Abstract
Purpose
The heterogeneity in sepsis is held responsible, in part, for the lack of precision treatment. Many attempts to identify subtypes of sepsis patients identify those with shared underlying biology or outcomes. To date, though, there has been limited effort to determine overlap across these previously identified subtypes. We aimed to determine the concordance of critically ill patients with sepsis classified by four previously described subtype strategies.
Methods
This secondary analysis of a multicenter prospective observational study included 522 critically ill patients with sepsis assigned to four previously established subtype strategies, primarily based on: (i) clinical data in the electronic health record (α, β, γ, and δ), (ii) biomarker data (hyper- and hypoinflammatory), and (iii–iv) transcriptomic data (Mars1–Mars4 and SRS1–SRS2). Concordance was studied between different subtype labels, clinical characteristics, biological host response aberrations, as well as combinations of subtypes by sepsis ensembles.
Results
All four subtype labels could be adjudicated in this cohort, with the distribution of the clinical subtype varying most from the original cohort. The most common subtypes in each of the four strategies were γ (61%), which is higher compared to the original classification, hypoinflammatory (60%), Mars2 (35%), and SRS2 (54%). There was no clear relationship between any of the subtyping approaches (Cramer’s V = 0.086–0.456). Mars2 and SRS1 were most alike in terms of host response biomarkers (p = 0.079–0.424), while other subtype strategies showed no clear relationship. Patients enriched for multiple subtypes revealed that characteristics and outcomes differ dependent on the combination of subtypes made.
Conclusion
Among critically ill patients with sepsis, subtype strategies using clinical, biomarker, and transcriptomic data do not identify comparable patient populations and are likely to reflect disparate clinical characteristics and underlying biology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
This study explores the comparison of sepsis subtypes using clinical and molecular techniques. It furthers the understanding of sepsis heterogeneity revealing that subtypes are likely to represent complimentary pathways to precision medicine approaches in sepsis. |
Introduction
Sepsis is a life-threatening organ dysfunction caused by a dysregulated host response to an infection [1]. The global burden of sepsis is estimated to be 49 million cases annually [2] of which half require intensive care unit (ICU) treatment [3]. Sepsis accounts for more than 11 million deaths [4], and significant long-term morbidity amongst survivors [5]. Despite progress in the understanding of sepsis pathophysiology, no specific treatment has proven successful [6, 7]. Amid calls for precision therapy, a greater understanding of the heterogeneity of sepsis is needed and a focus of clinicians, trialists, and funders [6, 8,9,10].
Recent approaches to measure sepsis heterogeneity used unsupervised computational methods on clinical, biomarker or gene expression data from observational studies or clinical trial datasets. Examples include subtypes from the Sepsis Endotyping in Emergency Care (SENECA) study [11], acute respiratory distress syndrome (ARDS) subphenotypes from the ARDSnet randomized trials [12, 13], and Molecular Diagnosis and Risk Stratification of Sepsis (MARS) and sepsis response signatures (SRS) endotypes based on transcriptomic data of whole blood leucocytes [14, 15]. At present, more than 100 sepsis subtypes are proposed, without awareness of overlap (or clinical implications), partly caused by the lack of data in a single cohort to compute each of the subtypes [16]. It is unknown whether each new subtype strategy is of added value for the patient or merely ‘reinventing the wheel’, by being comparable to already existing subtypes. This is essential information to inform predictive enrichment in future trials [10].
To address this knowledge gap, we sought to determine the concordance (i.e., agreement) between different subtype labels, outcomes, and biologic pathways of critically ill sepsis patients classified by previously proposed sepsis subtyping methods. We hypothesized that subtype strategies derived from different data sources signify unrelated subtypes, due to the complex nature of sepsis and fundamental differences between clinical, biomarker, and transcriptomic data. However, we also hypothesized that partial overlap will be present in the subtypes with most unfavorable outcome, mainly driven by disease severity. Moreover, we hypothesized that concordance was high when subtypes were based on the same data source.
Methods
We used a prior systematic review [16] to identify candidate subtype labels for critically ill sepsis patients (supplemental Table E1). We chose four subtype strategies shown to be reproducible, associated with differential treatment effects in prior work [11, 17,18,19,20], and reflective of multiple domains. These are a mainly clinical based domain [11], a combination of clinical and biological domains [21] and a transcriptomic domain [14, 15, 20]. To assign subtypes, we analyzed a previously selected cohort from the MARS study making use of clinical data, host response biomarkers, and transcriptomic data from whole blood leukocytes, all collected within 24 h after admission [14]. For each patient, a subtype label was applied for each of the four strategies. See online supplementary methods for information on host response biomarker measurements and blood leukocyte gene expression analysis.
