Abstract
Analysis of microRNAs (miRNAs) is important in cancer diagnostics and therapy. Conventional methods used to extract miRNA for analysis are generally time-consuming. A novel approach for rapid and sensitive extraction of miRNAs is urgently need for clinical applications. Herein, a novel strategy based on electrical potential-assisted DNA-RNA hybridization was designed for miRNA extraction. The entire extraction process was accomplished in approximately 3 min, which is much shorter than the commercial adsorption column method, at more than 60 min, or the TRIzol method, at more than 90 min. Additionally, the method offered the advantages of simplicity and specificity during the extraction process by electrical potential-assisted hybridization of single-stranded DNA and RNA. Taking let-7a as an example, satisfactory results were achieved for miRNA extraction in serum, demonstrating the applicability in miRNA nucleic acid amplification.
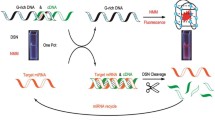
Graphical abstract








Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this manuscript and its supplementary information files.
Abbreviations
- miRNAs:
-
microRNAs
- RT-qPCR:
-
Reverse transcription quantitative polymerase chain reaction
- ASEA:
-
Accelerated strand exchange amplification
- TCEP:
-
Tris(2-carboxyethyl)phosphine hydrochloride
- MCH:
-
6-mercaptohexanol
- CV:
-
Cyclic voltammetry
- EIS:
-
Electrochemical impedance spectroscopy
References
Ban E, Chae D-K, Yoo YS, Song EJ. An improvement of miRNA extraction efficiency in human plasma. Anal Bioanal Chem. 2017;409(27):6397–404. https://doi.org/10.1007/s00216-017-0580-7.
Gu J, Qiao Z, He X, Yu Y, Lei Y, Tang J, Shi H, He D, Wang K. Enzyme-free amplified detection of miRNA based on target-catalyzed hairpin assembly and DNA-stabilized fluorescent silver nanoclusters. Analyst. 2020;145(15):5194–9. https://doi.org/10.1039/D0AN00545B.
Liu H, Tian T, Ji D, Ren N, Ge S, Yan M, Yu J. A graphene-enhanced imaging of microRNA with enzyme-free signal amplification of catalyzed hairpin assembly in living cells. Biosens Bioelectron. 2016;85:909–14. https://doi.org/10.1016/j.bios.2016.06.015.
Lavaee P, Taghdisi SM, Abnous K, Danesh NM, Khayyat LH, Jalalian SH. Fluorescent sensor for detection of miR-141 based on target-induced fluorescence enhancement and PicoGreen. Talanta. 2019;202:349–53. https://doi.org/10.1016/j.talanta.2019.04.084.
Li D, Luo Z, An H, Yang E, Wu M, Huang Z, Duan Y. Poly-adenine regulated DNA density on AuNPs to construct efficient DNA walker for microRNA-21 detection. Talanta. 2020;217:121056. https://doi.org/10.1016/j.talanta.2020.121056.
Kim H, Kang S, Park KS, Park HG. Enzyme-free and label-free miRNA detection based on target-triggered catalytic hairpin assembly and fluorescence enhancement of DNA-silver nanoclusters. Sens Actuators B: Chem. 2018;260:140–5. https://doi.org/10.1016/j.snb.2017.12.137.
Cury JA, Koo H. Extraction and purification of total RNA from Sreptococcus mutans biofilms. Anal Biochem. 2007;365(2):208–14. https://doi.org/10.1016/j.ab.2007.03.021.
Burgess KA, Workman VL, Elsawy MA, Miller AF, Oceandy D, Saiani A. RNA extraction from self-assembling peptide hydrogels to allow qPCR analysis of encapsulated cells. PLoS One. 2018;13(6):e0197517. https://doi.org/10.1371/journal.pone.0197517.
Zhao F, Lee EY, Shin Y. Improved reversible cross-linking-based solid-phase RNA extraction for pathogen diagnostics. Anal Chem. 2018;90(3):1725–33. https://doi.org/10.1021/acs.analchem.7b03493.
Yang H, Liu J, Huang S, Guo T, Deng L, Hua W. Selection and evaluation of novel reference genes for quantitative reverse transcription PCR (qRT-PCR) based on genome and transcriptome data in Brassica napus L. Gene. 2014;538(1):113–22. https://doi.org/10.1016/j.gene.2013.12.057.
Duy J, Koehler JW, Honko AN, Minogue TD. Optimized microRNA purification from TRIzol-treated plasma. BMC Genomics. 2015;16(1):95. https://doi.org/10.1186/s12864-015-1299-5.
Zhu C, Varona M, Anderson JL. Magnetic ionic liquids as solvents for RNA extraction and preservation. ACS Omega. 2020;5(19):11151–9. https://doi.org/10.1021/acsomega.0c01098.
