Abstract
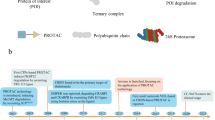
The canonical function of the endoplasmic reticulum-associated degradation (ERAD) system is to enforce quality control among membrane-associated proteins by targeting misfolded secreted, intra-organellar, and intramembrane proteins for degradation. However, increasing evidence suggests that ERAD additionally functions in maintaining appropriate levels of a subset of membrane-associated proteins. In this ‘quantity control’ capacity, ERAD responds to environmental cues to regulate the proteasomal degradation of specific ERAD substrates according to cellular need. In this review, we discuss in detail seven proteins that are targeted by the ERAD quantity control system. Not surprisingly, ERAD-mediated protein degradation is a key regulatory feature of a variety of ER-resident proteins, including HMG-CoA reductase, cytochrome P450 3A4, IP3 receptor, and type II iodothyronine deiodinase. In addition, the ERAD quantity control system plays roles in maintaining the proper stoichiometry of multi-protein complexes by mediating the degradation of components that are produced in excess of the limiting subunit. Perhaps somewhat unexpectedly, recent evidence suggests that the ERAD quantity control system also contributes to the regulation of plasma membrane-localized signaling receptors, including the ErbB3 receptor tyrosine kinase and the GABA neurotransmitter receptors. For these substrates, a proportion of the newly synthesized yet properly folded receptors are diverted for degradation at the ER, and are unable to traffic to the plasma membrane. Given that receptor abundance or concentration within the plasma membrane plays key roles in determining signaling efficiency, these observations may point to a novel mechanism for modulating receptor-mediated cellular signaling.
Similar content being viewed by others
References
Adeli K, Macri J, Mohammadi A, Kito M, Urade R, Cavallo D (1997) Apolipoprotein B is intracellularly associated with an ER-60 protease homologue in HepG2 cells. J Biol Chem 272:22489–22494
Altier C, Garcia-Caballero A, Simms B, You H, Chen L, Walcher J, Tedford HW, Hermosilla T, Zamponi GW (2011) The Cavβ subunit prevents RFP2-mediated ubiquitination and proteasomal degradation of L-type channels. Nat Neurosci 14:173–180
Alzayady KJ, Wojcikiewicz RJ (2005) The role of Ca2 + in triggering inositol 1,4,5-trisphosphate receptor ubiquitination. Biochem J 392:601–606
Alzayady KJ, Panning MM, Kelley GG, Wojcikiewicz RJ (2005) Involvement of the p97-Ufd1-Npl4 complex in the regulated endoplasmic reticulum-associated degradation of inositol 1,4,5-trisphosphate receptors. J Biol Chem 280:34530–34537
Amin DN, Campbell MR, Moasser MM (2010) The role of HER3, the unpretentious member of the HER family, in cancer biology and cancer therapeutics. Semin Cell Dev Biol 21:944–950
Arrojo e Drigo R, Bianco AC (2011) Type 2 deiodinase at the crossroads of thyroid hormone action. Int J Biochem Cell Biol 43:1432–1441
Arrojo e Drigo R, Fonseca TL, Werneck-de-Castro JP, Bianco AC (2013a) Role of the type 2 iodothyronine deiodinase (D2) in the control of thyroid hormone signaling. Biochim Biophys Acta 7:29
Arrojo e Drigo R, Egri P, Jo S, Gereben B, Bianco AC (2013b) The type II deiodinase is retrotranslocated to the cytoplasm and proteasomes via p97/Atx3 complex. Mol Endocrinol 27:2105–2115
Barel MT, Hassink GC, van Voorden S, Wiertz EJ (2006) Human cytomegalovirus-encoded US2 and US11 target unassembled MHC class I heavy chains for degradation. Mol Immunol 43:1258–1266
Baselga J, Swain SM (2009) Novel anticancer targets: revisiting ERBB2 and discovering ERBB3. Nat Rev Cancer 9:463–475
Benke D (2010) Mechanisms of GABAB receptor exocytosis, endocytosis, and degradation. Adv Pharmacol 58:93–111
Bettler B, Kaupmann K, Mosbacher J, Gassmann M (2004) Molecular structure and physiological functions of GABA(B) receptors. Physiol Rev 84:835–867
Borecky J, Vercesi AE (2005) Plant uncoupling mitochondrial protein and alternative oxidase: energy metabolism and stress. Biosci Rep 25:271–286
