Abstract
Glioblastomas (GBs) are one of the most aggressive and invasive intracranial cancers. Recently, it has been postulated that, among other factors, the hedgehog (HH) pathway may be a key factor in this phenomenon. Moreover, it has been reported that small-size silver nanoparticles (AgNPs) are characterized by a high cytotoxic effect towards GBs. However, their effect on the sonic hedgehog (SHH) pathway has never been demonstrated in any cancer cells. Therefore, the aim of the present study was to evaluate the impact of the anti-proliferative properties of 5-nm AgNPs on the SHH pathway in the GB cell line (U-87MG) in vitro. The results showed a time- and dose-dependent decrease in the metabolic activity in the U-87MG cells treated with AgNPs, with IC50 reaching 30.41 and 21.16 µg/mL after 24 h and 48 h, respectively, followed by an increase in the intracellular reactive oxygen species (ROS) level. The co-treatment of the cells with AgNPs and Robotnikinin (SHH inhibitor) abolished and/or strengthened the effect of AgNPs, especially on the SHH mRNA levels and on the PCNA, PTCH1, Gli1, and SUFU protein levels. Interestingly, no changes in the level of ERK1/2, Akt, and SRC kinase protein expression were detected, suggesting a direct impact of AgNPs and/or ROS on the inhibition of the canonical SHH pathway. However, more studies are needed due to the increase in the mTOR protein expression after the treatment of the cells with AgNPs, as in the Robotnikinin treatment. In conclusion, small-size AgNPs are able to inhibit the proliferation of GB cells in vitro by suppressing the canonical SHH pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glioblastomas (GBs) are glioma-derived cancers with a poor survival rate, estimated at approx. 1 year. Moreover, according to the World Health Organization (WHO), GBs are responsible for 2% of all the intracranial tumors nowadays. This phenomenon is linked with many aspects, such as the high selectivity of the blood–brain barrier (BBB) or the presence of the skull (Du et al. 2021; Li et al. 2016a). Moreover, GBs are characterized by high malignancy with enormous proliferation and migration potential resulting in the invasiveness of such cancer cells (Kim 2013). Among many pro-proliferative pathways in GBs, literature data show the pivotal role of the phosphoinositide 3-kinase/protein kinase B (PI3K/Akt), Janus kinase-signal transducer and activator of transcription (JAK-STAT), and mitogen-activated protein kinase/extracellular signal-regulated kinase (MAPK/ERK) pathways as key factors in the adhesive and invasive properties of such cancer cells (Ou et al. 2021; Ramaswamy et al. 2019; Li et al. 2016b). Although new anti-cancer strategies based on the inhibition of the above-mentioned pathways are being developed, the results obtained also in clinical trials are not satisfactory (Li et al. 2016b). As shown inter alia by Day et al., GBs are able to bypass the blocked proliferation-related pathways, which is one of the causes of the insufficiency of novel anti-cancer approaches (Day et al. 2020). Interestingly, the hedgehog pathway (HH) has recently been identified as another key factor in the aggressiveness of GBs, as described inter alia by Hung et al. (Hung et al. 2020).
HH is a conservative intracellular pathway, which is crucial in embryo development, cell division, and proliferation processes (Carballo et al. 2018). Physiologically, three activating ligands of HH have been described—sonic hedgehog (SHH), Indian hedgehog (IHH), and desert hedgehog (DHH). They are able to abolish the inhibitory effect of patched 1 and/or 2 (PTCH1/2) transmembrane proteins, which leads to activation of the smoothened (SMO) protein (Gergues et al. 2021). Subsequently, the signal is transduced intracellularly (with engagement of the SUFU protein), resulting in the activation of the zinc finger protein (Gli1), a well-described oncogene, which stimulates cells to proliferate (Zhu and Lo 2010). Interestingly, alterations in the HH pathway have been linked with development of high-grade GBs as well as increased cancer invasiveness (Takezaki et al. 2011; Wang et al. 2022). Many efforts have been made to use the HH pathway as a target in the anti-GB treatment, e.g., the inhibition of the activity of the SMO protein, which was shown by Bissey et al. to have failed during clinical trials (Bissey et al. 2020). This phenomenon has been linked with the overexpression of HH ligands, mainly SHH, by GBs, which is based on the auto- and paracrine regulation abolishing the HH inhibition at the SMO level (Bissey et al. 2020). Interestingly, a study has linked the ability of silver nanoparticles (AgNPs) to suppress the ATP-binding transporter (ABC), whose over-activation is a well-established feature of the multidrug-resistance (MDR) in cancer cells (Kovács et al. 2016).
In accordance with the definition used worldwide, AgNPs are structures with a size ranging between 1 to 100 nm characterized by high reactivity and internalization into the cell, resulting from their shape and size (Perde-Schrepler et al. 2019). AgNPs are usually described as pro-oxidative factors with an ability to impair the redox homeostasis by releasing high amounts of silver ions (Ag+), which in consequence leads to oxidative stress and apoptosis (Ullah et al. 2020). Interestingly, this phenomenon has been observed in both normal and cancer cells in vitro. However, Liang et al. have shown that AgNPs can act as potential sensitizers in temozolomide-resistant glioma cells (U-251) in vitro (Liang et al. 2017). Similarly, Liu et al. have proved that AgNPs enhance the anti-proliferative effect of radiotherapy in the U-251 glioma cell line (Liu et al. 2018a). Moreover, the toxic effect of these NPs has been reported repeatedly inter alia in human breast, lung, tongue, skin, brain, and hepatic cancer cell lines in vitro (Perde-Schrepler et al. 2019; Skóra et al. 2022, 2023; Hepokur et al. 2019; Faedmaleki et al. 2014). The aforementioned conclusions are promising, given the postulated ability of AgNPs to cross the BBB (Dan et al. 2015; Khan et al. 2019). However, none of the studies have determined the interaction between these NPs and the HH pathway, which is crucial due to its potential engagement in the failure of the current anti-cancer treatment. Additionally, human U-87MG cells are frequently used as a model for testing the anti-cancer properties of AgNPs in GBs due to inter alia their aggressiveness and proliferation properties; hence, these cells were chosen as a model in the present study (Louca et al. 2019).
The aim of this study was to determine the impact of 5-nm AgNPs on the SHH pathway in a glioblastoma cell line (U-87MG) in vitro. The SHH and its role in the anti-proliferative properties of AgNPs in this type of cancer was assessed as well as the potential mechanism of action of such NPs in comparison to Robotnikinin (a selective inhibitor of the SHH pathway). The metabolic activity, intracellular ROS level, and mRNA and protein expression were determined to understand the role of the SHH pathway in the AgNP cytotoxicity potential in GBs.
