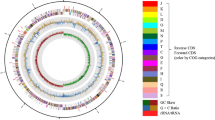
Abstract
Genus Niallia has recently been separated taxonomic group from the Bacillus based on conserved signature indels in the genome. Unlike bioremediation, its role in plant biomass hydrolysis has not garnered considerable attention. The present study investigates the genomic potential of a novel Niallia sp. CRN 25 for applications in lignocellulose hydrolysis, significant enzyme production, and bioremediation. The CRN 25 strain exhibits xylosidase, cellobiosidase, α-arabinosidase, and α-D-galactosidase activity as 0.03 U/ml whereas β-D-glucosidase and glucuronidase as 0.06 U/ml and 0.01 U/ml, respectively. Further genome sequencing reveals nine copies of GH43 gene coding for hemicellulose-specific xylanase enzyme attached to the CBM 6 domain for increased processivity. The presence of β-glucosidase and β-galactosidase indicates the possible application of CRN 25 in facilitating the valorization of plant biomass into value-added products. Apart from this, genes of FMN-dependent NADH-azoreductase, cytochrome P450, and nitrate reductase, playing a crucial role in bioremediation processes, were annotated. Biosynthetic gene clusters (BGCs), responsible for synthesizing specialized metabolites of terpenes and lasso peptides, were also found in the genome. Conclusively genomic sketch of Niallia sp. CRN 25 reveals versatile metabolic potential for diverse environmental applications.







Similar content being viewed by others
Data availability
The data that support the findings of this study are openly available in NCBI at https://www.ncbi.nlm.nih.gov, accession number JABRVO000000000.
References
Arkin AP, Cottingham RW, Henry CS et al (2018) KBase: the United States department of energy systems biology knowledgebase. Nat Biotechnol 36:566–569. https://doi.org/10.1038/nbt.4163
Behrendorff JBYH (2021) Reductive cytochrome P450 reactions and their potential role in bioremediation. Front Microbiol 12:837. https://doi.org/10.3389/FMICB.2021.649273/BIBTEX
Bertelli C, Laird MR, Williams KP et al (2017) IslandViewer 4: expanded prediction of genomic islands for larger-scale datasets. Nucleic Acids Res 45:W30–W35. https://doi.org/10.1093/nar/gkx343
Bhende RS, Jhariya U, Srivastava S et al (2022) Environmental distribution, metabolic fate, and degradation mechanism of chlorpyrifos: recent and future perspectives. Appl Biochem Biotechnol 194:2301–2335. https://doi.org/10.1007/S12010-021-03713-7
Blin K, Shaw S, Augustijn HE et al (2023) antiSMASH 7.0: new and improved predictions for detection, regulation, chemical structures and visualisation. Nucleic Acids Res 51:W46–W50. https://doi.org/10.1093/nar/gkad344
Bohra V, Tikariha H, Dafale NA (2019) genomically defined paenibacillus polymyxa ND24 for efficient cellulase production utilizing sugarcane bagasse as a substrate. Appl Biochem Biotechnol 187:266–281. https://doi.org/10.1007/s12010-018-2820-5
Cheng C, Hua ZC (2020) Lasso peptides: heterologous production and potential medical application. Front Bioeng Biotechnol 8:1131. https://doi.org/10.3389/FBIOE.2020.571165/BIBTEX
Deng Y, Faivre B, Back O et al (2020) Structural and functional characterization of 4-hydroxyphenylacetate 3-hydroxylase from Escherichia coli. ChemBioChem 21:163–170. https://doi.org/10.1002/cbic.201900277
El Bouraie M, El Din WS (2016) Biodegradation of reactive black 5 by Aeromonas hydrophila strain isolated from dye-contaminated textile wastewater. Sustain Environ Res 26:209–216. https://doi.org/10.1016/J.SERJ.2016.04.014
Gupta RS, Patel S, Saini N, Chen S (2020) Robust demarcation of 17 distinct bacillus species clades, proposed as novel bacillaceae genera, by phylogenomics and comparative genomic analyses: description of robertmurraya kyonggiensis sp. nov. and proposal for an emended genus bacillus limiting it only to the members of the subtilis and cereus clades of species. Int J Syst Evol Microbiol 70:5753–5798. https://doi.org/10.1099/IJSEM.0.004475/CITE/REFWORKS