Study design, patients, and definitions
The MARS cohort was a prospective observational study in the mixed ICUs of two tertiary teaching hospitals (Academic Medical Center in Amsterdam and University Medical Center in Utrecht) [14]. Consecutive patients above 18 years of age with an expected length of stay longer than 24 h admitted between January 2011 and January 2014 were included via an opt-out method approved by the medical ethical committees of the participating hospitals [14]. We analyzed a previously defined cohort of patients fulfilling Sepsis-2 (and Sepsis-3) criteria at ICU admission in whom clinical, biomarker and transcriptomic data was available making it possible to determine at least 3 out of 4 subtypes.
Sepsis subtype adjudication
First, clinical characteristics were used to identify the SENECA subtypes α, β, γ, and δ using the SENECA approach [11]. All clinical variables used for SENECA subtype derivation in SENECA were identified in the MARS cohort (supplemental Table E2). Then, subtypes were adjudicated by calculating the Euclidean distance from each patient to the centroid of each subtype from the SENECA derivation cohort [11].
To apply sepsis subtype labels based upon ARDS subphenotypes (termed ARDS subtypes “hyperinflammatory” and “hypoinflammatory”), we applied the previously published parsimonious model using plasma levels of bicarbonate, interleukin-8 (IL-8), and protein C [21].
To identify MARS and SRS subtypes, we used labels developed from whole blood leukocyte RNA expression. For MARS subtype adjudication, the original classification, which used consensus clustering method in the discovery cohort and a sepsis endotype classifier in the validation cohort, was applied [14]. SRS 1 or 2 was classified by applying the SepstratifieR method [22] using a previously derived 7-gene signature [23], with default parameters. For more information on subtype adjudication, see online supplementary methods.
Statistical analysis
We analyzed multiple domains of the sepsis subtypes; (i) reproducibility and clinical characteristics, (ii) concordance of subtype membership, (iii) biologic pathways of subtypes implicated through plasma protein biomarkers and whole blood leukocyte gene expression profiles, and (iv) development of subtype ensembles by adjudicating multiple subtypes to a single patient and, thus, creating subsets of patients.
First, the reproducibility and clinical characteristics of subtypes compared to the original studies was assessed. The overall distribution of the specific subtype, clinical characteristics, and outcomes (e.g., in-hospital mortality) were measured. Second, to study concordance between different subtype labels, proportions, alluvial plots and Cramer’s V, corrected for bias, were used. Cramer’s V is a measure of the strength of association between different subtype labels and was characterized as follows: 0–0.1 no association, 0.1–0.3 weak association, 0.3–0.5 moderate association, and > 0.5 strong association [24]. Third, biological domains of subtypes were described by plasma protein biomarker concentration on admission. Furthermore, whole blood leukocyte gene expression data were used to determine differentially expressed genes and to identify canonical signaling pathways in each subtype using Reactome database (R software package, reactome.db version 1.82) [25, 26]. Fourth, explorative analysis of subsets of patients was performed by identifying groups of patients with most subtype overlap, determined by (1) association strength, (2) overlapping biological pathways, and (3) ≥ 10 patients. Each enriched subset, called sepsis ensemble, was then compared as a “new subtype of multiple subtypes”.
In all analyses, a Mann–Whitney U or a Kruskal–Wallis test was used to analyze continuous nonparametric data, presented as median and interquartile ranges (IQR). Continuous parametric data, presented as means ± standard deviation (SD), were analyzed using a Student’s t test or analysis of variance when appropriate. Post hoc testing was performed using Dunn’s test of multiple comparisons using rank sums for nonparametric continuous data, a Tukey post hoc testing for parametric continuous data and a Bonferroni correction for categorical variables. Categorical data, presented as numbers (percentages), were analyzed using a chi-square test. All data were analyzed using R studio built under R version 4.0.3 (R Core Team 2013, Vienna, Austria). A P value < 0.05 was considered to be of statistical significance.