Materials and methods
Reagents
Phosphate buffer saline (PBS) and Dulbecco's Modified Eagle Medium (DMEM) were purchased from Corning (Corning, USA). Trypsin, resazurin sodium salt penicillin,, 2′,7′-dichlorodihydrofluorescein diacetate (H2DCF-DA), streptomycin, ethanol, ammonium persulfate (APS), methanol, N,N,N′,N′-tetramethyl ethylenediamine (TEMED), acrylamide/bisacrylamide, 30%, 37.5:1 ratio, glycine, Tris Base, Tris–HCl, sodium chloride (NaCl), silver nanoparticles (PVP-stabilized) with a small size (5 nm)—AgNPs, bovine serum albumin (BSA), sodium dodecyl sulfate (SDS) and Bradford reagent were purchased from Merck (Darmstadt, Germany). The High-Capacity cDNA Reverse Transcription Kit, Restore™ Western Blot Stripping Buffer, primers and TaqMan® probes complementary to sequences, encoding the GAPDH (Hs02758991_g1) and SHH (Hs00179843_m1), and goat anti-mouse and goat anti-rabbit HRP-conjugated antibodies were obtained from ThermoFisher (Waltham, USA). Fetal bovine serum (FBS), Fast Probe qPCR Master Mix (2x), Universal RNA Purification Kit, Perfect Tricolor Protein Ladder, and RIPA buffer were purchased from EURx (Gdańsk, Poland). The PVDF membrane, anti-GAPDH, anti-CAT1, and anti-SOD1 mouse primary antibodies were purchased from Santa Cruz Biotechnology (Santa Cruz, USA). Anti-PTCH1, anti-Gli1, anti-SHH, anti-SUFU, anti-Akt, anti-mTOR, anti-SRC, anti-ERK1/2, and anti-PCNA primary antibodies were obtained from ABClonal (Düsseldorf, Germany). Robotnikinin (cat. 13204) was purchased from Cayman Chemicals (Ann Arbor, USA). The PVP-stabilized AgNPs were chosen based on their high stability and low aggregation properties in aqueous solutions, comparing to, e.g., citrate-stabilized AgNPs proved previously in the literature report (Tejamaya et al. 2012).
Cell culture and treatment
The glioblastoma cell line (U-87MG, HTB-14™) was purchased from the American Type Culture Collection (ATCC). Cells were grown in DMEM with 10% FBS supplemented with 0.1% penicillin/streptomycin at 37℃ with 5% CO2 until reaching confluency. Afterwards, the U-87MG cells were trypsinized and seeded on 96-well plates (for resazurin reduction and H2DCF-DA assays), 12-well plates (for RT-qPCR), and a ⌀100 mm culture dish (for Western Blot) at the density of 4 × 103 cell/well, 1.2 × 105 cell/well, or 2.2 × 106 cell/well, respectively. Next, the cells were treated with increasing concentrations of AgNPs in a concentration range from 1 ng/mL to 100 µg/mL for 24 h and 48 h. Moreover, in the case of the RT-qPCR and Western Blot methods, the cells were treated with effective but non-lethal concentrations of the tested compounds (chosen based on the dose–response assays), namely 1 µg/mL of AgNPs and 10 µM of Robotnikinin for 6 h and 24 h (for RT-qPCR) or 24 h and 48 h (for Western Blot).
Resazurin reduction assay
The assay was performed as described by Skóra et al. w/o modifications (Skóra et al. 2023). After the 24-h and 48-h treatment of the U-87MG cells with certain compounds (as described in subsection Cell culture and treatment), the measurement of fluorescence was performed at respective excitation and emission wavelength, using a microplate reader (FilterMax F5).
Intracellular ROS level
The ROS level was quantified using 2′,7′-dichlorodihydrofluorescein diacetate (H2DCF-DA) as in Piechowiak et al. w/o modifications (Piechowiak et al. 2021). The measurement of fluorescence was performed at respective excitation and emission wavelength using a microplate reader (FilterMax F5).
Confocal microscopy
The Calcein-AM and Hoechst 33342 staining was used in this study to determine the ability of tested AgNPs to affect the morphology of cell and nucleus (e.g., apoptotic-like) in tested U-87MG cells. The method was performed according to Skóra et al. (Skóra et al. 2023). The cells were seeded on a culture dish (⌀35 mm) at the density of 1 × 105 cells/dish and sub-cultured for 24 h. Subsequently, the medium was exchanged to fresh one containing 1 µg/mL or 100 µg/mL of AgNPs. After 48 h, the cells were washed twice with warm PBS and the staining medium (serum-free), containing the 10 µM of Hoechst 33342, and 10 µM of Calcein-AM was applied for few min. Afterwards, fluorescence-based visualization was performed at certain excitation and emission wavelengths using a confocal microscope with a laser scanning module (ZEISS LSM700).
RT-qPCR
The RT-qPCR analysis was performed as in Szychowski et al. w/o modifications (Szychowski and Gmiński 2019). 1 µg/mL of AgNPs, 10 µM of Robotnikinin, or 1 µg/mL of AgNPs and 10 µM of Robotnikinin were applied in the experiment for 6 h (primary gene response) and 24 h (secondary gene response). Primers and TaqMan® probes, specific for sequences, encoding the GAPDH and SHH genes were applied. The results were expressed as an average fold (avg. fold) calculated based on the ΔΔCt obtained using the threshold value (Ct) for each sample during the exponential phase of the analyzed genes. GAPDH was used as a reference gene.
Western Blot
The Western Blot analysis was performed as in Skóra et al. without modifications (Skóra et al. 2023). 1 µg/mL of AgNPs, 10 µM of Robotnikinin, or 1 µg/mL of AgNPs and 10 µM of Robotnikinin were applied in the experiment for 24 h and 48 h. The specific primary and secondary HRP-conjugated antibodies were used in the dilutions described in the table below (Table 1). Moreover, the Restore™ Western Blot Stripping Buffer was used as described by the producer to reprobe the PVDF membranes with other primary antibodies with minor modifications (ThermoFisher). Briefly, after the enhanced chemiluminescence-based detection, the membranes were washed once with TBS for 5 min with shaking, followed by 20 min. incubation at 45 °C with shaking in the aforementioned stripping buffer. Subsequently, the buffer was removed, and the membrane was washed three times with TBS at RT for 5 min, followed by 1-h non-specific side blocking with 1% of BSA in TBST. Next, the blocking solution was removed, and the membrane was reprobed with another primary antibody o/n at 4 °C. The detection was performed using the secondary HRP-conjugated antibodies as described in the above-cited paper.
Statistical analyses
The data were expressed as means ± SD (standard deviations) of six (n = 6) or three (n = 3) repetitions of the experiments (specified in the caption of the graphs). The data were then used in the one-way analysis of variance (ANOVA) with Tukey’s post hoc test and denoted as *, **, *** for p < 0.05, p < 0.001, or p < 0.001, respectively, compared to control cells. The means denoted as # or $ were statistically different at p < 0.05 between certain groups (marked on the graphs).
Results
Metabolic activity and ROS production
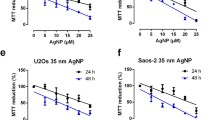
The U-87MG cells treated with 10 µg/mL, 50 µg/mL, and 100 µg/mL of AgNPs for 24 h were characterized by a 34.75%, 54.96%, and 72.44% decrease in the metabolic activity, respectively, compared to the control (Fig. 1A). In turn, after the treatment of the cells with 1 µg/mL, 10 µg/mL, 50 µg/mL, and 100 µg/mL of AgNPs, the metabolic activity decreased by 18.74%, 34.35%, 71.12%, and 76.38%, respectively, compared to the control (Fig. 1B). Similarly, the IC50 values increased during the time of the AgNPs’ exposure, reaching 30.41 µg/mL and 21.16 µg/mL after 24 h and 48 h, respectively (Fig. 1C).