Hanau S, Almugadam SH, Sapienza E et al (2020) Schematic overview of oligosaccharides, with survey on their major physiological effects and a focus on milk ones. Carbohydr Polymer Technol Appl 1:100013. https://doi.org/10.1016/j.carpta.2020.100013
Jha V, Purohit H, Dafale NA (2022) Designing an efficient consortium for improved crop productivity using phosphate stress adapted bacteria with multiple growth-promoting attributes. Geomicrobiol J 39:925–938. https://doi.org/10.1080/01490451.2022.2097340/SUPPL_FILE/UGMB_A_2097340_SM2993.DOC
Lange L, Connor KO, Arason S et al (2021) Developing a sustainable and circular bio-based economy in EU: by partnering across sectors, upscaling and using new knowledge faster, and for the benefit of climate, environment & biodiversity, and people & business. Front Bioeng Biotechnol. https://doi.org/10.3389/fbioe.2020.619066
Li E, Zhu Q, Pang D et al (2022) Analysis of Lactobacillus rhamnosus GG in mulberry galacto-oligosaccharide medium by comparative transcriptomics and metabolomics. Front Nutr 9:853271. https://doi.org/10.3389/FNUT.2022.853271
Liang Z, Zhi H, Fang Z, Zhang P (2021) Genetic engineering of yeast, filamentous fungi and bacteria for terpene production and applications in food industry. Food Res Int 147:110487. https://doi.org/10.1016/J.FOODRES.2021.110487
Liu Y, Pei R, Huang Z et al (2021) Green immobilization of CdS-Pt nanoparticles on recombinant Escherichia coli boosted by overexpressing cysteine desulfurase for photocatalysis application. Bioresour Technol Rep 16:100823. https://doi.org/10.1016/J.BITEB.2021.100823
Liu B, Zheng D, Zhou S et al (2022) VFDB 2022: a general classification scheme for bacterial virulence factors. Nucleic Acids Res 50:D912–D917. https://doi.org/10.1093/nar/gkab1107
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun 10:2182. https://doi.org/10.1038/s41467-019-10210-3
Mukkala S, Bramhachari PV, Reddy YHK (2022) The cellulosome: a fiber-degrading strategist of the rumen microbiome. Underst Microb Interact Agric Environ. https://doi.org/10.1007/978-981-19-3696-8_11
Omenesa Idris M, Asshifa MNN, Nasir Mohamad Ibrahim M, Ali Yaqoob A (2023) Sustainable microbial fuel cell functionalized with a bio-waste: a feasible route to formaldehyde bioremediation along with bioelectricity generation. Chem Eng J 455:140781. https://doi.org/10.1016/j.cej.2022.140781
Ontañon OM, Bedő S, Ghio S et al (2021) Optimisation of xylanases production by two cellulomonas strains and their use for biomass deconstruction. Appl Microbiol Biotechnol 105:4577–4588. https://doi.org/10.1007/S00253-021-11305-Y/FIGURES/4
Ottoni JR, Bernal SPF, Marteres TJ et al (2022) Cultured and uncultured microbial community associated with biogas production in anaerobic digestion processes. Arch Microbiol 204:1–13. https://doi.org/10.1007/S00203-022-02819-8/FIGURES/4
Patel D, Patil KS, Madamwar D, Desai C (2022) Electrogenic degradation of reactive red 152 dye by Niallia circulans DC10 and its genome sequence analysis reveals genes mediating dye degradation and anodic electron transfer. J Water Process Eng 47:102690. https://doi.org/10.1016/J.JWPE.2022.102690
Sathitkowitchai W, Wongsurawat T, Jenjaroenpun P et al (2022) Complete genome sequences of mannanase-producing Bacillus and niallia strains isolated from the intestine of the black tiger shrimp (Penaeus monodon). Microbiol Resour Announc. https://doi.org/10.1128/MRA.00112-22
Shi Q, Abdel-Hamid AM, Sun Z et al (2023) Carbohydrate-binding modules facilitate the enzymatic hydrolysis of lignocellulosic biomass: releasing reducing sugars and dissociative lignin available for producing biofuels and chemicals. Biotechnol Adv 65:108126. https://doi.org/10.1016/J.BIOTECHADV.2023.108126