Results
Patients
Among 8332 patients in the MARS study, 2499 patients were admitted with sepsis and 522 patients had the possibility to adjudicate at least 3 out of 4 subtypes. Mean age was 61.2 years (SD 14.6), male sex was most common (n = 297, 56.9%) and the median sequential organ failure assessment (SOFA) score was 7 (IQR [5–9]) on ICU admission. Patients were commonly mechanically ventilated in the first 24 h of ICU admission (n = 421, 80.7%) and in-hospital mortality rate was 30.1% (Table 1). Nearly half (n = 226, 43.3%) were admitted with a pulmonary infection, while abdominal infections were less common (n = 130, 25%).
Reproducibility and clinical characteristics of sepsis subtypes
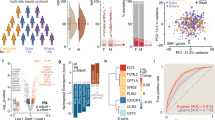
After SENECA subtype adjudication using the SENECA approach in this critically ill cohort, 316 patients were classified as γ (60.5%), 188 as δ (36%), 12 as β (2.3%), and 6 as α (1.1%). The distribution of subtypes, especially α and β, was different from the original SENECA cohort (supplemental Fig. 1). The patterns of clinical data and host response were reproduced in the γ- and δ-type (supplemental Table 3). For example, γ-type patients had higher markers of inflammation, with the highest C-reactive protein (CRP) levels (169 mg/L [87–267], p = 0.007) and core temperature (38.2 (1.1) °C, p = 0.001). δ-type patients had the most abnormal cardiovascular, liver function and hemodynamic markers, and highest in-hospital mortality (n = 82 (43.6%), p < 0.001) compared to other subtypes (supplemental Table 3).
For ARDS subtypes, more patients were adjudicated to the hypoinflammatory subtype (59.9%) than to the hyperinflammatory subtype (40.1%) (supplemental Table 4). Patients classified as hyperinflammatory showed higher SOFA scores (9 [7–11] vs 6 [4–8], p < 0.001) and higher levels of CRP on admission (180 mg/mL [99–273] versus 138 mg/mL [62–234], p = 0.009). Markers reflective of cardiovascular and hemodynamic aberrations were more abnormal in the hyperinflammatory group with lower bicarbonate, increased heartrate, increased serum lactate and lower systolic blood pressure (supplemental Table 4, p = < 0.001–0.002). The hyperinflammatory subtype showed increased mortality compared to hypoinflammatory (37.5% vs 19.7%, p < 0.001).
Two transcriptomic labels were used to classify sepsis patients using whole blood RNA expression profiles. First, patients were adjudicated to the MARS subtypes, identical to the original discovery and validation cohort (supplemental Table 5, [14]). SRS subtypes yielded similar distributions as the original derivation cohort (SRS1 45.6% vs 40.7%). Similar as previously reported and compared to SRS2, SOFA score was highest (8 [5–10] vs 7 [4–9], p = 0.002) and bicarbonate was lowest (18.7 mEq/L (5.6) vs 21.5 mEq/L (6.1), p < 0.001) in SRS1. However, there was no significant mortality difference between the SRS subtypes in the MARS cohort (supplemental Table 6).
Concordance between subtypes
There was no relationship between any of the subtyping approaches (Fig. 1, supplemental Table 7–8). For example, when comparing SENECA versus ARDS subtypes, 71.3% of γ-type patients were classified as hypoinflammatory. Meanwhile, of the δ-type patients, 64% were classified as hyperinflammatory resulting in a moderate association between the SENECA and ARDS strategies (Cramer’s V = 0.366). The clinical characteristics of γ-type were similar to those classified as hypoinflammatory, as well as the 30-day mortality (supplemental Figs. 2–3, supplemental Table 3–4).
When comparing SENECA vs. MARS and SRS labels, δ-type patients were mostly distributed amongst Mars1 and Mars2 patients and the majority of γ-type patients were classified as Mars2. This suggests a weak association between the two approaches (Cramer’s V = 0.091). The same weak association was true for the SRS subtype labels (Cramer’s V = 0.086). Subsequently, clinical characteristics classified by SENECA subtypes showed no clear relation to clinical characteristics classified by MARS or SRS subtypes (Table 1, supplemental Fig. 2, supplemental Table 3, 5–6). Only δ-type patients and Mars1 appeared to have similar survival rates (supplemental Fig. 3).
When comparing ARDS subtypes vs. MARS and SRS subtypes, a moderate association between the strategies was found (Fig. 1, Cramer’s V = 0.351–0.382). The hypoinflammatory patients were most comparable with Mars3 (36.9%) and/or SRS2 (69.5%), which agrees with the clinical characteristics (Fig. 1 and supplemental Fig. 2).