Metabolic activity (A–B) and the ROS production level (D–F) in the U-87MG cells treated with AgNPs in the concentration range from 1 ng/mL to 100 µg/mL for 6 h (D), 24 h (A, E), and 48 h (B, F). The calculated IC50 values for the respective time treatments shown in the right panel (C) were calculated based on the resazurin reduction measurement results. The means ± SD denoted as *, **, and *** are statistically different from the control at p < 0.05, p < 0.001, and p < 0.001, respectively
After the 6-h treatment of the U-87MG cells with 100 ng/mL and 1 µg/mL of AgNPs, a 15.56% and 10.73% increase in the ROS production was observed, respectively, compared to the control, while the 10 µg/mL, 50 µg/mL, and 100 µg/mL concentrations of these NPs decreased the ROS level by 11.74%, 45.34%, and 62.60%, respectively, compared to the control (Fig. 1D). After the 24-h treatment, the ROS production in the cells treated with 100 ng/mL and 1 µg/mL of AgNPs increased by 24.56% and 30.19%, respectively, compared to the control (Fig. 1E). In turn, the 10 µg/mL, 50 µg/mL, and 100 µg/mL concentrations of AgNPs induced a 13.79%, 55.81%, and 66.90% decrease in this parameter in the U-87MG cells, respectively, compared to the control (Fig. 1E). Similarly, the cells treated with 100 ng/mL and 1 µg/mL of AgNPs for 48 h were characterized by a 14.22% and 28.21% increase in the intracellular ROS level, respectively, compared to the control (Fig. 1E). On the other hand, a 19.04%, 53.14%, and 66.01% decrease in this parameter was observed, compared to the control, respectively (Fig. 1E).
Co-treatment of cells
The U-87MG cells treated with AgNPs, Robotnikinin, or co-treated with AgNPs/Robotnikinin were characterized by no significant changes in the metabolic activity after 24 h (Fig. 2A).
Metabolic activity of the U-87MG cells treated with 1 μg/mL of AgNPs, 10 μM of Robotnikinin, and/or co-treated with AgNPs/Robotnikinin for 24 h (A) and 48 h (B). The means ± SD denoted as *, **, and *** are statistically different from the control at p < 0.05, p < 0.001, and p < 0.001, respectively. #denotes statistical differences between certain groups at p < 0.05
In turn, after 48 h, a 14.46% decrease in the metabolic activity was observed in the AgNP-treated cells, compared to the control, while the cells co-treated with AgNPs/Robotnikinin showed a 53.00% increase in this parameter, compared to the control (Fig. 2B). Additionally, a 67.46% statistically different effect between AgNPs- and AgNP/Robotnikinin-treated cells was observed (Fig. 2B). The apoptotic-like changes in nuclei after treatment with AgNPs have been observed (Fig. 3).
SHH mRNA expression
The U-87MG cells treated for 6 h with AgNPs and Robotnikinin were characterized by a 27.05% and 33.60% decrease in the SHH mRNA expression, respectively, compared to the control (Fig. 4A). In turn, a 22.61% increase in the expression of this gene was observed in cells treated with AgNPs/Robotnikinin, compared to the control (Fig. 4A). Moreover, a 49.69% statistically different SHH gene expression was observed in the AgNP-treated cells, compared to the AgNP/Robotnikinin-treated cells (Fig. 4B). After 24 h, a significant 24.06% and 21.13% decrease in the SHH mRNA expression was noted in the U-87MG cells treated with AgNPs and AgNPs/Robotnikinin, respectively, compared to the control (Fig. 4B). No significant changes in the expression of this gene were observed after the treatment of the tested cells with Robotnikinin (Fig. 4B).
SHH mRNA expression in the U-87MG cells treated with AgNPs, Robotnikinin, and AgNPs/Robotnikinin for 6 h (A) and 24 h (B). The means ± SD (standard deviations) denoted as *, **, and *** are statistically different, compared to the control cells at p < 0.05, p < 0.01, and p < 0.001, respectively. The data marked as # or $ are statistically different between certain groups at p < 0.05
Protein expression level
After the 24-h U-87MG treatment with Robotnikinin, the ERK1/2 protein expression increased by 8.86%, compared to the control, while the cells co-treated with AgNPs/Robotnikinin showed a 7.19% decrease in the expression of this protein, compared to the control (Fig. 5A). Moreover, the effect on the AgNP/Robotnikinin-treated cells was statistically lower by 10.77%, compared to the AgNP-treated cells (Fig. 5A). In turn, after 48 h, the ERK1/2 protein expression in the U-87MG cells treated with Robotnikinin and AgNPs/Robotnikinin was decreased by 25.71% and 28.99%, respectively, compared to the control (Fig. 5C). The effect of the AgNP/Robotnikinin co-treatment on the ERK1/2 protein expression was 30.55% lower than in the AgNP-treated cells (Fig. 5C).
ERK1/2 (A, C), SHH (B, D), PCNA (E, G), and SRC (F, H) protein expression after the treatment of the U-87MG cells with 1 μg/mL of AgNPs, 10 μM of Robotnikinin, and AgNPs/Robotnikinin for 24 h (A, C, E, G) and 48 h (B, D, F, H). The means ± SD (standard deviations) denoted as *, **, and *** are statistically different, compared to the control cells at p < 0.05, p < 0.01, and p < 0.001, respectively. The data marked as # and $ are statistically different between certain groups at p < 0.05
The U-87MG cells treated with AgNPs for 24 h were characterized by a 34.94% increase in the SHH protein expression, compared to the control (Fig. 5B). Conversely, 52.88% and 69.78% decrease in the expression of this protein was observed in the cells treated with Robotnikinin and AgNPs/Robotnikinin, respectively, compared to the control (Fig. 4B). A 104.74% statistically different effect between the AgNP- and AgNP/Robotnikinin-treated cells was noticed (Fig. 5B). After 48 h, the SHH protein expression increased by 197.62% in the AgNP-treated U-87MG cells, compared to the control, while the decrease in the AgNP/Robotnikinin-treated cells reached 54.68%, compared to the control (Fig. 5D). This effect differed by 252.30% from that in the AgNP-treated cells (Fig. 5D).
After 24 h, no significant changes in the PCNA protein expression were observed in the U-87MG cells (Fig. 5E). In turn, after 48 h, the cells treated with AgNPs, Robotnikinin, and AgNPs/Robotnikinin were characterized by an 11.80%, 32.69%, and 56.52% decrease in the PCNA protein expression, respectively, compared to the control (Fig. 5G). The effect of the AgNP treatment on the PCNA protein expression was statistically higher (by 44.72%) than in the AgNP/Robotnikinin-treated cells (Fig. 5G).
The U-87MG cells co-treated with AgNPs/Robotnikinin for 24 h were characterized by a 27.29% decrease in the SRC protein expression, compared to the control (Fig. 5F). In turn, the 48-h exposure of the tested cells to AgNPs/Robotnikinin decreased the expression of this protein by 36.62%, compared to the control (Fig. 5H). This effect differed by 39.91% from that in the AgNP-treated cells (Fig. 5H).