Boren J, Olin K, Lee I, Chait A, Wight TN, Innerarity TL (1998) Identification of the principal proteoglycan-binding site in LDL. A single-point mutation in apo-B100 severely affects proteoglycan interaction without affecting LDL receptor binding. J Clin Invest 101:2658–2664
Bosanac I, Alattia JR, Mal TK, Chan J, Talarico S, Tong FK, Tong KI, Yoshikawa F, Furuichi T, Iwai M, Michikawa T, Mikoshiba K, Ikura M (2002) Structure of the inositol 1,4,5-trisphosphate receptor binding core in complex with its ligand. Nature 420:696–700
Botero D, Gereben B, Goncalves C, De Jesus LA, Harney JW, Bianco AC (2002) Ubc6p and ubc7p are required for normal and substrate-induced endoplasmic reticulum-associated degradation of the human selenoprotein type 2 iodothyronine monodeiodinase. Mol Endocrinol 16:1999–2007
Bowery NG, Bettler B, Froestl W, Gallagher JP, Marshall F, Raiteri M, Bonner TI, Enna SJ (2002) International Union of Pharmacology. XXXIII. Mammalian gamma-aminobutyric acid(B) receptors: structure and function. Pharmacol Rev 54:247–264
Brent GA (2012) Mechanisms of thyroid hormone action. J Clin Invest 122:3035–3043
Brodsky JL, Skach WR (2011) Protein folding and quality control in the endoplasmic reticulum: recent lessons from yeast and mammalian cell systems. Curr Opin Cell Biol 23:464–475
Burr ML, Cano F, Svobodova S, Boyle LH, Boname JM, Lehner PJ (2011) HRD1 and UBE2J1 target misfolded MHC class I heavy chains for endoplasmic reticulum-associated degradation. Proc Natl Acad Sci USA 108:2034–2039
Cannon B, Nedergaard J (2011) Nonshivering thermogenesis and its adequate measurement in metabolic studies. J Exp Biol 214:242–253
Cao J, Wang J, Qi W, Miao HH, Ge L, DeBose-Boyd RA, Tang JJ, Li BL, Song BL (2007) Ufd1 is a cofactor of gp78 and plays a key role in cholesterol metabolism by regulating the stability of HMG-CoA reductase. Cell Metab 6:115–128
Carraway KL 3rd (2010) E3 ubiquitin ligases in ErbB receptor quantity control. Semin Cell Dev Biol 21:936–943
Carvalho P, Stanley AM, Rapoport TA (2010) Retrotranslocation of a misfolded luminal ER protein by the ubiquitin-ligase Hrd1p. Cell 143:579–591
Chen Y, Le Caherec F, Chuck SL (1998) Calnexin and other factors that alter translocation affect the rapid binding of ubiquitin to apoB in the Sec61 complex. J Biol Chem 273:11887–11894
Choi K, Kim H, Kang H, Lee SY, Lee SJ, Back SH, Lee SH, Kim MS, Lee JE, Park JY, Kim J, Kim S, Song JH, Choi Y, Lee S, Lee HJ, Kim JH, Cho S (2014) Regulation of diacylglycerol acyltransferase 2 protein stability by gp78-associated endoplasmic-reticulum-associated degradation. FEBS J 281:3048–3060
Chun KT, Bar-Nun S, Simoni RD (1990) The regulated degradation of 3-hydroxy-3-methylglutaryl-CoA reductase requires a short-lived protein and occurs in the endoplasmic reticulum. J Biol Chem 265:22004–22010
Curcio-Morelli C, Zavacki AM, Christofollete M, Gereben B, de Freitas BC, Harney JW, Li Z, Wu G, Bianco AC (2003) Deubiquitination of type 2 iodothyronine deiodinase by von Hippel-Lindau protein-interacting deubiquitinating enzymes regulates thyroid hormone activation. J Clin Invest 112:189–196
Darom A, Bening-Abu-Shach U, Broday L (2010) RNF-121 is an endoplasmic reticulum-membrane E3 ubiquitin ligase involved in the regulation of beta-integrin. Mol Biol Cell 21:1788–1798
DeBose-Boyd RA (2008) Feedback regulation of cholesterol synthesis: sterol-accelerated ubiquitination and degradation of HMG CoA reductase. Cell Res 18:609–621
Dentice M, Bandyopadhyay A, Gereben B, Callebaut I, Christoffolete MA, Kim BW, Nissim S, Mornon JP, Zavacki AM, Zeold A, Capelo LP, Curcio-Morelli C, Ribeiro R, Harney JW, Tabin CJ, Bianco AC (2005) The Hedgehog-inducible ubiquitin ligase subunit WSB-1 modulates thyroid hormone activation and PTHrP secretion in the developing growth plate. Nat Cell Biol 7:698–705
Diamonti AJ, Guy PM, Ivanof C, Wong K, Sweeney C, Carraway KL 3rd (2002) An RBCC protein implicated in maintenance of steady-state neuregulin receptor levels. Proc Natl Acad Sci U S A 99:2866–2871
Dixon JL, Furukawa S, Ginsberg HN (1991) Oleate stimulates secretion of apolipoprotein B-containing lipoproteins from Hep G2 cells by inhibiting early intracellular degradation of apolipoprotein B. J Biol Chem 266:5080–5086