Drula R, Ott LF, Berindan-Neagoe I, Pantel K, Calin GA. MicroRNAs from liquid biopsy derived extracellular vesicles: recent advances in detection and characterization methods. Cancers. 2020;12(8). https://doi.org/10.3390/cancers12082009.
LdS FA, Amorim RG, Scopel WL, Scheicher RH. Controlled current confinement in interfaced 2D nanosensor for electrical identification of DNA. Phys Chem Chem Phys. 2019;21(45):24884–90. https://doi.org/10.1039/c9cp03950c.
Jambrec D, Gebala M, La Mantia F, Schuhmann W. Potential-assisted DNA immobilization as a prerequisite for fast and controlled formation of DNA monolayers. Angew Chem Int Ed Engl. 2015;54(50):15064–8. https://doi.org/10.1002/anie.201506672.
Miranda-Castro R, Palchetti I, de-Los-Santos-Alvarez N (2020) The translational potential of electrochemical DNA-based liquid biopsy. Front Chem 8:143. https://doi.org/10.3389/fchem.2020.00143.
Sanchez-Salcedo R, Miranda-Castro R, de-Los-Santos-Alvarez N, Lobo-Castanon MJ (2021) Dual electrochemical genosensor for early diagnosis of prostate cancer through lncRNAs detection. Biosens Bioelectron 192:113520. https://doi.org/10.1016/j.bios.2021.113520.
Tymoczko J, Schuhmann W, Gebala M. Electrical potential-assisted DNA hybridization. How to mitigate electrostatics for surface DNA hybridization. ACS Appl Mater Interfaces. 2014;6(24):21851–8. https://doi.org/10.1021/am5027902.
Cai W, Peck JR, van der Weide DW, Hamers RJ. Direct electrical detection of hybridization at DNA-modified silicon surfaces. Biosens Bioelectron. 2004;19(9):1013–9. https://doi.org/10.1016/j.bios.2003.09.009.
Liu Q, Ma C, Liu X-P, Wei Y-P, Mao C-J, Zhu J-J. A novel electrochemiluminescence biosensor for the detection of microRNAs based on a DNA functionalized nitrogen doped carbon quantum dots as signal enhancers. Biosens Bioelectron. 2017;92:273–9. https://doi.org/10.1016/j.bios.2017.02.027.
Miao P, Wang B, Meng F, Yin J, Tang Y. Ultrasensitive detection of microRNA through rolling circle amplification on a DNA tetrahedron decorated electrode. Bioconjug Chem. 2015;26(3):602–7. https://doi.org/10.1021/acs.bioconjchem.5b00064.
Tavallaie R, McCarroll J, Le Grand M, Ariotti N, Schuhmann W, Bakker E, Tilley RD, Hibbert DB, Kavallaris M, Gooding JJ. Nucleic acid hybridization on an electrically reconfigurable network of gold-coated magnetic nanoparticles enables microRNA detection in blood. Nat Nanotechnol. 2018;13(11):1066–71. https://doi.org/10.1038/s41565-018-0232-x.
Rafiee-Pour HA, Behpour M, Keshavarz M. A novel label-free electrochemical miRNA biosensor using methylene blue as redox indicator: application to breast cancer biomarker miRNA-21. Biosens Bioelectron. 2016;77:202–7. https://doi.org/10.1016/j.bios.2015.09.025.
Dong J, Chen G, Wang W, Huang X, Peng H, Pu Q, Du F, Cui X, Deng Y, Tang Z. Colorimetric PCR-based microRNA detection method based on small organic dye and single enzyme. Anal Chem. 2018;90(12):7107–11. https://doi.org/10.1021/acs.analchem.8b01111.
Xu H, Zhang S, Ouyang C, Wang Z, Wu D, Liu Y, Jiang Y, Wu ZS. DNA nanostructures from palindromic rolling circle amplification for the fluorescent detection of cancer-related microRNAs. Talanta. 2019;192:175–81. https://doi.org/10.1016/j.talanta.2018.07.090.
Jimenez LA, Gionet-Gonzales MA, Sedano S, Carballo JG, Mendez Y, Zhong W. Extraction of microRNAs from biological matrices with titanium dioxide nanofibers. Anal Bioanal Chem. 2018;410(3):1053–60. https://doi.org/10.1007/s00216-017-0649-3.
Sosnowski RG, Tu E, Butler WF, O'Connell JP, Heller MJ. Rapid determination of single base mismatch mutations in DNA hybrids by direct electric field control. Proc Natl Acad Sci U S A. 1997;94(4):1119–23. https://doi.org/10.1073/pnas.94.4.1119.