The CAT1 protein expression after the treatment of the U-87MG cells with AgNPs, Robotnikinin, and AgNPs/Robotnikinin was significantly increased by 14.75%, 25.99%, and 41.33%, respectively, compared to the control (Fig. 6A). Moreover, a significant 26.58% increase in the expression of this protein was observed in the cells treated with AgNPs/Robotnikinin, compared to the AgNP-treated cells (Fig. 6A). In turn, after 48 h, the cells were characterized by an 18.90% and 16.25% decrease in the CAT1 protein expression in the variants with AgNPs and Robotnikinin, respectively (Fig. 6C). The observed effect in the AgNP-treated cells was statistically different (by 15.14%), compared to the AgNP/Robotnikinin-treated cells (Fig. 6C).
CAT1 (A, C), mTOR (B, D), Akt (E, G), PTCH1 (F, H), SOD1 (I, K), Gli1 (J, L), and SUFU (M, N) protein expression after the treatment of the U-87MG cells with 1 μg/mL of AgNPs, 10 μM of Robotnikinin, and AgNPs/Robotnikinin for 24 h (A, B, E, F, I, J, M) and 48 h (C, D, G, H, K, L, N). The means ± SD (standard deviations) denotes as *, **, and *** are statistically different, compared to the control cells at p < 0.05, p < 0.01, and p < 0.001, respectively. The data marked as # and $ are statistically different between certain groups at p < 0.05
After 24 h, the U-87MG cells treated with AgNPs, Robotnikinin, and AgNPs/Robotnikinin were characterized by a 22.40%, 13.40%, and 21.43% increase in the mTOR protein expression, respectively, compared to the control (Fig. 6B). In turn, a statistically different 14.50% increase in the AgNP-treated cells was observed after 48 h, compared to the control (Fig. 6D).
The Akt protein expression was decreased by 19.17% and 30.51% in the Robotnikinin and AgNP/Robotnikinin-treated cells after 24 h, respectively, compared to the control (Fig. 6E). The effect of the AgNP/Robotnikinin co-treatment in the U-87MG cells was statistically different than in the AgNP-treated cells (33.19%) (Fig. 6E). In turn, after the 48-h treatment, the cells treated with AgNPs, Robotnikinin, and AgNPs/Robotnikinin were characterized by a 20.15%, 10.49%, and 23.24% decrease in the Akt protein expression, respectively, compared to the control (Fig. 6G).
After 24 h, the PTCH1 protein expression was increased by 10.03% and 29.68% in the U-87MG cells treated with AgNPs and Robotnikinin, respectively, compared to the control (Fig. 6F). In contrast, a 31.07% decrease in the expression of this protein was observed in the AgNP/Robotnikinin co-treated cells, compared to the control (Fig. 6F). This effect was by 41.11% different from that in the AgNP-treated cells (Fig. 6F). In turn, after the 48-h treatment of the cells with AgNPs, Robotnikinin, and AgNPs/Robotnikinin, the PTCH1 protein expression increased by 40.27%, 261.69%, and 58.06%, respectively, compared to the control (Fig. 6H). The effect caused by AgNPs was statistically lower (by 17.79%) than in the AgNP/Robotnikinin-treated cells (Fig. 6H).
The SOD1 protein expression was increased by 29.98%, 37.63%, and 48.33% in the AgNP-, Robotnikinin-, and AgNP/Robotnikinin-treated cells, respectively, compared to the control, after 24 h (Fig. 6I). Similarly, a 30.34% and 38.22% increase in the expression of this protein was observed in the cells treated with Robotnikinin and AgNPs/Robotnikinin, respectively, compared to the control, while a 47.96% decrease in the SOD1 protein expression in the AgNP-treated cells was observed, compared to the control, after 48 h (Fig. 6K). The effect caused by AgNPs and AgNPs/Robotnikinin differed by 86.18% (Fig. 6K).
After 24 h, the Gli1 protein expression in the U-87MG cells treated with AgNPs was decreased by 15.55%, compared to the control, while the AgNPs/Robotnikinin co-treatment caused a 10.42% increase in the expression of this protein, compared to the control (Fig. 6J). The observed effect was statistically different between these groups, and the difference was estimated at 25.97% (Fig. 6J). In turn, after 48 h, the Gli1 protein expression increased by 11.98%, 66.25%, and 43.36% in all the tested groups exposed to AgNPs, Robotnikinin, and AgNPs/Robotnikinin, respectively, compared to the control (Fig. 6L). Additionally, the effect caused by AgNPs and AgNPs/Robotnikinin differed between these two groups by 31.38% (Fig. 6L).
The SUFU protein expression after the 24-h treatment of the U-87MG cells with AgNPs, Robotnikinin, and AgNPs/Robotnikinin decreased by 23.76%, 28.54%, and 42.98%, respectively, compared to the control (Fig. 6 M). The effect between AgNPs and AgNP/Robotnikinin-treated cells differed by 19.23% (Fig. 6M). In turn, after the 48-h treatment, the cells treated with AgNPs were characterized by a 38.76% decrease in the SUFU protein expression, compared to the control (Fig. 6N). This effect was statistically different (by 43.75%) than in the AgNP/Robotnikinin-treated cells (Fig. 6N).
Discussion
AgNPs are well-established cytotoxic factors, whose biological activity is strictly related to their high pro-oxidative properties leading to redox imbalance and, in consequence, oxidative stress (Maurer and Meyer 2016). The present results showed a significant increase in the intracellular ROS level only after 6 h of treatment, which was strengthened with the time of exposure (especially in the concentration range between 100 ng/mL and 1 µg/mL). These findings are similar to the results reported by Onodera et al., who proved the ability of 1-nm AgNPs applied at the 5 µg/mL concentration to increase the ROS level in BALB/3T3 A31-1–1 cells (Onodera et al. 2015). Furthermore, Kang et al. showed that exposure to a low 2 µg/mL concentration of AgNPs (2.3 nm) significantly increased ROS production in the murine dendritic cell line (DC2.4) (Kang et al. 2012). In turn, in our study, a massive decrease in the ROS production level was observed in some of the tested concentrations of AgNPs—10 µg/mL, 50 µg/mL, and 100 µg/mL, which was presumably caused by their high cytotoxic effect. Indeed, the resazurin reduction assay showed that, in the microgram concentrations, AgNPs were able to decrease the metabolic activity by approx. 50%, resulting in IC50 values of 30.41 µg/mL and 21.16 µg/mL in the 24-h and 48-h AgNP exposure variants, respectively. Moreover, our further studies indicated a decrease in the PCNA protein expression (well-established proliferation marker) after 48 h in the U-87MG treated with AgNPs, which proves the aforementioned phenomenon. Similarly, our previous study showed that small-size AgNPs were able to decrease KI67 protein expression in human normal fibroblasts (BJ), lung adenocarcinoma cells (A549), and squamous tongue carcinoma (SCC-15) (Skóra et al. 2022). Moreover, Chairuangkitti et al. proved that AgNPs (100 nm) were able to arrest A549 cells in the sub-G1 cell cycle phase with a decrease in PCNA protein expression, which is associated with the ROS-mediated pathway (Chairuangkitti et al. 2013). The toxicity of AgNPs was shown inter alia by Liang et al. in U-87MG cells and by Salazar-García et al. in rat GB cells (C6) (Liang et al. 2017; Salazar-García et al. 2020). Additionally, our results of confocal microscopy analysis showed the ability of tested AgNPs to induce apoptotic-like changes in the nucleus in tested cell line. Therefore, the results obtained in this study are consistent with the current state of knowledge and show the high anti-cancer potential of AgNPs to be used in GB therapy; however, this has never been tested in the context of the HH pathway.