Dunn R, Hicke L (2001) Multiple roles for Rsp5p-dependent ubiquitination at the internalization step of endocytosis. J Biol Chem 276:25974–25981
Engelman JA, Cantley LC (2006) The role of the ErbB family members in non-small cell lung cancers sensitive to epidermal growth factor receptor kinase inhibitors. Clin Cancer Res 12:4372s–4376s
Erickson SL, O’Shea KS, Ghaboosi N, Loverro L, Frantz G, Bauer M, Lu LH, Moore MW (1997) ErbB3 is required for normal cerebellar and cardiac development: a comparison with ErbB2-and heregulin-deficient mice. Development 124:4999–5011
Faouzi S, Medzihradszky KF, Hefner C, Maher JJ, Correia MA (2007) Characterization of the physiological turnover of native and inactivated cytochromes P450 3A in cultured rat hepatocytes: a role for the cytosolic AAA ATPase p97? Biochemistry 46:7793–7803
Finley D (2009) Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu Rev Biochem 78:477–513
Fisher EA, Lapierre LR, Junkins RD, McLeod RS (2008) The AAA-ATPase p97 facilitates degradation of apolipoprotein B by the ubiquitin-proteasome pathway. J Lipid Res 49:2149–2160
Fisher EA, Khanna NA, McLeod RS (2011) Ubiquitination regulates the assembly of VLDL in HepG2 cells and is the committing step of the apoB-100 ERAD pathway. J Lipid Res 52:1170–1180
Fry WH, Simion C, Sweeney C, Carraway KL 3rd (2011) Quantity control of the ErbB3 receptor tyrosine kinase at the endoplasmic reticulum. Mol Cell Biol 31:3009–3018
Fujita K (2004) Food-drug interactions via human cytochrome P450 3A (CYP3A). Drug Metabol Drug Interact 20:195–217
Furuichi T, Mikoshiba K (1995) Inositol 1,4,5-trisphosphate receptor-mediated Ca2+ signaling in the brain. J Neurochem 64:953–960
Furukawa S, Sakata N, Ginsberg HN, Dixon JL (1992) Studies of the sites of intracellular degradation of apolipoprotein B in Hep G2 cells. J Biol Chem 267:22630–22638
Gardner RG, Shearer AG, Hampton RY (2001) In vivo action of the HRD ubiquitin ligase complex: mechanisms of endoplasmic reticulum quality control and sterol regulation. Mol Cell Biol 21:4276–4291
Garza RM, Sato BK, Hampton RY (2009) In vitro analysis of Hrd1p-mediated retrotranslocation of its multispanning membrane substrate 3-hydroxy-3-methylglutaryl (HMG)-CoA reductase. J Biol Chem 284:14710–14722
Gassmann M, Bettler B (2012) Regulation of neuronal GABA(B) receptor functions by subunit composition. Nat Rev Neurosci 13:380–394
Gereben B, Goncalves C, Harney JW, Larsen PR, Bianco AC (2000) Selective proteolysis of human type 2 deiodinase: a novel ubiquitin-proteasomal mediated mechanism for regulation of hormone activation. Mol Endocrinol 14:1697–1708
Ginsberg HN, Fisher EA (2009) The ever-expanding role of degradation in the regulation of apolipoprotein B metabolism. J Lipid Res 50:2
Glickman MH, Ciechanover A (2002) The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev 82:373–428
Goldstein JL, Brown MS (1990) Regulation of the mevalonate pathway. Nature 343:425–430
Gregersen N, Bross P, Vang S, Christensen JH (2006) Protein misfolding and human disease. Annu Rev Genom Hum Genet 7:103–124
Guo X, Shen S, Song S, He S, Cui Y, Xing G, Wang J, Yin Y, Fan L, He F, Zhang L (2011) The E3 ligase Smurf1 regulates Wolfram syndrome protein stability at the endoplasmic reticulum. J Biol Chem 286:18037–18047
Gusarova V, Caplan AJ, Brodsky JL, Fisher EA (2001) Apoprotein B degradation is promoted by the molecular chaperones hsp90 and hsp70. J Biol Chem 276:24891–24900
Hamburger AW (2008) The role of ErbB3 and its binding partners in breast cancer progression and resistance to hormone and tyrosine kinase directed therapies. J Mammary Gland Biol Neoplasia 13:225–233
Hampton RY, Rine J (1994) Regulated degradation of HMG-CoA reductase, an integral membrane protein of the endoplasmic reticulum, in yeast. J Cell Biol 125:299–312
Hartman IZ, Liu P, Zehmer JK, Luby-Phelps K, Jo Y, Anderson RG, DeBose-Boyd RA (2010) Sterol-induced dislocation of 3-hydroxy-3-methylglutaryl coenzyme A reductase from endoplasmic reticulum membranes into the cytosol through a subcellular compartment resembling lipid droplets. J Biol Chem 285:19288–19298
