]) and O. zaguliaevi, suggests that Wolbachia introgression probably occurred among these species [24].
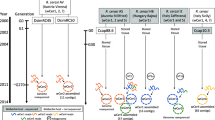
We now propose two possible evolutionary history models of the symbiotic association between Ostrinia species and Wolbachia based on the preceding discussion. In the first model (Fig. 6a), (i) Wolbachia infection was established in the ancestral population prior to Clade III Ostrinia speciation. (ii) Subsequently, Clade III Ostrinia species such as O. furnacalis, O. scapulalis, and others diversified, and at least one species retained Wolbachia infection. (iii) Wolbachia was recently transmitted via introgression among O. furnacalis, O. scapulalis, and possibly other Clade III species. Alternatively, in the second model (Fig. 6b), (i) Wolbachia infection was established in an unidentified Ostrinia relative which had diverged from Clade III phylogeny. (ii) Subsequently, infection was transmitted to Clade III Ostrinia species via introgression. (iii) Wolbachia was transmitted recently via introgression among Clade III Ostrinia species as in the first model.
A proposed model for evolutionary history of Wolbachia infection in Ostrinia moths. Each circle represents an assumed individual species in evolutionary time scale. Wolbachia-infected subpopulations are depicted in a dark color. Black arrowheads indicate when the divergence of mitochondrial haplotypes currently found in infected and uninfected Clade III Ostrinia moths began. a The establishment of Wolbachia infection can be traced back to the beginning of mitochondrial divergence at the maximum, or b more recently by assuming introgressive transfer from an unidentified (possibly extinct) relative
While the outline of evolutionary scenarios was inferred, there are still unanswered questions that should be addressed in future work. First, what is the real origin of Wolbachia in Ostrinia moths? The phylogenetic tree clearly depicts the evolutionary relationship between Wolbachia associated with Ostrinia and other moth- and parasitoid wasp-associated Wolbachia, implying the possibility of interspecies transfer. However, since Wolbachia was estimated to be present in roughly half of all terrestrial arthropods [2], the current analysis contained only a fraction of Wolbachia’s total diversity. A thorough survey of Wolbachia diversity within the Lepidoptera and wasp clades, as well as possibly other clades, is required to characterize interspecies dynamics of Wolbachia. Ostrinia-associated strains with a different origin than wFur and wSca may provide insight into interspecies dynamics. For instance, Wolbachia strains belonging to Supergroup A have been reported from O. furnacalis populations in China [77].
Second, we sought to determine the direction of O. furnacalis and O. scapulalis introgression. Unidirectional mitochondrial introgression from O. furnacalis into O. scapulalis has been observed in Chinese populations [20]. This indicates that the direction’s introgressive Wolbachia transmission was more likely. Given the high degree of homology between wFur and wSca or the hosts’ mitogenomes, and the resulting assumption of recent introgression, such transmission and hybridization may frequently occur among Ostrinia species. Indeed, it is unclear whether infected Ostrinia populations are genetically isolated clearly along with classical species such as O. furnacalis and O. scapulalis or they are spread across multiple species with incomplete reproductive isolation.
In conclusion, we determined two complete genomes of male-killing Wolbachia in two closely related lepidopteran hosts. These sequences were used to characterize nearly minimal genome evolution of Wolbachia, which was only possible through comparison of high-quality complete genomes. The mitochondrial genome analysis facilitated our investigation of the evolutionary relationship between Ostrinia moths and Wolbachia. It revealed Wolbachia’s complex dynamics across multiple hosts, including the infection establishment possibly before allied species diversification and the introgressive transmission following host speciation. The genomic data on male-killing Wolbachia obtained in this study will serve as a foundation for future research on host-symbiont interaction, which is crucial to assessing evolutionary impacts of potential use of this symbiont for a biological control agent.
Data Availability
All data are available in the main text or the supplemental materials. The genome sequences of wFur and wSca have been deposited in GenBank under accession numbers CP096925 and CP096926, respectively. The mitochondrial genomes of wFur-infected O. furnacalis and wSca-infected O. scapulalis have been deposited in GenBank under accession numbers ON645200 and ON645201, respectively.