It has been reported that ROS may affect the SHH–NOX4–HIF1α pathway in cerebellar progenitor cells (Eyrich et al. 2019). Therefore, Robotnikinin was used in the next part of this study. This compound has been identified as a selective inhibitor of SHH, whose mechanism of action is related to the blocking of the ability of SHH to bind to PTCH1 (Stanton and Peng 2010). The co-treatment of the U-87MG cells with AgNPs and Robotnikinin abolished the cytotoxic effect of AgNPs, which may suggest some correlation between the SHH pathway and the mechanism of action of AgNPs. However, due to the rather overall non-specific character of the resazurin reduction assay, the SHH mRNA expression was assessed. Our data showed that AgNPs had an impact on the SHH gene expression in a similar way as Robotnikinin, reducing the mRNA expression of the gene. A massive decrease in the expression of certain genes is often correlated with an increase in the expression of the corresponding protein, which in turn acts as a negative regulator of the expression of this gene (Ivanov 2019). Indeed, the measured SHH protein expression showed an increase in this parameter in both time intervals after the treatment with AgNPs. As cited above, AgNPs act mainly by inducing high amounts of ROS, which is consistent with results reported by Dai et al., who showed activation of the SHH pathway in H2O2-treated primary rat cortical neurons, resulting in a time-dependent increase in the SHH protein expression as part of the protective mechanism (Dai et al. 2011). On the other hand, Kim et al. concluded that oxidative stress inhibited SHH-induced osteogenic differentiation of multipotent bone marrow stromal cells (Kim et al. 2010). Therefore, it can be hypothesized that AgNPs exert the inhibitory effect on the SHH pathway in a ROS-dependent manner. Interestingly, in our study, Robotnikinin and/or AgNPs/Robotnikinin showed an opposite effect, i.e., a decrease in the SHH protein expression, which may suggest a different or excluding mechanism of action between these two compounds.
The predicted ROS-dependent mechanism of AgNPs and its effect on the SHH pathway is consistent with the results of the SOD1 and CAT1 protein expression, which are well-established markers of oxidative stress (Patlolla et al. 2009). After the treatment of the U-87MG cells with AgNPs for 24 h, the SOD1 and CAT1 protein expression was significantly increased, followed by a decrease in the expression of these proteins after 48 h, which indicated the ability of the tested NPs to induce oxidative stress. A similar tendency was shown for Robotnikinin. Although the pro-oxidative properties of AgNPs in GB (DBTRG-05MG) cells were shown inter alia by Akyuva and Nazıroğlu, the direct effect of Robotnikinin on oxidative stress induction has never been tested (Akyuva and Nazıroğlu 2023). However, recently (2022), Karadağ and Başbinar have shown an inhibitory effect of Robotnikinin and Vismodegib (SHH inhibitor at the SMO level) on U-87MG proliferation and invasiveness, which is consistent with the results shown in this study (Karadağ and Basbinar 2022). Nevertheless, to investigate whether AgNPs affect the canonical or non-canonical HH pathway, the expression of specific downstream proteins was measured.
As shown by the literature data, the canonical HH pathway is based on the mutual interaction between PTCH1, SMO, SUFU, and Gli-1 proteins (Teperino et al. 2014). Both Gli1 and PTCH1 play a crucial role in the HH pathway, which is related to the binding of SHH to PTCH1 and transducing this signal intracellularly with Gli1, upregulating certain pro-proliferative genes (Cohen et al. 2015). Interestingly, Ji et al. have proved that the inhibition of the SHH pathway by cyclopamine (at the SMO level) results in increased DNA damage and downregulation of certain downstream proteins in the SHH pathway, inter alia Gli1 and PTCH1 (Ji et al. 2012). Similar data were shown in our study. The tested AgNPs were able to decrease the Gli1 protein expression (transcriptional factor of the SHH pathway) after 24 h, in contrast to Robotnikinin, which did not affect this parameter. Nevertheless, an inverse effect was observed in the co-treated cells. Moreover, the PTCH1 protein expression was significantly increased after the treatment with AgNPs for 24 h and 48 h. Furthermore, this effect was statistically different after the co-treatment of the U-87MG cells with AgNPs and Robotnikinin. These results are consistent with those reported by Wang et al., who showed the ability of cyclopamine to decrease the PTCH1, SMO, and Gli1 mRNA expression in human pancreatic cancer (PANC-1) and human colon cancer (HT-29, LoVo, and HCT-116) cell lines in vitro (Wang et al. 2014). On the other hand, some SHH inhibitors seem to act in a cell-specific manner, which was confirmed by the results obtained by Carballo et al., who proved an effect of cyclopamine on PTCH1 protein expression in GBM95 cells but not in GBM02 and GBM03 cells (Carballo et al. 2020). Interestingly, it was shown in the present study that AgNPs were able to decrease the SUFU protein expression in both tested time intervals. SUFU is classified as a regulator of the Gli1-dependent pro-proliferative ability of the HH pathway (Yan et al. 2021). The decrease in SUFU and Gli1 presented in this study proved the cytotoxic properties of AgNPs, shown inter alia by the PCNA protein expression and by the resazurin reduction assay, and are consistent with the results reported by Wang et al. and Liu et al. (Liu et al. 2018a; Wang et al. 2013). However, this study is the first to show that AgNPs can reduce the proliferation of U-87MG cells with engagement of the Gli1 transcriptional factor. Moreover, the results shown above also prove the ability of AgNPs to block the SHH pathway first and later decrease the downstream effectors of this pathway-related protein, leading to a decrease in the SHH mRNA expression, probably by blocking the PTCH1 ability to bind SHH. These results are crucial, given the recent findings of the overexpression of SHH and/or Gli1 in GBs and other cancers, leading to lower survival rates in patients (Budimir et al. 2022; Cui et al. 2010).
It has been reported that the HH pathway can also be activated as a result of the activity of certain kinases, inter alia ERK1/2, Akt, and/or mTOR, i.e., the non-canonical pathway (Wang et al. 2012). The results obtained in this study showed that the tested small-size AgNPs did not affect the ERK1/2 protein expression in any time interval, in contrast to Robotnikinin, which alone and in the co-treatment with the AgNPs reduced the expression of this protein. This is consistent with the results shown by Liu et al., who proved that cyclopamine decreased the proliferation of fibroblast-like synoviocytes by reducing ERK1/2 phosphorylation (Liu et al. 2018b). This also suggests that AgNPs exert an anti-proliferative effect on GBs cells independently of ERK1/2, but rather with engagement of the canonical HH pathway. This is opposite to the results reported by Castiglioni et al., who discovered that the engagement of ERK1/2 and its phosphorylation state were crucial for induction of AgNP cytotoxicity in bladder carcinoma cells (T24) (Castiglioni et al. 2015). Similarly, Rinna et al. showed that, in addition to oxidative stress, AgNPs were able to induce DNA damage, which was strengthened after ERK1/2 blocking in human epithelial embryonic cells (Rinna et al. 2015). The differences between the cited papers and the present results may be an effect of the different cell model used in the studies as well as the higher diameter of AgNPs (20 nm and 35 nm vs. 5 nm tested in this study).