Hatakeyama J, Wald JH, Rafidi H, Cuevas A, Sweeney C, Carraway KL 3rd (2016) The ER structural protein Rtn4A stabilizes and enhances signaling through the receptor tyrosine kinase ErbB3. Sci Signal 9:rar65
Hayashi T, Hayashi E, Fujimoto M, Sprong H, Su TP (2012) The lifetime of UDP-galactose:ceramide galactosyltransferase is controlled by a distinct endoplasmic reticulum-associated degradation (ERAD) regulated by sigma-1 receptor chaperones. J Biol Chem 287:43156–43169
Hegde RS, Ploegh HL (2010) Quality and quantity control at the endoplasmic reticulum. Curr Opin Cell Biol 22:437–446
Hetz C (2012) The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Biol 13:89–102
Higo T, Hattori M, Nakamura T, Natsume T, Michikawa T, Mikoshiba K (2005) Subtype-specific and ER lumenal environment-dependent regulation of inositol 1,4,5-trisphosphate receptor type 1 by ERp44. Cell 120:85–98
Horimoto S, Ninagawa S, Okada T, Koba H, Sugimoto T, Kamiya Y, Kato K, Takeda S, Mori K (2013) The unfolded protein response transducer ATF6 represents a novel transmembrane-type endoplasmic reticulum-associated degradation substrate requiring both mannose trimming and SEL1L protein. J Biol Chem 288:31517–31527
Hrizo SL, Gusarova V, Habiel DM, Goeckeler JL, Fisher EA, Brodsky JL (2007) The Hsp110 molecular chaperone stabilizes apolipoprotein B from endoplasmic reticulum-associated degradation (ERAD). J Biol Chem 282:32665–32675
Hughes EA, Hammond C, Cresswell P (1997) Misfolded major histocompatibility complex class I heavy chains are translocated into the cytoplasm and degraded by the proteasome. Proc Natl Acad Sci USA 94:1896–1901
Hughes BT, Nwosu CC, Espenshade PJ (2009) Degradation of sterol regulatory element-binding protein precursor requires the endoplasmic reticulum-associated degradation components Ubc7 and Hrd1 in fission yeast. J Biol Chem 284:20512–20521
Hynes NE, MacDonald G (2009) ErbB receptors and signaling pathways in cancer. Curr Opin Cell Biol 21:177–184
Inoue S, Bar-Nun S, Roitelman J, Simoni RD (1991) Inhibition of degradation of 3-hydroxy-3-methylglutaryl-coenzyme A reductase in vivo by cysteine protease inhibitors. J Biol Chem 266:13311–13317
Ishikura S, Weissman AM, Bonifacino JS (2010) Serine residues in the cytosolic tail of the T-cell antigen receptor alpha-chain mediate ubiquitination and endoplasmic reticulum-associated degradation of the unassembled protein. J Biol Chem 285:23916–23924
Jackson-Fisher AJ, Bellinger G, Breindel JL, Tavassoli FA, Booth CJ, Duong JK, Stern DF (2008) ErbB3 is required for ductal morphogenesis in the mouse mammary gland. Breast Cancer Res 10:18
Jaenicke LA, Brendebach H, Selbach M, Hirsch C (2011) Yos9p assists in the degradation of certain nonglycosylated proteins from the endoplasmic reticulum. Mol Biol Cell 22:2937–2945
Jo Y, Debose-Boyd RA (2010) Control of cholesterol synthesis through regulated ER-associated degradation of HMG CoA reductase. Crit Rev Biochem Mol Biol 45:185–198
Jo Y, Lee PC, Sguigna PV, DeBose-Boyd RA (2011) Sterol-induced degradation of HMG CoA reductase depends on interplay of two Insigs and two ubiquitin ligases, gp78 and Trc8. Proc Natl Acad Sci USA 108:20503–20508
Jo Y, Hartman IZ, DeBose-Boyd RA (2013) Ancient ubiquitous protein-1 mediates sterol-induced ubiquitination of 3-hydroxy-3-methylglutaryl CoA reductase in lipid droplet-associated endoplasmic reticulum membranes. Mol Biol Cell 24:169–183
Johnson PR, Swanson R, Rakhilina L, Hochstrasser M (1998) Degradation signal masking by heterodimerization of MATalpha2 and MATa1 blocks their mutual destruction by the ubiquitin-proteasome pathway. Cell 94:217–227
Khan MT, Joseph SK (2003) Proteolysis of type I inositol 1,4,5-trisphosphate receptor in WB rat liver cells. Biochem J 375:603–611
Kikkert M, Doolman R, Dai M, Avner R, Hassink G, van Voorden S, Thanedar S, Roitelman J, Chau V, Wiertz E (2004) Human HRD1 is an E3 ubiquitin ligase involved in degradation of proteins from the endoplasmic reticulum. J Biol Chem 279:3525–3534