Edman CF, Raymond DE, Wu DJ, Tu E, Sosnowski RG, Butler WF, Nerenberg M, Heller MJ. Electric field directed nucleic acid hybridization on microchips. Nucleic Acids Res. 1997;25(24):4907–14. https://doi.org/10.1093/nar/25.24.4907.
Fixe F, Branz HM, Louro N, Chu V, Prazeres DM, Conde JP. Electric-field assisted immobilization and hybridization of DNA oligomers on thin-film microchips. Nanotechnology. 2005;16(10):2061–71. https://doi.org/10.1088/0957-4484/16/10/014.
Yao M, Lv X, Deng Y, Rasheed M. Specific and simultaneous detection of micro RNA 21 and let-7a by rolling circle amplification combined with lateral flow strip. Anal Chim Acta. 2019;1055:115–25. https://doi.org/10.1016/j.aca.2018.12.040.
Wang ZH, Xu CJ. Research Progress of MicroRNA in early detection of ovarian Cancer. Chin Med J. 2015;128(24):3363–70. https://doi.org/10.4103/0366-6999.171459.
Johnson-Buck A, Su X, Giraldez MD, Zhao M, Tewari M, Walter NG. Kinetic fingerprinting to identify and count single nucleic acids. Nat Biotechnol. 2015;33(7):730–2. https://doi.org/10.1038/nbt.3246.
Daneshpour M, Karimi B, Omidfar K. Simultaneous detection of gastric cancer-involved miR-106a and let-7a through a dual-signal-marked electrochemical nanobiosensor. Biosens Bioelectron. 2018;109:197–205. https://doi.org/10.1016/j.bios.2018.03.022.
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, Lao KQ, Livak KJ, Guegler KJ. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005;33(20):e179. https://doi.org/10.1093/nar/gni178.
Zhang Z, Zhang L, Wang Y, Yao J, Wang T, Weng Z, Yang L, Xie G. Ultrasensitive electrochemical biosensor for attomolar level detection of let 7a based on toehold mediated strand displacement reaction circuits and molecular beacon mediated circular strand displacement polymerization. Anal Chim Acta. 2021;1147:108–15. https://doi.org/10.1016/j.aca.2020.12.057.
Li M, Liu M, Ma C, Shi C. Rapid DNA detection and one-step RNA detection catalyzed by Bst DNA polymerase and narrow-thermal-cycling. Analyst. 2020;145(15):5118–22. https://doi.org/10.1039/D0AN00975J.
Faria HAM, Zucolotto V. Label-free electrochemical DNA biosensor for zika virus identification. Biosens Bioelectron. 2019;131:149–55. https://doi.org/10.1016/j.bios.2019.02.018.
Rant U, Arinaga K, Tornow M, Kim YW, Netz RR, Fujita S, Yokoyama N, Abstreiter G. Dissimilar kinetic behavior of electrically manipulated single- and double-stranded DNA tethered to a gold surface. Biophys J. 2006;90(10):3666–71. https://doi.org/10.1529/biophysj.105.078857.
Deraney RN, Schneider L, Tripathi A. Synergistic use of electroosmotic flow and magnetic forces for nucleic acid extraction. Analyst. 2020;145(6):2412–9. https://doi.org/10.1039/c9an02191d.
Guo Y, Bosompem A, Zhong X, Clark T, Shyr Y, Kim AS. A comparison of microRNA sequencing reproducibility and noise reduction using mirVana and TRIzol isolation methods. Int J Comput Biol Drug Design. 2014;7(2-3):102–12. https://doi.org/10.1504/ijcbdd.2014.061642.
Kim Y-K, Yeo J, Kim B, Ha M, Kim VN. Short structured RNAs with low GC content are selectively lost during extraction from a small number of cells. Mol Cell. 2012;46(6):893–5. https://doi.org/10.1016/j.molcel.2012.05.036.
Juan D, Alexe G, Antes T, Liu H, Madabhushi A, Delisi C, Ganesan S, Bhanot G, Liou LS. Identification of a microRNA panel for clear-cell kidney cancer. Urology. 2010;75(4):835–41. https://doi.org/10.1016/j.urology.2009.10.033.
Acknowledgements
The study was supported by grants from the National Key Research and Development Programs of China (2018YFE0113300), National Natural Science Foundation of China (81801264) and Key Project of Shandong Provincial Natural Science Foundation (ZR2020KH030).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The studies were approved by the appropriate ethics committee and were performed in accordance with ethical standards.
Competing interests
The authors declare that there are no competing interests associated with the manuscript.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 429 kb)
Rights and permissions
About this article
Cite this article
Zhao, X., Li, Y., Sun, R. et al. Electrical potential-assisted DNA-RNA hybridization for rapid microRNA extraction. Anal Bioanal Chem 414, 3529–3539 (2022). https://doi.org/10.1007/s00216-022-03979-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-03979-8