When comparing MARS and SRS labels, almost all Mars3 patients were classified as SRS2 (90.7%). However, the remaining transcriptomic labels showed little concordance resulting in a moderate association (Fig. 1, Cramer’s V = 0.456).
Relationship between subtyping strategies and plasma protein biomarkers and immune cell gene expression mechanistic pathway analysis
Among 466 patients with measured host response biomarkers (89% of the cohort, Fig. 2, supplemental Table 9–12), we observed that inflammation markers were most elevated in δ, hyperinflammatory, Mars2 and SRS1. However, the hyperinflammatory ARDS subtype was significantly different from all other subtypes (p < 0.01). In line with the distribution between subtypes, the hypoinflammatory subtype was most similar to γ-type, Mars3 and SRS2 (p = 0.001–0.439). Coagulation markers, including D-dimer and antithrombin, and endothelial cell activation markers, including angiopoietin and fractalkine, were most abnormal in δ-type, hyperinflammatory, Mars2, and SRS1, although almost all subtypes showed similar concentrations. Overall, Mars2 and SRS1 were most comparable in biomarker levels across all subtypes in all domains (p = 0.079–0.424).
Host response biomarkers classified by sepsis subtype. Boxplots displaying biomarker concentrations (pg/ml), after log10 transformation. Horizontal line is the median concentration of healthy volunteers (n = 25). Biomarkers are grouped by domain (Inflammation, Coagulation, and Endothelial dysfunction). IL Interleukin, MMP matrix metalloproteinase, ICAM intercellular adhesion molecule, ANG angiopoietin
There were substantial alterations in blood leukocyte gene expression among all sepsis subtypes compared to healthy controls. Yet, only few genes were specific within each subtype label (supplemental Figs. 4, 5). When comparing subtypes, δ-type and Mars1 shared the most unique genes (supplemental Figs. 6, 7). Pathway analysis confirmed previously implicated mechanisms [14, 15]. For example, downregulation in major pathways in the adaptive and innate immune system were observed in Mars1 and SRS2 (Fig. 3), but also in the δ-type and hyperinflammatory subtype. Pathways involved in hemostasis were mostly not significant, whereas metabolism pathways were overexpressed and similar in the hyperinflammatory and Mars2 subtypes.
Blood transcriptional responses between subtypes for targeted pathways. Heatmaps showing the Benjamini–Hochberg adjusted P value correcting for all existing Reactome database pathways, and direction using the Normalized Enrichment Score (red = + NES, blue = − NES). Canonical signaling pathways were grouped into major pathways, indicated in bold
Sepsis ensembles
In an exploratory analysis among 64 possible subtype combinations, 17 ensembles contained at least 10 patients (supplemental Fig. 8), with membership ranging from 10 to 65 patients (median 20, IQR 15). These ensembles varied in characteristics and outcome, and were dependent on the combination of subtyping approaches (Fig. 4, supplemental Fig. 9). For example, hospital mortality ranged from 8% to 46% as the Mars subtype changed from Mars4 to Mars1, according to whether that patient also belonged to γ-type, hypoinflammatory and SRS1. In another example, although not significantly, endothelial markers and mortality decreased as the SRS subtype changed in combination with δ-type, hyperinflammatory and Mars2, but not when combined with Mars1 (supplemental Fig. 9).
Circular visualization of sepsis ensembles. Circle plot, where each section represents a patient subset of at least 10 patients. From in to out, group size, hospital mortality, boxplot of clinical variables, heatmap of plasma protein biomarkers and a heatmap of pathway analysis of the transcriptome are displayed. In the boxplot, the variables are scaled and the horizontal dotted line is 0. Each row in the heatmap represents a biomarker, ordered and colored according to Fig. 2. The pathways are the 5 major pathways in Fig. 3
Discussion
In this secondary analysis of the MARS prospective cohort study, we describe a comparison of sepsis subtypes, based on clinical, biomarker or transcriptomic data in critically ill patients with sepsis. Subtypes could be assigned successfully, but the distribution of the SENECA subtypes and mortality in SRS subtypes was notably different from the original cohorts. Concordance between SENECA, ARDS, MARS and SRS subtypes was moderate to low, concluding that a different set of patients was identified in each subtype strategy. Host response biomarkers and transcriptomic data showed the most similarity between Mars2 and SRS1. When patients were assigned multiple subtypes, this created subgroups of patient with a broad variability in characteristics and in-hospital mortality.