Kim SM, Acharya P, Engel JC, Correia MA (2010) Liver cytochrome P450 3A ubiquitination in vivo by gp78/autocrine motility factor receptor and C terminus of Hsp70-interacting protein (CHIP) E3 ubiquitin ligases: physiological and pharmacological relevance. J Biol Chem 285:35866–35877
Kliewer SA, Umesono K, Mangelsdorf DJ, Evans RM (1992) Retinoid X receptor interacts with nuclear receptors in retinoic acid, thyroid hormone and vitamin D3 signalling. Nature 355:446–449
Komander D, Rape M (2012) The ubiquitin code. Annu Rev Biochem 81:203–229
Laney JD, Hochstrasser M (2003) Ubiquitin-dependent degradation of the yeast Mat(alpha)2 repressor enables a switch in developmental state. Genes Dev 17:2259–2270
Lecureux LW, Wattenberg BW (1994) The regulated degradation of a 3-hydroxy-3-methylglutaryl-coenzyme A reductase reporter construct occurs in the endoplasmic reticulum. J Cell Sci 107:2635–2642
Leichner GS, Avner R, Harats D, Roitelman J (2009) Dislocation of HMG-CoA reductase and Insig-1, two polytopic endoplasmic reticulum proteins, en route to proteasomal degradation. Mol Biol Cell 20:3330–3341
Lemoine NR, Barnes DM, Hollywood DP, Hughes CM, Smith P, Dublin E, Prigent SA, Gullick WJ, Hurst HC (1992) Expression of the ERBB3 gene product in breast cancer. Br J Cancer 66:1116–1121
Lerner M, Corcoran M, Cepeda D, Nielsen ML, Zubarev R, Ponten F, Uhlen M, Hober S, Grander D, Sangfelt O (2007) The RBCC gene RFP2 (Leu5) encodes a novel transmembrane E3 ubiquitin ligase involved in ERAD. Mol Biol Cell 18:1670–1682
Liang JS, Kim T, Fang S, Yamaguchi J, Weissman AM, Fisher EA, Ginsberg HN (2003) Overexpression of the tumor autocrine motility factor receptor Gp78, a ubiquitin protein ligase, results in increased ubiquitinylation and decreased secretion of apolipoprotein B100 in HepG2 cells. J Biol Chem 278:23984–23988
Liao M, Faouzi S, Karyakin A, Correia MA (2006) Endoplasmic reticulum-associated degradation of cytochrome P450 CYP3A4 in Saccharomyces cerevisiae: further characterization of cellular participants and structural determinants. Mol Pharmacol 69:1897–1904
Lu JP, Wang Y, Sliter DA, Pearce MM, Wojcikiewicz RJ (2011) RNF170 protein, an endoplasmic reticulum membrane ubiquitin ligase, mediates inositol 1,4,5-trisphosphate receptor ubiquitination and degradation. J Biol Chem 286:24426–24433
Lukacs GL, Verkman AS (2012) CFTR: folding, misfolding and correcting the DeltaF508 conformational defect. Trends Mol Med 18:81–91
Magadan JG, Perez-Victoria FJ, Sougrat R, Ye Y, Strebel K, Bonifacino JS (2010) Multilayered mechanism of CD4 downregulation by HIV-1 Vpu involving distinct ER retention and ERAD targeting steps. PLoS Pathog 6:1000869
Meacham GC, Patterson C, Zhang W, Younger JM, Cyr DM (2001) The Hsc70 co-chaperone CHIP targets immature CFTR for proteasomal degradation. Nat Cell Biol 3:100–105
Meigs TE, Roseman DS, Simoni RD (1996) Regulation of 3-hydroxy-3-methylglutaryl-coenzyme A reductase degradation by the nonsterol mevalonate metabolite farnesol in vivo. J Biol Chem 271:7916–7922
Meusser B, Hirsch C, Jarosch E, Sommer T (2005) ERAD: the long road to destruction. Nat Cell Biol 7:766–772
Mikoshiba K (2007) IP3 receptor/Ca2 + channel: from discovery to new signaling concepts. J Neurochem 102:1426–1446
Morito D, Hirao K, Oda Y, Hosokawa N, Tokunaga F, Cyr DM, Tanaka K, Iwai K, Nagata K (2008) Gp78 cooperates with RMA1 in endoplasmic reticulum-associated degradation of CFTRDeltaF508. Mol Biol Cell 19:1328–1336
Moriyama T, Wada M, Urade R, Kito M, Katunuma N, Ogawa T, Simoni RD (2001) 3-hydroxy-3-methylglutaryl coenzyme A reductase is sterol-dependently cleaved by cathepsin L-type cysteine protease in the isolated endoplasmic reticulum. Arch Biochem Biophys 386:205–212
Murray BP, Correia MA (2001) Ubiquitin-dependent 26S proteasomal pathway: a role in the degradation of native human liver CYP3A4 expressed in Saccharomyces cerevisiae? Arch Biochem Biophys 393:106–116
Nagai A, Kadowaki H, Maruyama T, Takeda K, Nishitoh H, Ichijo H (2009) USP14 inhibits ER-associated degradation via interaction with IRE1alpha. Biochem Biophys Res Commun 379:995–1000