References
Goryacheva I, Andrianov B (2021) Reproductive parasitism in insects. The interaction of host and bacteria. Biol Commun 66:17–27. https://doi.org/10.21638/spbu03.2021.103
Weinert LA, Araujo-Jnr EV, Ahmed MZ, Welch JJ (2015) The incidence of bacterial endosymbionts in terrestrial arthropods. Proc R Soc B 282:20150249. https://doi.org/10.1098/rspb.2015.0249
Kaur R, Shropshire JD, Cross KL et al (2021) Living in the endosymbiotic world of Wolbachia: a centennial review. Cell Host Microbe 29:879–893. https://doi.org/10.1016/j.chom.2021.03.006
Dyer KA, Jaenike J (2004) Evolutionarily stable infection by a male-killing endosymbiont in Drosophila innubila: molecular evidence from the host and parasite genomes. Genet 168:1443–1455. https://doi.org/10.1534/genetics.104.027854
Hurst GDD, Johnson AP, Schulenburg JHGVD, Fuyama Y (2000) Male-killing Wolbachia in Drosophila: A temperature-sensitive trait with a threshold bacterial density. Genet 156:699–709. https://doi.org/10.1093/genetics/156.2.699
Sheeley SL, McAllister BF (2009) Mobile male-killer: similar Wolbachia strains kill males of divergent Drosophila hosts. Heredity 102:286–292. https://doi.org/10.1038/hdy.2008.126
Elnagdy S, Messing S, Majerus MEN (2013) Two strains of male-killing Wolbachia in a ladybird Coccinella undecimpunctata from a hot climate. PLoS One 8:e54218. https://doi.org/10.1371/journal.pone.0054218
Hurst GDD, Jigging FM, Von Der Schulenburg JHG et al (1999) Male-killing Wolbachia in two species of insect. Proc R Soc B 266:735–740. https://doi.org/10.1098/rspb.1999.0698
Jiggins FM, Hurst GDD, Majerus MEN (1998) Sex ratio distortion in Acraea encedon (Lepidoptera: Nymphalidae) is caused by a male-killing bacterium. Hered 81:87–91. https://doi.org/10.1046/j.1365-2540.1998.00357.x
Jiggins FM, Hurst GDD, Dolman CE, Majerus MEN (2000) High-prevalence male-killing Wolbachia in the butterfly Acraea encedana. J Evol Biol 13:495–501. https://doi.org/10.1046/j.1420-9101.2000.00180.x
Dyson EA, Kamath MK, Hurst GDD (2002) Wolbachia infection associated with all-female broods in Hypolimnas bolina (Lepidoptera: Nymphalidae): evidence for horizontal transmission of a butterfly male killer. Hered 88:166–171. https://doi.org/10.1038/sj.hdy.6800021
Arai H, Lin SR, Nakai M et al (2020) Closely related male-killing and nonmale-killing Wolbachia strains in the oriental tea tortrix Homona magnanima. Microb Ecol 79:1011–1020. https://doi.org/10.1007/s00248-019-01469-6
Fukui T, Kawamoto M, Shoji K et al (2015) The endosymbiotic bacterium Wolbachia selectively kills male hosts by targeting the masculinizing gene. PLoS Pathog 11:1005048. https://doi.org/10.1371/journal.ppat.1005048
Kageyama D, Traut W (2004) Opposite sex-specific effects of Wolbachia and interference with the sex determination of its host Ostrinia scapulalis. Proc R Soc B 271:251–258. https://doi.org/10.1098/rspb.2003.2604
Frolov AN, Bourguet D, Ponsard S (2007) Reconsidering the taxomony of several Ostrinia species in the light of reproductive isolation: a tale for Ernst Mayr. Biol J Linn Soc 91:49–72. https://doi.org/10.1111/j.1095-8312.2007.00779.x
Ishikawa Y, Takanashi T, Kim CG et al (1999) Ostrinia spp. in Japan: their host plants and sex pheromones. Entomol Exp Appl 91:237–244. https://doi.org/10.1046/j.1570-7458.1999.00489.x
Mutuura A, Munroe E (1970) Taxonomy and distribution of the European corn borer and allied species: genus Ostrinia (Lepidoptera: Pyralidae). Mem Entomol Soc Canada 102:1–112. https://doi.org/10.4039/entm10271fv
Yang Z, Plotkin D, Landry JF et al (2021) Revisiting the evolution of Ostrinia moths with phylogenomics (Pyraloidea: Crambidae: Pyraustinae). Syst Entomol 46:827–838. https://doi.org/10.1111/syen.12491
Kim C, Hoshizaki S, Huang Y et al (1999) Usefulness of mitochondrial COII gene sequences in examining phylogenetic relationships in the Asian corn borer, Ostrinia furnacalis, and allied species (Lepidoptera : Pyralidae). Appl Entomol Zool 34:405–412. https://doi.org/10.1303/aez.34.405x
