Abstract
Introduction and hypothesis
The study is aimed at bioinformatically deciphering immune cell infiltration, signature genes, and their correlations in POP.
Methods
Three microarray datasets were included. Matrixes representing the uterosacral ligament were merged as a test matrix and the others representing vaginal tissues were merged as a validation matrix. The single-sample Gene Set Enrichment Analysis (ssGSEA) algorithm was performed to evaluate immune cell infiltration. Correlations among differential immune cells were revealed by Spearman’s rank correlation. Differentially expressed genes (DEGs) were screened by both “Batch correction” and “RobustRankAggreg” methods. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes were conducted for functional analysis. Hub genes were identified through cytoHubba of Cytoscape, and further validated by a validation matrix and clinical samples as signature genes. Correlations of differential immune cells with signature genes were analyzed by Spearman’s rank correlation.
Results
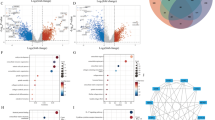
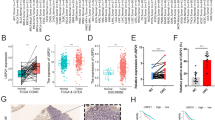
Five differential immune cells (macrophages, monocytes, regulatory T cell [Treg], type 1 T cell [Th1], and natural killer T cells [NKT]) were identified and eight pairs of immune cells had significant correlations. Screened 230 DEGs were extracellular matrix (ECM) and immune related. Eleven hub genes were initially identified and five of them (LOX, IL-6, SDC1, ICAM1, and CD38) were validated as signature genes. Significant correlations of differential immune cells with signature genes were shown in twelve pairs, especially Th1–IL6, NKT–IL6, Th1–ICAM1, macrophage–IL6, and macrophage–LOX pairs.
Conclusions
Pelvic organ prolapse could be considered immune related. Significantly infiltrated immune cells may contribute to the development of POP through close involvement with ECM- and immune-related signature genes.





Similar content being viewed by others
Data availability
The datasets (GSE28660, GSE12852, GSE53868) analyzed in the study are publicly available from the GEO DataSets (https://www.ncbi.nlm.nih.gov/).
References
Gong R, Xia Z. Collagen changes in pelvic support tissues in women with pelvic organ prolapse. Eur J Obstet Gynecol Reprod Biol. 2019;234:185–9.
Maldonado PA, Wai CY. Pelvic organ prolapse: new concepts in pelvic floor anatomy. Obstet Gynecol Clin N Am. 2016;43(1):15–26.
Kieserman-Shmokler C, Swenson CW, Chen L, Desmond LM, Ashton-Miller JA, DeLancey JO. From molecular to macro: the key role of the apical ligaments in uterovaginal support. Am J Obstet Gynecol. 2020;222(5):427–36.
Deprest JA, Cartwright R, Dietz HP, Brito LGO, Koch M, Allen-Brady K, et al. International Urogynecological Consultation (IUC): pathophysiology of pelvic organ prolapse (POP). Int Urogynecol J. 2022;33(7):1699–710.
Weintraub AY, Glinter H, Marcus-Braun N. Narrative review of the epidemiology, diagnosis and pathophysiology of pelvic organ prolapse. Int Braz J Urol. 2020;46(1):5–14.
Bildircin D, Kokcu A, Celik H, Sagir D, Kefeli M. Comparison of connective tissue components in the uterine ligaments between women with and without pelvic organ prolapse. Minerva Ginecol. 2014;66(2):201–8.
Gong R, Ji Y, Zhao Y, Xia Z. Changes in β-catenin expression in the anterior vaginal wall tissues of women with pelvic organ prolapse: a potential pathophysiological mechanism. Female Pelvic Med Reconstr Surg. 2020;26(11):e54–61.
Li Y, Nie N, Gong L, Bao F, An C, Cai H, et al. Structural, functional and molecular pathogenesis of pelvic organ prolapse in patient and Loxl1 deficient mice. Aging (Albany NY). 2021;13(24):25886–902.
Zhu YP, Xie T, Guo T, Sun ZJ, Zhu L, Lang JH. Evaluation of extracellular matrix protein expression and apoptosis in the uterosacral ligaments of patients with or without pelvic organ prolapse. Int Urogynecol J. 2021;32(8):2273–81.
Orlicky DJ, Guess MK, Bales ES, Rascoff LG, Arruda JS, Hutchinson-Colas JA, et al. Using the novel pelvic organ prolapse histologic quantification system to identify phenotypes in uterosacral ligaments in women with pelvic organ prolapse. Am J Obstet Gynecol. 2021;224(1):67.e1–e18.
Li Y, Zhang QY, Sun BF, Ma Y, Zhang Y, Wang M, et al. Single-cell transcriptome profiling of the vaginal wall in women with severe anterior vaginal prolapse. Nat Commun. 2021;12(1):87.