Narjoz C, Marisa L, Imbeaud S, Paris A, Delacroix H, Beaune P, De Waziers I (2009) Genomic consequences of cytochrome P450 2C9 overexpression in human hepatoma cells. Chem Res Toxicol 22:779–787
Nedergaard J, Golozoubova V, Matthias A, Asadi A, Jacobsson A, Cannon B (2001) UCP1: the only protein able to mediate adaptive non-shivering thermogenesis and metabolic inefficiency. Biochim Biophys Acta 1:82–106
Ogu CC, Maxa JL (2000) Drug interactions due to cytochrome P450. Proc 13:421–423
Olofsson SO, Boren J (2012) Apolipoprotein B secretory regulation by degradation. Arterioscler Thromb Vasc Biol 32:1334–1338
Omura T, Kaneko M, Okuma Y, Orba Y, Nagashima K, Takahashi R, Fujitani N, Matsumura S, Hata A, Kubota K, Murahashi K, Uehara T, Nomura Y (2006) A ubiquitin ligase HRD1 promotes the degradation of Pael receptor, a substrate of Parkin. J Neurochem 99:1456–1469
Ortiz de Montellano PR (2005) Cytochrome P450: structure, mechanism, and biochemistry, 3rd edn. Kluwer, New York
Ota T, Gayet C, Ginsberg HN (2008) Inhibition of apolipoprotein B100 secretion by lipid-induced hepatic endoplasmic reticulum stress in rodents. J Clin Invest 118:316–332
Pabarcus MK, Hoe N, Sadeghi S, Patterson C, Wiertz E, Correia MA (2009) CYP3A4 ubiquitination by gp78 (the tumor autocrine motility factor receptor, AMFR) and CHIP E3 ligases. Arch Biochem Biophys 483:66–74
Pan M, Cederbaum AI, Zhang YL, Ginsberg HN, Williams KJ, Fisher EA (2004) Lipid peroxidation and oxidant stress regulate hepatic apolipoprotein B degradation and VLDL production. J Clin Invest 113:1277–1287
Pearce MM, Wang Y, Kelley GG, Wojcikiewicz RJ (2007) SPFH2 mediates the endoplasmic reticulum-associated degradation of inositol 1,4,5-trisphosphate receptors and other substrates in mammalian cells. J Biol Chem 282:20104–20115
Pearce MM, Wormer DB, Wilkens S, Wojcikiewicz RJ (2009) An endoplasmic reticulum (ER) membrane complex composed of SPFH1 and SPFH2 mediates the ER-associated degradation of inositol 1,4,5-trisphosphate receptors. J Biol Chem 284:10433–10445
Petras SF, Lindsey S, Harwood HJ Jr (1999) HMG-CoA reductase regulation: use of structurally diverse first half-reaction squalene synthetase inhibitors to characterize the site of mevalonate-derived nonsterol regulator production in cultured IM-9 cells. J Lipid Res 40:24–38
Printsev I, Yen L, Sweeney C, Carraway KL 3rd (2014) Oligomerization of the Nrdp1 E3 ubiquitin ligase is necessary for efficient autoubiquitination but not ErbB3 ubiquitination. J Biol Chem 289:8570–8578
Qiu W, Kohen-Avramoglu R, Rashid-Kolvear F, Au CS, Chong TM, Lewis GF, Trinh DK, Austin RC, Urade R, Adeli K (2004) Overexpression of the endoplasmic reticulum 60 protein ER-60 downregulates apoB100 secretion by inducing its intracellular degradation via a nonproteasomal pathway: evidence for an ER-60-mediated and pCMB-sensitive intracellular degradative pathway. Biochemistry 43:4819–4831
Qiu W, Kohen-Avramoglu R, Mhapsekar S, Tsai J, Austin RC, Adeli K (2005) Glucosamine-induced endoplasmic reticulum stress promotes ApoB100 degradation: evidence for Grp78-mediated targeting to proteasomal degradation. Arterioscler Thromb Vasc Biol 25:571–577
Ravid T, Kreft SG, Hochstrasser M (2006) Membrane and soluble substrates of the Doa10 ubiquitin ligase are degraded by distinct pathways. EMBO J 25:533–543
Ravuri C, Svineng G, Huseby NE (2013) Differential regulation of γ-glutamyltransferase and glutamate cysteine ligase expression after mitochondrial uncoupling: γ-glutamyltransferase is regulated in an Nrf2- and NFκB-independent manner. Free Radic Res 47:394–403
Riethmacher D, Sonnenberg-Riethmacher E, Brinkmann V, Yamaai T, Lewin GR, Birchmeier C (1997) Severe neuropathies in mice with targeted mutations in the ErbB3 receptor. Nature 389:725–730
Roberts BJ (1997) Evidence of proteasome-mediated cytochrome P-450 degradation. J Biol Chem 272:9771–9778
Rochat B (2005) Role of cytochrome P450 activity in the fate of anticancer agents and in drug resistance: focus on tamoxifen, paclitaxel and imatinib metabolism. Clin Pharmacokinet 44:349–366
Rotin D, Kumar S (2009) Physiological functions of the HECT family of ubiquitin ligases. Nat Rev Mol Cell Biol 10:398–409