Bourguet D, Ponsard S, Streiff R et al (2014) ‘Becoming a species by becoming a pest’ or how two maize pests of the genus Ostrinia possibly evolved through parallel ecological speciation events. Mol Ecol 23:325–342. https://doi.org/10.1111/mec.12608
Kageyama D, Nishimura G, Hoshizaki S, Ishikawa Y (2002) Feminizing Wolbachia in an insect, Ostriniafurnacalis (Lepidoptera: Crambidae). Hered 88:444–449. https://doi.org/10.1038/sj.hdy.6800077
Wang Y, Kim KS, Guo W et al (2017) Introgression between divergent corn borer species in a region of sympatry: Implications on the evolution and adaptation of pest arthropods. Mol Ecol 26:6892–6907. https://doi.org/10.1111/mec.14387
Kageyama D, Nishimura G, Hoshizaki S, Ishikawa Y (2003) Two kinds of sex ratio distorters in a moth, Ostrinia scapulalis. Genome 46:974–982. https://doi.org/10.1139/g03-083
Kageyama D, Nishimura G, Ohno S et al (2004) Wolbachia infection and an all-female trait in Ostrinia orientalis and Ostrinia zaguliaevi. Entomol Exp Appl 111:79–83. https://doi.org/10.1111/j.0013-8703.2004.00153.x
Charlat S, Davies N, Roderick GK, Hurst GDD (2007) Disrupting the timing of Wolbachia-induced male-killing. Biol Lett 3:154–156. https://doi.org/10.1098/rsbl.2006.0584
Kiuchi T, Koga H, Kawamoto M et al (2014) A single female-specific piRNA is the primary determiner of sex in the silkworm. Nature 509:633–636. https://doi.org/10.1038/nature13315
Fukui T, Kiuchi T, Shoji K et al (2018) In vivo masculinizing function of the Ostrinia furnacalis Masculinizer gene. Biochem Biophys Res Commun 503:1768–1772. https://doi.org/10.1016/j.bbrc.2018.07.111
Sugimoto TN, Kayukawa T, Shinoda T et al (2015) Misdirection of dosage compensation underlies bidirectional sex-specific death in Wolbachia-infected Ostrinia scapulalis. Insect Biochem Mol Biol 66:72–76. https://doi.org/10.1016/j.ibmb.2015.10.001
Le Page DP, Metcalf JA, Bordenstein SR et al (2017) Prophage WO genes recapitulate and enhance Wolbachia-induced cytoplasmic incompatibility. Nature 543:243–247. https://doi.org/10.1038/nature21391
Perlmutter JI, Bordenstein SR, Unckless RL et al (2019) The phage gene wmk is a candidate for male killing by a bacterial endosymbiont. PLoS Pathog 15:e1007936. https://doi.org/10.1371/journal.ppat.1007936
Cooper BS, Vanderpool D, Conner WR et al (2019) Wolbachia acquisition by Drosophila yakuba-clade hosts and transfer of incompatibility loci between distantly related Wolbachia. Genet 212:1399–1419. https://doi.org/10.1534/genetics.119.302349
Turelli M, Cooper BS, Richardson KM et al (2018) Rapid global spread of wRi-like Wolbachia across multiple Drosophila. Curr Biol 28:963–971. https://doi.org/10.1016/j.cub.2018.02.015
Wolfe TM, Bruzzese DJ, Klasson L et al (2021) Comparative genome sequencing reveals insights into the dynamics of Wolbachia in native and invasive cherry fruit flies. Mol Ecol 30:6259–6272. https://doi.org/10.1111/mec.15923
Duplouy A, Iturbe-Ormaetxe I, Beatson SA et al (2013) Draft genome sequence of the male-killing Wolbachia strain wBol1 reveals recent horizontal gene transfers from diverse sources. BMC Genomics 14:20. https://doi.org/10.1186/1471-2164-14-20
Hill T, Unckless RL, Perlmutter JI (2021) Positive selection and horizontal gene transfer in the genome of a male-killing Wolbachia. Mol Biol Evol 39:msab303. https://doi.org/10.1093/molbev/msab303
Metcalf JA, Jo M, Bordenstein SR et al (2014) Recent genome reduction of Wolbachia in Drosophila recens targets phage WO and narrows candidates for reproductive parasitism. PeerJ 2:e529. https://doi.org/10.7717/peerj.529
Ahmed MZ, Araujo-Jnr EV, Welch JJ, Kawahara AY (2015) Wolbachia in butterflies and moths: Geographic structure in infection frequency. Front Zool 12:16. https://doi.org/10.1186/s12983-015-0107-z
Ilinsky Y, Kosterin OE (2017) Molecular diversity of Wolbachia in Lepidoptera: prevalent allelic content and high recombination of MLST genes. Mol Phylogenet Evol 109:164–179. https://doi.org/10.1016/j.ympev.2016.12.034