Interestingly, as shown by Katoh in 2009, Gli1 can be activated by the PI3K/Akt pathway, which is related to the ability of Akt to stabilize Gli1 (Katoh 2009). In our study, the tested AgNPs did not exert an impact on the Akt protein expression after the 24-h treatment, in contrast to the effect caused by Robotnikinin. Therefore, it is justified to assume that AgNPs inhibit the proliferation of GBs based via the canonical rather than non-canonical pathway, which supports the hypothesis of the different mechanisms of action between AgNPs and Robotnikinin. Indeed, Adnan et al. demonstrated that small-size AgNPs (6–20 nm) were able to be uptaken in U-251 cells after only 2 h, which proved their ability to induce oxidative stress in a short-time treatment, as shown in this study as well (Adnan et al. 2020). Because our data did not show a decrease in the Akt protein expression 24 h after the AgNP treatment but the impact of the nanoparticles on the SHH mRNA expression was observed after only 6 h, followed by the massive increase in the SHH protein expression, it is justified to assume that the impact of AgNPs on the HH pathway is ERK1/2- and Akt-independent. The further decrease in the Akt protein expression was probably an effect of the AgNP- and Gli1-dependent cytotoxicity. This is consistent with the results shown by Elekofehinti, who proved that AgNPs were able to decrease AKT mRNA protein expression in Wistar rats (Elekofehinti et al. 2021). Moreover, the present results showed an increase in the mTOR protein expression after the treatment of the U-87MG cells with AgNPs, which is consistent with the results reported in the literature inter alia by Li et al. in neuroblastoma cells (SH-SY5Y) or by Chang et al. in mouse hippocampal neurons (HT-22), suggesting the potential of the tested AgNPs to induce autophagy (Li et al. 2019; Chang et al. 2021). However, taking the above results and the cited studies into account, this study is the first to show that small-size AgNPs (5 nm) are able to act as inhibitors of the HH pathway, probably via an ROS-dependent pathway. These interactions subsequently lead to a reduced PTCH1 ability to bind SHH, which results in the suppression of the downstream proteins of the HH pathway, including the Gli1 transcriptional factor. Moreover, the non-canonical activation of HH is not likely to be engaged in the AgNP mechanism of action. The present results also prove the postulated suitability of these NPs in the GB treatment, especially in the context of the potential SHH role in the MDR phenomenon in such cells. Based on the results, the proposed mechanism of tested NPs' action were shown below (Fig. 7). However, more comprehensive studies are needed to fully elucidate the AgNPs’ role in this field.
Proposed mechanism of action of AgNPs in U-87MG cells. After the uptake, AgNPs cause an increase in the cellular ROS level, which subsequently affects the PTCH1 receptor, resulting in a decrease in the ability to bind SHH. Consequently, U-87MG cells are characterized by overexpression of the SHH protein (compensation effect), resulting in a decrease in the SHH mRNA expression (probably by negative feedback). In consequence, the HH downstream proteins (SMO, SUFU, and Gli1) are downregulated, resulting in reduced U-87MG proliferation
Conclusions
The study shows for the first time that AgNPs are able to suppress the HH pathway at the mRNA and protein expression level based on the ROS-dependent pathway in GBs cells. The tested AgNPs were able to decrease the ability of SHH to bind to PTCH1 and downregulate the downstream proteins related to the canonical HH pathway, which resulted in a decrease in the proliferation of the U-87MG cells through suppression of SUFU-Gli1. Moreover, the observed effect was not correlated with the non-canonical HH pathway. The present results prove the potential suitability of small-size AgNPs in anti-cancer therapy, especially in the case of the recently proved Gli1 overexpression in GB cells.
Data availability
The data are available from the corresponding author on reasonable request.
Abbreviations
- ABC:
-
ATP-binding cassette transporter
- AgNPs:
-
Silver nanoparticles
- Ag+ :
-
Silver ions
- BBB:
-
Blood–brain barrier
- DHH:
-
Desert hedgehog
- FBS:
-
Fetal bovine serum
- GBs:
-
Glioblastomas
- H2DCF-DA:
-
2′,7′-Dichlorodihydrofluorescein diacetate
- HH:
-
Hedgehog
- IHH:
-
Indian hedgehog
- NRF2:
-
Nuclear Factor Erythroid 2 Like 2
- Robotnikinin:
-
N-[(4-chlorophenyl)methyl]-5,12-dioxo-2R-phenyl-1-oxa-4-azacyclododec-8E-ene-6S-acetamide
- PCNA:
-
Proliferating cell nuclear antigen
- PTCH1:
-
Protein patched homolog 1
- ROS:
-
Reactive oxygen species
- SHH:
-
Sonic hedgehog
- SMO:
-
Smoothened protein
- SUFU:
-
Suppressor of fused homolog protein
References
Adnan M, Obyedul Kalam Azad M, Madhusudhan A et al (2020) Simple and cleaner system of silver nanoparticle synthesis using kenaf seed and revealing its anticancer and antimicrobial potential. Nanotechnology. https://doi.org/10.1088/1361-6528/ab7d72
Akyuva Y, Nazıroğlu M (2023) Silver nanoparticles potentiate antitumor and oxidant actions of cisplatin via the stimulation of TRPM2 channel in glioblastoma tumor cells. Chem Biol Interact 369:110261. https://doi.org/10.1016/j.cbi.2022.110261
Bissey P-A, Mathot P, Guix C et al (2020) Blocking SHH/Patched interaction triggers tumor growth inhibition through patched-induced apoptosis. Cancer Res 80:1970–1980. https://doi.org/10.1158/0008-5472.CAN-19-1340
Budimir I, Tomasović-Lončarić Č, Kralik K et al (2022) Higher expressions of SHH and AR are associated with a positive receptor status and have impact on survival in a cohort of croatian breast cancer patients. Life 12:1559. https://doi.org/10.3390/life12101559
Carballo GB, Honorato JR, de Lopes GPF, de Spohr TCL, S e, (2018) A highlight on Sonic hedgehog pathway. Cell Commun Signal 16:11. https://doi.org/10.1186/s12964-018-0220-7
Carballo GB, Matias D, Ribeiro JH et al (2020) Cyclopamine sensitizes glioblastoma cells to temozolomide treatment through Sonic hedgehog pathway. Life Sci 257:118027. https://doi.org/10.1016/j.lfs.2020.118027
Castiglioni S, Cazzaniga A, Perrotta C, Maier JAM (2015) Silver nanoparticles-induced cytotoxicity requires ERK activation in human bladder carcinoma cells. Toxicol Lett 237:237–243. https://doi.org/10.1016/j.toxlet.2015.06.1707