Rutledge AC, Qiu W, Zhang R, Kohen-Avramoglu R, Nemat-Gorgani N, Adeli K (2009) Mechanisms targeting apolipoprotein B100 to proteasomal degradation: evidence that degradation is initiated by BiP binding at the N terminus and the formation of a p97 complex at the C terminus. Arterioscler Thromb Vasc Biol 29:579–585
Rutledge AC, Qiu W, Zhang R, Urade R, Adeli K (2013) Role of cysteine-protease CGHC motifs of ER-60, a protein disulfide isomerase, in hepatic apolipoprotein B100 degradation. Arch Biochem Biophys 537:104–112
Saeed M, Suzuki R, Watanabe N, Masaki T, Tomonaga M, Muhammad A, Kato T, Matsuura Y, Watanabe H, Wakita T, Suzuki T (2011) Role of the endoplasmic reticulum-associated degradation (ERAD) pathway in degradation of hepatitis C virus envelope proteins and production of virus particles. J Biol Chem 286:37264–37273
Saliba RS, Michels G, Jacob TC, Pangalos MN, Moss SJ (2007) Activity-dependent ubiquitination of GABA(A) receptors regulates their accumulation at synaptic sites. J Neurosci 27:13341–13351
Saliba RS, Pangalos M, Moss SJ (2008) The ubiquitin-like protein Plic-1 enhances the membrane insertion of GABAA receptors by increasing their stability within the endoplasmic reticulum. J Biol Chem 283(27):18538–18544
Sato R, Imanaka T, Takatsuki A, Takano T (1990) Degradation of newly synthesized apolipoprotein B-100 in a pre-Golgi compartment. J Biol Chem 265:11880–11884
Shearer AG, Hampton RY (2005) Lipid-mediated, reversible misfolding of a sterol-sensing domain protein. EMBO J 24:149–159
Shimada T, Yamazaki H, Mimura M, Inui Y, Guengerich FP (1994) Interindividual variations in human liver cytochrome P-450 enzymes involved in the oxidation of drugs, carcinogens and toxic chemicals: studies with liver microsomes of 30 Japanese and 30 Caucasians. J Pharmacol Exp Ther 270:414–423
Sibilia M, Kroismayr R, Lichtenberger BM, Natarajan A, Hecking M, Holcmann M (2007) The epidermal growth factor receptor: from development to tumorigenesis. Differentiation 75:770–787
Sniderman A, Couture P, de Graaf J (2010) Diagnosis and treatment of apolipoprotein B dyslipoproteinemias. Nat Rev Endocrinol 6:335–346
Snyder PM (2002) The epithelial Na + channel: cell surface insertion and retrieval in Na+ homeostasis and hypertension. Endocr Rev 23:258–275
Song BL, Javitt NB, DeBose-Boyd RA (2005a) Insig-mediated degradation of HMG CoA reductase stimulated by lanosterol, an intermediate in the synthesis of cholesterol. Cell Metab 1:179–189
Song BL, Sever N, DeBose-Boyd RA (2005b) Gp78, a membrane-anchored ubiquitin ligase, associates with Insig-1 and couples sterol-regulated ubiquitination to degradation of HMG CoA reductase. Mol Cell 19:829–840
Steinsapir J, Harney J, Larsen PR (1998) Type 2 iodothyronine deiodinase in rat pituitary tumor cells is inactivated in proteasomes. J Clin Invest 102:1895–1899
Stern DF (2008) ERBB3/HER3 and ERBB2/HER2 duet in mammary development and breast cancer. J Mammary Gland Biol Neoplasia 13:215–223
Suzuki M, Otsuka T, Ohsaki Y, Cheng J, Taniguchi T, Hashimoto H, Taniguchi H, Fujimoto T (2012) Derlin-1 and UBXD8 are engaged in dislocation and degradation of lipidated ApoB-100 at lipid droplets. Mol Biol Cell 23:800–810
Tanaka RD, Li AC, Fogelman AM, Edwards PA (1986) Inhibition of lysosomal protein degradation inhibits the basal degradation of 3-hydroxy-3-methylglutaryl coenzyme A reductase. J Lipid Res 27:261–273
Tyler RE, Pearce MM, Shaler TA, Olzmann JA, Greenblatt EJ, Kopito RR (2012) Unassembled CD147 is an endogenous endoplasmic reticulum-associated degradation substrate. Mol Biol Cell 23:4668–4678
Vashist S, Ng DT (2004) Misfolded proteins are sorted by a sequential checkpoint mechanism of ER quality control. J Cell Biol 165:41–52
Wang HF, Figueiredo Pereira ME, Correia MA (1999) Cytochrome P450 3A degradation in isolated rat hepatocytes: 26S proteasome inhibitors as probes. Arch Biochem Biophys 365:45–53
Wang Y, Liao M, Hoe N, Acharya P, Deng C, Krutchinsky AN, Correia MA (2009a) A role for protein phosphorylation in cytochrome P450 3A4 ubiquitin-dependent proteasomal degradation. J Biol Chem 284:5671–5684