Baldo L, Ayoub NA, Hayashi CY et al (2008) Insight into the routes of Wolbachia invasion: High levels of horizontal transfer in the spider genus Agelenopsis revealed by Wolbachia strain and mitochondrial DNA diversity. Mol Ecol 17:557–569. https://doi.org/10.1111/j.1365-294X.2007.03608.x
Charlat S, Duplouy A, Hornett EA et al (2009) The joint evolutionary histories of Wolbachia and mitochondria in Hypolimnas bolina. BMC Evol Biol 9:64. https://doi.org/10.1186/1471-2148-9-64
Sucháčková Bartoňová A, Konvička M, Marešová J et al (2021) Wolbachia affects mitochondrial population structure in two systems of closely related Palaearctic blue butterflies. Sci Rep 11:3019. https://doi.org/10.1038/s41598-021-82433-8
Raychoudhury R, Baldo L, Oliveira DCSG, Werren JH (2009) Modes of acquisition of Wolbachia: horizontal transfer, hybrid introgression, and codivergence in the Nasonia species complex. Evol 63:165–183. https://doi.org/10.1111/j.1558-5646.2008.00533.x
Meany MK, Conner WR, Richter SV et al (2019) Loss of cytoplasmic incompatibility and minimal fecundity effects explain relatively low Wolbachia frequencies in Drosophila mauritiana. Evol 73:1278–1295. https://doi.org/10.1111/evo.13745
Sugimoto TN, Fujii T, Kayukawa T et al (2010) Expression of a doublesex homologue is altered in sexual mosaics of Ostrinia scapulalis moths infected with Wolbachia. Insect Biochem Mol Biol 40:847–854. https://doi.org/10.1016/j.ibmb.2010.08.004
Koren S, Walenz BP, Berlin K et al (2017) Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res 27:722–736. https://doi.org/10.1101/gr.215087.116
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 1303.3997 [q-bio.GN] https://doi.org/10.48550/arXiv.1303.3997
Danecek P, Bonfield JK, Liddle J et al (2021) Twelve years of SAMtools and BCFtools. Gigasci 10:giab008. https://doi.org/10.1093/gigascience/giab008
Walker BJ, Abeel T, Shea T et al (2014) Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9:e112963. https://doi.org/10.1371/journal.pone.0112963
Gao F, Luo H, Zhang CT (2013) DoriC 5.0: an updated database of oriC regions in both bacterial and archaeal genomes. Nucleic Acids Res 41:D90–D93. https://doi.org/10.1093/nar/gks990
Ioannidis P, Hotopp JCD, Sapountzis P et al (2007) New criteria for selecting the origin of DNA replication in Wolbachia and closely related bacteria. BMC Genomics 8:182. https://doi.org/10.1186/1471-2164-8-182
Tatusova T, Dicuccio M, Badretdin A et al (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Thorvaldsdóttir H, Robinson JT, Mesirov JP (2013) Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14:178–192. https://doi.org/10.1093/bib/bbs017
Jain C, Rodriguez-R LM, Phillippy AM et al (2018) High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun 9:5114. https://doi.org/10.1038/s41467-018-07641-9
Marçais G, Delcher AL, Phillippy AM et al (2018) MUMmer4: a fast and versatile genome alignment system. PLoS Comput Biol 14:e1005944. https://doi.org/10.1371/journal.pcbi.1005944
Shen W, Le S, Li Y, Hu F (2016) SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS One 11:e0163962. https://doi.org/10.1371/journal.pone.0163962
Potter SC, Luciani A, Eddy SR et al (2018) HMMER web server: 2018 update. Nucleic Acids Res 46:W200–W204. https://doi.org/10.1093/nar/gky448
Simão FA, Waterhouse RM, Ioannidis P et al (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinform 31:3210–3212. https://doi.org/10.1093/bioinformatics/btv351
Emms DM, Kelly S (2018) OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol 20:238. https://doi.org/10.1186/s13059-019-1832-y
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinforma 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Nguyen LT, Schmidt HA, Von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Hoang DT, Chernomor O, Von Haeseler A et al (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Kalyaanamoorthy S, Minh BQ, Wong TKF et al (2017) ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Meng G, Li Y, Yang C, Liu S (2019) MitoZ: a toolkit for animal mitochondrial genome assembly, annotation and visualization. Nucleic Acids Res 47:e63. https://doi.org/10.1093/nar/gkz173