Chairuangkitti P, Lawanprasert S, Roytrakul S et al (2013) Silver nanoparticles induce toxicity in A549 cells via ROS-dependent and ROS-independent pathways. Toxicol in Vitro. https://doi.org/10.1016/j.tiv.2012.08.021
Chang X, Wang X, Li J et al (2021) Silver nanoparticles induced cytotoxicity in HT22 cells through autophagy and apoptosis via PI3K/AKT/mTOR signaling pathway. Ecotoxicol Environ Saf 208:111696. https://doi.org/10.1016/j.ecoenv.2020.111696
Cohen M, Kicheva A, Ribeiro A et al (2015) Ptch1 and Gli regulate Shh signalling dynamics via multiple mechanisms. Nat Commun 6:6709. https://doi.org/10.1038/ncomms7709
Cui D, Xu Q, Wang K, Che X (2010) Gli1 is a potential target for alleviating multidrug resistance of gliomas. J Neurol Sci 288:156–166. https://doi.org/10.1016/j.jns.2009.09.006
Dai R-L, Zhu S-Y, Xia Y-P et al (2011) Sonic hedgehog protects cortical neurons against oxidative stress. Neurochem Res 36:67–75. https://doi.org/10.1007/s11064-010-0264-6
Dan M, Xu L, Shao A et al (2015) Silver nanoparticles induce tight junction disruption and astrocyte neurotoxicity in a rat blood–brain barrier primary triple coculture model. Int J Nanomed. https://doi.org/10.2147/IJN.S85265
Day EK, Sosale NG, Xiao A et al (2020) Glioblastoma cell resistance to EGFR and MET inhibition can be overcome via blockade of FGFR-SPRY2 bypass signaling. Cell Rep 30:3383-3396.e7. https://doi.org/10.1016/j.celrep.2020.02.014
Du J, Zhang A, Li J et al (2021) Doxorubicin-induced cognitive impairment: The mechanistic Insights. Front Oncol. https://doi.org/10.3389/fonc.2021.673340
Elekofehinti OO, Oyedokun VO, Iwaloye O et al (2021) Momordica charantia silver nanoparticles modulate SOCS/JAK/STAT and P13K/Akt/PTEN signalling pathways in the kidney of streptozotocin-induced diabetic rats. J Diabetes Metab Disord 20:245–260. https://doi.org/10.1007/s40200-021-00739-w
Eyrich NW, Potts CR, Robinson MH et al (2019) Reactive oxygen species signaling promotes hypoxia-inducible factor 1α stabilization in sonic hedgehog-driven cerebellar progenitor cell proliferation. Mol Cell Biol. https://doi.org/10.1128/MCB.00268-18
Faedmaleki F, Shirazi FH, Salarian AA, et al (2014) Toxicity effect of silver nanoparticles on mice liver primary cell culture and HepG2 cell line. Iran J Pharm Res https://doi.org/10.22037/ijpr.2014.1436
Gergues M, Savanur VH, Prado D, et al (2021) Therapeutic approaches to overcome temozolomide resistance in glioblastoma. In: Glioblastoma Resistance to Chemotherapy: Molecular Mechanisms and Innovative Reversal Strategies. Elsevier, pp 507–545
Hepokur C, Kariper İA, Mısır S et al (2019) Silver nanoparticle/capecitabine for breast cancer cell treatment. Toxicol Vitr. https://doi.org/10.1016/j.tiv.2019.104600
Hung H-C, Liu C-C, Chuang J-Y et al (2020) Inhibition of sonic hedgehog signaling suppresses glioma stem-like cells likely through inducing autophagic cell death. Front Oncol. https://doi.org/10.3389/fonc.2020.01233
Ivanov D (2019) Notch signaling-induced oscillatory gene expression may drive neurogenesis in the developing retina. Front Mol Neurosci. https://doi.org/10.3389/fnmol.2019.00226
Ji H, Miao J, Zhang X et al (2012) Inhibition of sonic hedgehog signaling aggravates brain damage associated with the down-regulation of Gli1, Ptch1 and SOD1 expression in acute ischemic stroke. Neurosci Lett 506:1–6. https://doi.org/10.1016/j.neulet.2011.11.027
Kang K-A, Jung H-Y, Lim J-S (2012) Cell death by polyvinylpyrrolidine-coated silver nanoparticles is mediated by ROS-dependent signaling. Biomol Ther 20:399–405. https://doi.org/10.4062/biomolther.2012.20.4.399
Karadağ A, Başbinar Y (2022) Novel approach to the hedgehog signaling pathway: combined treatment of SMO and PTCH inhibitors. J Basic Clin Heal Sci https://doi.org/10.30621/jbachs.1193720
Katoh (2009) Integrative genomic analyses on GLI1: Positive regulation of GLI1 by Hedgehog-GLI, TGFβ-Smads, and RTK-PI3K-AKT signals, and negative regulation of GLI1 by Notch-CSL-HES/HEY, and GPCR-Gs-PKA signals. Int J Oncol https://doi.org/10.3892/ijo_00000328
Khan AM, Korzeniowska B, Gorshkov V et al (2019) Silver nanoparticle-induced expression of proteins related to oxidative stress and neurodegeneration in an in vitro human blood-brain barrier model. Nanotoxicology 13:221–239. https://doi.org/10.1080/17435390.2018.1540728
Kim Y (2013) Regulation of cell proliferation and migration in glioblastoma: new therapeutic approach. Front Oncol. https://doi.org/10.3389/fonc.2013.00053
Kim W-K, Meliton V, Bourquard N et al (2010) Hedgehog signaling and osteogenic differentiation in multipotent bone marrow stromal cells are inhibited by oxidative stress. J Cell Biochem 111:1199–1209. https://doi.org/10.1002/jcb.22846
Kovács D, Szőke K, Igaz N et al (2016) Silver nanoparticles modulate ABC transporter activity and enhance chemotherapy in multidrug resistant cancer. Nanomed Nanotechnol Biol Med. https://doi.org/10.1016/j.nano.2015.10.015
Li L, Li L, Zhou X et al (2019) Silver nanoparticles induce protective autophagy via Ca 2+ /CaMKKβ/AMPK/mTOR pathway in SH-SY5Y cells and rat brains. Nanotoxicology 13:369–391. https://doi.org/10.1080/17435390.2018.1550226
Li Y, Zheng X, Gong M, Zhang J (2016a) Delivery of a peptide-drug conjugate targeting the blood brain barrier improved the efficacy of paclitaxel against glioma. Oncotarget 7:79401–79407. https://doi.org/10.18632/oncotarget.12708
Li X, Wu C, Chen N, et al (2016b) PI3K/Akt/mTOR signaling pathway and targeted therapy for glioblastoma. Oncotarget 7:33440–33450. https://doi.org/10.18632/oncotarget.7961
Liang P, Shi H, Zhu W, et al (2017) Silver nanoparticles enhance the sensitivity of temozolomide on human glioma cells. Oncotarget 8:7533–7539. https://doi.org/10.18632/oncotarget.13503
Liu Z, Tan H, Zhang X et al (2018a) Enhancement of radiotherapy efficacy by silver nanoparticles in hypoxic glioma cells. Artif Cells Nanomed Biotechnol 46:922–930. https://doi.org/10.1080/21691401.2018.1518912
Liu F, Feng XX, Zhu SL et al (2018b) Sonic Hedgehog signaling pathway mediates proliferation and migration of fibroblast-like synoviocytes in rheumatoid arthritis via MAPK/ERK signaling pathway. Front Immunol. https://doi.org/10.3389/fimmu.2018.02847