Compared to prior reports, SENECA subtypes of sepsis were similar in their clinical characteristics [11]. However, the distribution of the SENECA subtypes in this critically ill population was different from those reported in the original Emergency Department (ED) population. For example, the rates of α and β were < 10%, far lower than 20–30% in broader study cohorts. Even though the distribution of SENECA subtypes was dependent on case mix differences, the clinical characteristics of the individual SENECA subtype members were consistent with prior work [11]. Notably, we found no clear relationship between the distribution of SENECA subtypes nor clinical characteristics with any of the transcriptomic subtypes. For example, we did not observe concordance between δ patients and Mars1, even though both subtypes tend toward higher organ dysfunction and greater mortality, which was not in line with our hypothesis. However, host response biomarker aberrations were most similar between γ-type and Mars4, and to a lesser extent between δ-type and Mars2. This could be driven in part by disease severity, as δ-type and Mars2 had the highest SOFA score.
The most overlap in the distribution of sepsis subtypes was between subtype strategies derived from the same data source (MARS and SRS subtypes). In addition, there was also a moderate association between ARDS subtypes and all other subtypes. In the absence of concordance between clinical and molecular classifiers, it may be that biomarkers form a key link between clinical and transcriptomic approaches to subtyping sepsis. Furthermore, biologic subtypes also show a difference in gene expression profiles and prior work has shown that SENECA subtypes are modifiable when new biomarker data is added [27, 28]. Although there is no ground truth to subtype labels or gold standard to compare, the inclusion of biologic features such as plasma protein biomarkers may be feasible, and unveil subtypes suitable for evaluation in clinical trials [29].
These findings suggest that multiple subtype strategies reflect different, perhaps complimentary pieces to the subtyping story. Similar work in traumatic injury and oncology reveals that combinations of clinical, biomarker, and transcriptomic subtypes will reveal important ensembles, however small, that are prognostic or predictive of treatment response and have added value compared to a single subtype [30, 31]. As a proposed solution to investigate the complimentary nature of the subtypes, we explored sepsis subtype ensembles as combinations. In our exploratory analysis and the examples given, we show that combining subtypes seem to give a more precise estimate of the outcome compared to a single subtype. Nevertheless, there are many challenges with this approach. First, this stratification approach requires large sample sizes. Dividing the homogeneous subgroups into more homogeneous subgroups will quickly lead to loss of power and low-grade evidence if the sample size is not adjusted. Second, feasibility of acquiring the subtypes is important. Considering a bed-side approach, it could be more realistic to identify SENECA subtypes over ARDS or MARS and SRS subtypes, regarding the time and effort that it costs. However, routinely available clinical variables only indirectly reflect the underlying injury process and biomarker data is essential in identifying subtypes with heterogeneity of treatment effect [32]. Third, as data layers increase, so does the need for dimension reduction, to encounter overlap, duplicity and overfitting. In this study, the γ-type is, amongst others, characterized by high clinical inflammatory markers such as CRP and an upregulated immune system in the transcriptomic data; however, host response inflammatory biomarkers such as IL-6 and IL-8 are lower compared to the δ-type. This indicates a mismatch in the data relationship, creating a challenge in the integration of omics and non-omics data. In the long run, the ultimate goal should be the same for every method: to stratify patients into more homogenous subsets with an identifiable and verifiable treatable trait [33]. Taken together, the ensembles explored here demonstrate feasibility of existing subtype labels to jointly identify unique patients. It is unknown though how these subsets will identify treatable traits.
This study has several limitations. First, SENECA subtypes were applied using a reduced feature set compared to the original work [11]. This limitation is previously shown to have modest, if any, impact on classification and subtype characteristics. Second, the prospective enrollment of the MARS population occurred prior to updated clinical practice guidelines and this analysis is performed in a single cohort. This could limit generalizability. Third, not all published sepsis subtypes were feasible to include in this analysis. We selected four highly cited approaches that reflected different features, analytic methods, and were potentially linked to treatment. Fourth, this result may not apply to low-income and middle-income countries or patients cared for in non-ICU settings.
Conclusion
Among critically ill patients with sepsis, subtype strategies using clinical, biomarker, and transcriptomic data do not identify comparable patient populations and are likely to reflect disparate clinical characteristics and underlying biology.
Availability of data, code, and materials
Gene expression datasets are available at the Gene Expression Omnibus public repository of NCBI under accession number GSE65682. Other data of the current study are available on reasonable request.