Chernomor O, Von Haeseler A, Minh BQ (2016) Terrace aware data structure for phylogenomic inference from supermatrices. Syst Biol 65:997–1008. https://doi.org/10.1093/sysbio/syw037
Katsuma S, Hirota K, Matsuda-Imai N et al (2022) A Wolbachia factor for male killing in lepidopteran insects. Nat Commun 13:6764. https://doi.org/10.1038/s41467-022-34488-y
Ote M, Ueyama M, Yamamoto D (2016) Wolbachia protein TomO targets nanos mRNA and restores germ stem cells in Drosophila Sex-lethal mutants. Curr Biol 26:2223–2232. https://doi.org/10.1016/j.cub.2016.06.054
Lindsey ARI, Werren JH, Richards S, Stouthamer R (2016) Comparative genomics of a parthenogenesis-inducing Wolbachia symbiont. G3 Genes. Genomes, Genet 6:2113–2123. https://doi.org/10.1534/g3.116.028449
Gschloessl B, Audiot P, Nidelet S et al (2020) Complete mitogenome data from a European specimen of Ostrinia scapulalis (Walker, 1859) (Lepidoptera, Pyraloidea, Crambidae, Pyraustinae). Data Br 33:106427. https://doi.org/10.1016/j.dib.2020.106427
Luo Q, Zhou N, Yang Z (2021) Complete mitochondrial genome of Ostrinia kasmirica (Lepidoptera: Crambidae). Mitochondrial DNA Part B 6:2316–2318. https://doi.org/10.1080/23802359.2021.1950058
Comandatore F, Cordaux R, Bandi C et al (2015) Supergroup C Wolbachia, mutualist symbionts of filarial nematodes, have a distinct genome structure. Open Biol 5:150099. https://doi.org/10.1098/rsob.150099
Newton ILG, Clark ME, Kent BN et al (2016) Comparative genomics of two closely related Wolbachia with different reproductive effects on hosts. Genome Biol Evol 8:1526–1542. https://doi.org/10.1093/gbe/evw096
Al-Khodor S, Price CT, Kalia A, Abu Kwaik Y (2010) Functional diversity of ankyrin repeats in microbial proteins. Trends Microbiol 18:132–139. https://doi.org/10.1016/j.tim.2009.11.004
Ahmed MZ, Li SJ, Xue X et al (2015) The intracellular bacterium Wolbachia uses parasitoid wasps as phoretic vectors for efficient horizontal transmission. PLoS Pathog 11:e1004672. https://doi.org/10.1371/journal.ppat.1004672
Huigens ME, Luck RF, Klaassen RHG et al (2000) Infectious parthenogenesis. Nature 405:178–179. https://doi.org/10.1038/35012066
Sakai R, Fukuzawa M, Nakano R et al (2009) Alternative suppression of transcription from two desaturase genes is the key for species-specific sex pheromone biosynthesis in two Ostrinia moths. Insect Biochem Mol Biol 39:62–67. https://doi.org/10.1016/j.ibmb.2008.10.001
Li J, Wang ZY, Bourguet D, He KL (2013) Wolbachia infection in populations of Ostrinia furnacalis: diversity, prevalence, phylogeny and evidence for horizontal transmission. J Integr Agric 12:283–295. https://doi.org/10.1016/S2095-3119(13)60227-0
Acknowledgements
We thank the Institute for Sustainable Agro-ecosystem Services, The University of Tokyo, for collecting Ostrinia moths. Computations were partially performed on the NIG supercomputer at ROIS National Institute of Genetics.
Funding
Open access funding provided by The University of Tokyo. This work was supported by Grants-in-Aid for Scientific Research on Innovative Areas “Spectrum of the Sex: a continuity of phenotypes between female and male” (17H06431) to S.K. and T.K., Grants-in-Aid for Scientific Research (A) (22H00366) to S.K., and U TOKYO Sustainable Agriculture Education Program to T.M.
Author information
Authors and Affiliations
Contributions
T.M., T.K., and S.K. designed and performed the experiments. T.M. and H.H. performed bioinformatics analysis. T.F. performed sex pheromone analysis. T.M. and S.K. wrote the manuscript with intellectual input from all authors. S.K. supervised the project.
Corresponding author
Ethics declarations
Competing Interest
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Muro, T., Hikida, H., Fujii, T. et al. Two Complete Genomes of Male-Killing Wolbachia Infecting Ostrinia Moth Species Illuminate Their Evolutionary Dynamics and Association with Hosts. Microb Ecol 86, 1740–1754 (2023). https://doi.org/10.1007/s00248-023-02198-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-023-02198-7