Sista Kameshwar AK, Qin W (2018) Understanding the structural and functional properties of carbohydrate esterases with a special focus on hemicellulose deacetylating acetyl xylan esterases. Mycology 9:273–295. https://doi.org/10.1080/21501203.2018.1492979
Srivastava S, Dafale NA, Purohit HJ (2020) Functional genomics assessment of lytic polysaccharide mono-oxygenase with glycoside hydrolases in Paenibacillus dendritiformis CRN18. Int J Biol Macromol 164:3729–3738. https://doi.org/10.1016/j.ijbiomac.2020.08.147
Srivastava S, Dafale NA, Jakhesara SJ et al (2021) Unraveling the camel rumen microbiome through metaculturomics approach for agriculture waste hydrolytic potential. Arch Microbiol 203:107–123. https://doi.org/10.1007/s00203-020-02010-x
Srivastava S, Bombaywala S, Jakhesara SJ et al (2023) Potential of camel rumen derived Bacillus subtilis and Bacillus velezensis strains for application in plant biomass hydrolysis. Mol Genet Genomics 298:361–374. https://doi.org/10.1007/s00438-022-01987-y
Su X, Meng F, Liu Y et al (2022) Molecular cloning and functional characterization of a β-Glucosidase gene to produce platycodin D in Platycodon grandiflorus. Front Plant Sci 13:955628. https://doi.org/10.3389/FPLS.2022.955628/FULL
Subbotina NM, Chernykh AM, Taranov AI et al (2021) Gentisate 1,2-dioxygenase from the gram-positive bacteria Rhodococcus opacus 1CP: identical active sites vs. different substrate selectivities. Biochimie 180:90–103. https://doi.org/10.1016/J.BIOCHI.2020.10.016
Thorat V, Kirdat K, Tiwarekar B et al (2022) Paenibacillus albicereus sp. nov. and Niallia alba sp. nov., isolated from digestive syrup. Arch Microbiol 204:1–11. https://doi.org/10.1007/S00203-021-02749-X/FIGURES/3
Travkin VM, Solyanikova IP (2021) Salicylate or phthalate: the main intermediates in the bacterial degradation of naphthalene. Processes 9:1862. https://doi.org/10.3390/PR9111862
van Staden ADP, van Zyl WF, Trindade M et al (2021) Therapeutic application of Lantibiotics and other Lanthipeptides: old and new findings. Appl Environ Microbiol. https://doi.org/10.1128/AEM.00186-21/SUPPL_FILE/AEM00186-21_SUPP_1_SEQ5.PDF
Xia L, Miao Y, Cao A et al (2022) Biosynthetic gene cluster profiling predicts the positive association between antagonism and phylogeny in Bacillus. Nat Commun 13:1–11. https://doi.org/10.1038/s41467-022-28668-z
Yin Y, Mao X, Yang J et al (2012) dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 40:W445–W451. https://doi.org/10.1093/nar/gks479
Zhao W, Huang Y, Wang M et al (2018) Post-endogenous denitrification and phosphorus removal in an alternating anaerobic/oxic/anoxic (AOA) system treating low carbon/nitrogen (C/N) domestic wastewater. Chem Eng J 339:450–458. https://doi.org/10.1016/J.CEJ.2018.01.096
Acknowledgements
The authors highly acknowledge The Director, CSIR-NEERI, Nagpur, for providing facilities to carry out this work. The manuscript has been checked for plagiarism by Knowledge Resource Centre, CSIR-NEERI, Nagpur, India, and assigned KRC No.: CSIR-NEERI/KRC/2023/APRIL/EBGD/1.
Funding
The authors have not received any financial support for the research, authorship, and publication of this article.
Author information
Authors and Affiliations
Contributions
SS: Writing original manuscript, experiments, in-silico analysis, visualization; ND: Conceptualization, resources, supervision, review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no financial or commercial conflict of interest.
Ethical approval
This study is a genome analysis based on the hydrolytic and bioremediation potential of Niallia sp. and does not directly involve animals or humans as research subjects.
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Srivastava, S., Dafale, N.A. Genomic dissection of Niallia sp. for potential application in lignocellulose hydrolysis and bioremediation. Arch Microbiol 206, 2 (2024). https://doi.org/10.1007/s00203-023-03728-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03728-0