Wang Y, Pearce MM, Sliter DA, Olzmann JA, Christianson JC, Kopito RR, Boeckmann S, Gagen C, Leichner GS, Roitelman J, Wojcikiewicz RJ (2009b) SPFH1 and SPFH2 mediate the ubiquitination and degradation of inositol 1,4,5-trisphosphate receptors in muscarinic receptor-expressing HeLa cells. Biochim Biophys Acta 11:12
Wang F, Olson EM, Shyng SL (2012a) Role of Derlin-1 protein in proteostasis regulation of ATP-sensitive potassium channels. J Biol Chem 287:10482–10493
Wang Y, Guan S, Acharya P, Liu Y, Thirumaran RK, Brandman R, Schuetz EG, Burlingame AL, Correia MA (2012b) Multisite phosphorylation of human liver cytochrome P450 3A4 enhances Its gp78- and CHIP-mediated ubiquitination: a pivotal role of its Ser-478 residue in the gp78-catalyzed reaction. Mol Cell Proteomics 11:17
Wang X, Yu YY, Myers N, Hansen TH (2013) Decoupling the role of ubiquitination for the dislocation versus degradation of major histocompatibility complex (MHC) class I proteins during endoplasmic reticulum-associated degradation (ERAD). J Biol Chem 288:23295–23306
Webster JM, Tiwari S, Weissman AM, Wojcikiewicz RJ (2003) Inositol 1,4,5-trisphosphate receptor ubiquitination is mediated by mammalian Ubc7, a component of the endoplasmic reticulum-associated degradation pathway, and is inhibited by chelation of intracellular Zn2+. J Biol Chem 278:38238–38246
Wojcikiewicz RJ, Nahorski SR (1991) Chronic muscarinic stimulation of SH-SY5Y neuroblastoma cells suppresses inositol 1,4,5-trisphosphate action. Parallel inhibition of inositol 1,4,5-trisphosphate-induced Ca2+ mobilization and inositol 1,4,5-trisphosphate binding. J Biol Chem 266:22234–22241
Wojcikiewicz RJ, Oberdorf JA (1996) Degradation of inositol 1,4,5-trisphosphate receptors during cell stimulation is a specific process mediated by cysteine protease activity. J Biol Chem 271:16652–16655
Wojcikiewicz RJ, Furuichi T, Nakade S, Mikoshiba K, Nahorski SR (1994) Muscarinic receptor activation down-regulates the type I inositol 1,4,5-trisphosphate receptor by accelerating its degradation. J Biol Chem 269:7963–7969
Wojcikiewicz RJ, Ernst SA, Yule DI (1999) Secretagogues cause ubiquitination and down-regulation of inositol 1, 4,5-trisphosphate receptors in rat pancreatic acinar cells. Gastroenterology 116:1194–1201
Wu T, Zhao F, Gao B, Tan C, Yagishita N, Nakajima T, Wong PK, Chapman E, Fang D, Zhang DD (2014) Hrd1 suppresses Nrf2-mediated cellular protection during liver cirrhosis. Genes Dev 28:708–722
Yamaguchi J, Conlon DM, Liang JJ, Fisher EA, Ginsberg HN (2006) Translocation efficiency of apolipoprotein B is determined by the presence of beta-sheet domains, not pause transfer sequences. J Biol Chem 281:27063–27071
Yen L, Cao Z, Wu X, Ingalla ER, Baron C, Young LJ, Gregg JP, Cardiff RD, Borowsky AD, Sweeney C, Carraway KL 3rd (2006) Loss of Nrdp1 enhances ErbB2/ErbB3-dependent breast tumor cell growth. Cancer Res 66:11279–11286
Zavacki AM, Arrojo EDR, Freitas BC, Chung M, Harney JW, Egri P, Wittmann G, Fekete C, Gereben B, Bianco AC (2009) The E3 ubiquitin ligase TEB4 mediates degradation of type 2 iodothyronine deiodinase. Mol Cell Biol 29:5339–5347
Zemoura K, Benke D (2014) Proteasomal degradation of gamma-aminobutyric acidB receptors is mediated by the interaction of the GABAB2 C terminus with the proteasomal ATPase Rtp6 and regulated by neuronal activity. J Biol Chem 289:7738–7746
Zemoura K, Schenkel M, Acuna MA, Yevenes GE, Zeilhofer HU, Benke D (2013) Endoplasmic reticulum-associated degradation controls cell surface expression of gamma-aminobutyric acid, type B receptors. J Biol Chem 288:34897–34905
Acknowledgments
Salary support for the authors was provided by NIH predoctoral fellowship CA165546 (IP), and by NIH Grants CA123541, CA166412 and CA178681 (KLC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest.
Rights and permissions
About this article
Cite this article
Printsev, I., Curiel, D. & Carraway, K.L. Membrane Protein Quantity Control at the Endoplasmic Reticulum. J Membrane Biol 250, 379–392 (2017). https://doi.org/10.1007/s00232-016-9931-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-016-9931-0