Zhou Q, Hong L, Wang J. Identification of key genes and pathways in pelvic organ prolapse based on gene expression profiling by bioinformatics analysis. Arch Gynecol Obstet. 2018;297(5):1323–32.
Zhao Y, Xia Z, Lin T, Yin Y. Significance of hub genes and immune cell infiltration identified by bioinformatics analysis in pelvic organ prolapse. PeerJ. 2020;8:e9773.
Liu P, Liu D, Chen F, Luo L, Jin Y, Peng J, et al. Effect of Nrf2 on phenotype changes of macrophages in the anterior vaginal wall of women with pelvic organ prolapse. Female Pelvic Med Reconstr Surg. 2022. https://doi.org/10.1097/SPV.0000000000001212.
Artsen AM, Rytel M, Liang R, King GE, Meyn L, Abramowitch SD, et al. Mesh induced fibrosis: the protective role of T regulatory cells. Acta Biomater. 2019;96:203–10.
Roumier T, Capron M, Dombrowicz D, Faveeuw C. Pathogen induced regulatory cell populations preventing allergy through the Th1/Th2 paradigm point of view. Immunol Res. 2008;40(1):1–17.
Wynn TA. Fibrotic disease and the TH1/TH2 paradigm. Nat Rev Immunol. 2004;4(8):583–94.
Duan YG, Gong J, Yeung WSB, Haidl G, Allam JP. Natural killer and NKT cells in the male reproductive tract. J Reprod Immunol. 2020;142:103178.
Kobak W, Lu J, Hardart A, Zhang C, Stanczyk FZ, Felix JC. Expression of lysyl oxidase and transforming growth factor beta2 in women with severe pelvic organ prolapse. J Reprod Med. 2005;50(11):827–31.
Zhang SQ, Zhang LL, Yu H. Expression of elastin, lysyl oxidase and elafin in the cardinal ligament of women with pelvic organ prolapse. Zhonghua Fu Chan Ke Za Zhi. 2008;43(9):675–9.
Liu X, Zhao Y, Pawlyk B, Damaser M, Li T. Failure of elastic fiber homeostasis leads to pelvic floor disorders. Am J Pathol. 2006;168(2):519–28.
Qiu J, Qin M, Fan B, Chen X. Klotho protein reduced the expression of matrix metalloproteinase-1 (MMP-1) and matrix metalloproteinase-3 (MMP-3) in fibroblasts from patients with pelvic organ prolapse (POP) by down-regulating the phosphorylation of ERK1/2. Med Sci Monit. 2019;25:3815–24.
Takeuchi T, Yoshida H, Tanaka S. Role of interleukin-6 in bone destruction and bone repair in rheumatoid arthritis. Autoimmun Rev. 2021;20(9):102884.
Angsana J, Chen J, Smith S, Xiao J, Wen J, Liu L, et al. Syndecan-1 modulates the motility and resolution responses of macrophages. Arterioscler Thromb Vasc Biol. 2015;35(2):332–40.
Roebuck KA, Finnegan A. Regulation of intercellular adhesion molecule-1 (CD54) gene expression. J Leukoc Biol. 1999;66(6):876–88.
Piedra-Quintero ZL, Wilson Z, Nava P, Guerau-de-Arellano M. CD38: an immunomodulatory molecule in inflammation and autoimmunity. Front Immunol. 2020;11:597959.
Acknowledgement
We are grateful for the GEO database and all suppliers who uploaded datasets of GSE28660, GSE12852, GSE151188.
Funding
This work was supported by the National Natural Science Foundation of China (No. 82071630 and 81771560 granted to X. Tong; No. 81702745 granted to Y. Yang; No. 81873827 granted to Y. Guo); Tongji Hospital National Natural Science Foundation Training Project (No. GJPY2119 granted to X. Tong). The funding organizations did not contribute to any of the following: design of the study, data collection, data analysis, interpretation of data, and writing the manuscript or decision to publish.
Author information
Authors and Affiliations
Contributions
C. Wu: manuscript writing, bioinformatics analysis; Z. Zhou: experiment performance and data analysis; Y. Yang and H. Li: sample collection; G. Yi: manuscript revision; X. Tong: project development and manuscript revision.
Corresponding authors
Ethics declarations
Ethics approval
The study protocol was approved by the Ethics Committee of Tongji Hospital, Tongji University School of Medicine (Number: K-2021-1115). All participants received the written informed consent.
Conflicts of interest
None.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, C., Zhou, Z., Yang, Y. et al. Bioinformatically deciphering immune cell infiltration and signature genes in pelvic organ prolapse. Int Urogynecol J 34, 1091–1101 (2023). https://doi.org/10.1007/s00192-022-05378-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00192-022-05378-0