Louca M, Stylianou A, Minia A et al (2019) Ras suppressor-1 (RSU-1) promotes cell invasion in aggressive glioma cells and inhibits it in non-aggressive cells through STAT6 phospho-regulation. Sci Rep 9:7782. https://doi.org/10.1038/s41598-019-44200-8
Maurer LL, Meyer JN (2016) A systematic review of evidence for silver nanoparticle-induced mitochondrial toxicity. Environ Sci Nano. https://doi.org/10.1039/C5EN00187K
Onodera A, Nishiumi F, Kakiguchi K et al (2015) Short-term changes in intracellular ROS localisation after the silver nanoparticles exposure depending on particle size. Toxicol Rep 2:574–579. https://doi.org/10.1016/j.toxrep.2015.03.004
Ou A, Ott M, Fang D, Heimberger A (2021) The role and therapeutic targeting of JAK/STAT signaling in glioblastoma. Cancers (basel) 13:437. https://doi.org/10.3390/cancers13030437
Patlolla AK, Barnes C, Yedjou C et al (2009) Oxidative stress, DNA damage, and antioxidant enzyme activity induced by hexavalent chromium in Sprague-Dawley rats. Environ Toxicol 24:66–73. https://doi.org/10.1002/tox.20395
Perde-Schrepler M, Florea A, Brie I et al (2019) Size-dependent cytotoxicity and genotoxicity of silver nanoparticles in cochlear cells in vitro. J Nanomater. https://doi.org/10.1155/2019/6090259
Piechowiak T, Skóra B, Grzelak-Błaszczyk K, Sójka M (2021) Extraction of antioxidant compounds from blueberry fruit waste and evaluation of their in vitro biological activity in Human Keratinocytes (HaCaT). Food Anal Methods 14:2317–2327. https://doi.org/10.1007/s12161-021-02056-7
Ramaswamy P, Nanjaiah ND, Borkotokey M (2019) Role of MEK-ERK signaling mediated adhesion of glioma cells to extracellular matrix: Possible implication on migration and proliferation. Ann Neurosci 26:52–56. https://doi.org/10.5214/ans.0972.7531.260203
Rinna A, Magdolenova Z, Hudecova A et al (2015) Effect of silver nanoparticles on mitogen-activated protein kinases activation: role of reactive oxygen species and implication in DNA damage. Mutagenesis 30:59–66. https://doi.org/10.1093/mutage/geu057
Salazar-García S, García-Rodrigo JF, Martínez-Castañón GA et al (2020) Silver nanoparticles (AgNPs) and zinc chloride (ZnCl2) exposure order determines the toxicity in C6 rat glioma cells. J Nanoparticle Res 22:253. https://doi.org/10.1007/s11051-020-04984-7
Skóra B, Piechowiak T, Szychowski KA (2022) Epidermal Growth Factor-labeled liposomes as a way to target the toxicity of silver nanoparticles into EGFR-overexpressing cancer cells in vitro. Toxicol Appl Pharmacol 443:116009. https://doi.org/10.1016/j.taap.2022.116009
Skóra B, Matuszewska P, Masicz M et al (2023) Crosstalk between the aryl hydrocarbon receptor (AhR) and the peroxisome proliferator-activated receptor gamma (PPARγ) as a key factor in the metabolism of silver nanoparticles in neuroblastoma (SH-SY5Y) cells in vitro. Toxicol Appl Pharmacol 458:116339. https://doi.org/10.1016/j.taap.2022.116339
Stanton BZ, Peng LF (2010) Small-molecule modulators of the Sonic Hedgehog signaling pathway. Mol BioSyst 6:44–54. https://doi.org/10.1039/B910196A
Szychowski KA, Gmiński J (2019) Specific role of N-methyl-D-aspartate (NMDA) receptor in elastin-derived VGVAPG peptide-dependent calcium homeostasis in mouse cortical astrocytes in vitro. Sci Rep 9:20165. https://doi.org/10.1038/s41598-019-56781-5
Takezaki T, Hide T, Takanaga H et al (2011) Essential role of the Hedgehog signaling pathway in human glioma-initiating cells. Cancer Sci 102:1306–1312. https://doi.org/10.1111/j.1349-7006.2011.01943.x
Tejamaya M, Römer I, Merrifield RC, Lead JR (2012) Stability of citrate, PVP, and PEG coated silver nanoparticles in ecotoxicology media. Environ Sci Technol 46:7011–7017. https://doi.org/10.1021/es2038596
Teperino R, Aberger F, Esterbauer H et al (2014) Canonical and non-canonical Hedgehog signalling and the control of metabolism. Semin Cell Dev Biol 33:81–92. https://doi.org/10.1016/j.semcdb.2014.05.007
Ullah I, Khalil AT, Ali M et al (2020) Green-synthesized silver nanoparticles induced apoptotic cell death in MCF-7 breast cancer cells by generating reactive oxygen species and activating caspase 3 and 9 enzyme activities. Oxid Med Cell Longev 2020:1–14. https://doi.org/10.1155/2020/1215395
Wang Y, Ding Q, Yen C-J et al (2012) The crosstalk of mTOR/S6K1 and Hedgehog pathways. Cancer Cell 21:374–387. https://doi.org/10.1016/j.ccr.2011.12.028
Wang R, Chen C, Yang W et al (2013) Enhancement effect of cytotoxicity response of silver nanoparticles combined with thermotherapy on c6 rat glioma cells. J Nanosci Nanotechnol 13:3851–3854. https://doi.org/10.1166/jnn.2013.7156
Wang H, Ke F, ZHENG J, (2014) Hedgehog-glioma-associated oncogene homolog-1 signaling in colon cancer cells and its role in the celecoxib-mediated anti-cancer effect. Oncol Lett 8:2203–2208. https://doi.org/10.3892/ol.2014.2439
Wang H, Lai Q, Wang D et al (2022) Hedgehog signaling regulates the development and treatment of glioblastoma (Review). Oncol Lett 24:294. https://doi.org/10.3892/ol.2022.13414
Yan Z, Cheng M, Hu G et al (2021) Positive feedback of SuFu negating protein 1 on Hedgehog signaling promotes colorectal tumor growth. Cell Death Dis 12:199. https://doi.org/10.1038/s41419-021-03487-0
Zhu H, Lo H-W (2010) The human glioma-associated oncogene homolog 1 (GLI1) family of transcription factors in gene regulation and diseases. Curr Genom 11:238–245. https://doi.org/10.2174/138920210791233108
Funding
This work was supported by funds from the Ministry of Education and Science in Poland (Project No. SKN/SN/534295/2022) and statutory funds of the University of Information Technology and Management in Rzeszow (DS: 503-07-01-31 and DS: 503-07-01-77).
Author information
Authors and Affiliations
Contributions
BS—conceptualization, methodology, formal analysis, investigation, resources, visualization, writing—original draft, writing—review & editing, funding acquisition, and project administration; MM, PN, JL, PS, JB, and PM—investigation; KSz—investigation, supervising, and writing—review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Skóra, B., Masicz, M., Nowak, P. et al. Suppression of sonic hedgehog pathway-based proliferation in glioblastoma cells by small-size silver nanoparticles in vitro. Arch Toxicol 97, 2385–2398 (2023). https://doi.org/10.1007/s00204-023-03552-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-023-03552-x