References
Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M et al (2016) The third international consensus definitions for sepsis and septic shock (Sepsis-3). JAMA 315(8):801–810
Rudd KE, Johnson SC, Agesa KM, Shackelford KA, Tsoi D, Kievlan DR et al (2020) Global, regional, and national sepsis incidence and mortality, 1990–2017: analysis for the Global Burden of Disease Study. Lancet 395(10219):200–211
Angus DC, Linde-Zwirble WT, Lidicker J, Clermont G, Carcillo J, Pinsky MR (2001) Epidemiology of severe sepsis in the United States: analysis of incidence, outcome, and associated costs of care. Crit Care Med 29(7):1303–1310
Vincent JL, Marshall JC, Namendys-Silva SA, Francois B, Martin-Loeches I, Lipman J et al (2014) Assessment of the worldwide burden of critical illness: the Intensive Care Over Nations (ICON) audit. Lancet Respir Med 2(5):380–386
Yende S, Austin S, Rhodes A, Finfer S, Opal S, Thompson T et al (2016) Long-term quality of life among survivors of severe sepsis: analyses of two international trials. Crit Care Med 44(8):1461–1467
Cohen J, Vincent JL, Adhikari NK, Machado FR, Angus DC, Calandra T et al (2015) Sepsis: a roadmap for future research. Lancet Infect Dis 15(5):581–614
Perner A, Gordon AC, Angus DC, Lamontagne F, Machado F, Russell JA et al (2017) The intensive care medicine research agenda on septic shock. Intensive Care Med 43(9):1294–1305
Marshall JC (2014) Why have clinical trials in sepsis failed? Trends Mol Med 20(4):195–203
Talisa VB, Yende S, Seymour CW, Angus DC (2018) Arguing for adaptive clinical trials in sepsis. Front Immunol 9:1502
Stanski NL, Wong HR (2020) Prognostic and predictive enrichment in sepsis. Nat Rev Nephrol 16(1):20–31
Seymour CW, Kennedy JN, Wang S, Chang CH, Elliott CF, Xu Z et al (2019) Derivation, validation, and potential treatment implications of novel clinical phenotypes for sepsis. JAMA J Am Med Assoc 321(20):2003–2017
Calfee CS, Delucchi K, Parsons PE, Thompson BT, Ware LB, Matthay MA (2014) Subphenotypes in acute respiratory distress syndrome: latent class analysis of data from two randomised controlled trials. Lancet Respir Med 2(8):611–620
Sinha P, Kerchberger VE, Willmore A, Chambers J, Zhuo H, Abbott J et al (2023) Identifying molecular phenotypes in sepsis: an analysis of two prospective observational cohorts and secondary analysis of two randomised controlled trials. Lancet Respir Med. https://doi.org/10.1016/S2213-2600(23)00237-0
Scicluna BP, van Vught LA, Zwinderman AH, Wiewel MA, Davenport EE, Burnham KL et al (2017) Classification of patients with sepsis according to blood genomic endotype: a prospective cohort study. Lancet Respir Med 5(10):816–826
Davenport EE, Burnham KL, Radhakrishnan J, Humburg P, Hutton P, Mills TC et al (2016) Genomic landscape of the individual host response and outcomes in sepsis: a prospective cohort study. Lancet Respir Med 4:259
DeMerle KM, Angus DC, Baillie JK, Brant E, Calfee CS, Carcillo J et al (2021) Sepsis subclasses: a framework for development and interpretation. Crit Care Med 49(5):748–759
Calfee CS, Delucchi KL, Sinha P, Matthay MA, Hackett J, Shankar-Hari M et al (2018) Acute respiratory distress syndrome subphenotypes and differential response to simvastatin: secondary analysis of a randomised controlled trial. Lancet Respir Med 6(9):691–698
Famous KR, Delucchi K, Ware LB, Kangelaris KN, Liu KD, Thompson BT et al (2017) Acute respiratory distress syndrome subphenotypes respond differently to randomized fluid management strategy. Am J Respir Crit Care Med 195(3):331–338
Sinha P, Delucchi KL, Thompson BT, McAuley DF, Matthay MA, Calfee CS et al (2018) Latent class analysis of ARDS subphenotypes: a secondary analysis of the statins for acutely injured lungs from sepsis (SAILS) study. Intensive Care Med 44(11):1859–1869
Antcliffe DB, Burnham KL, Al-Beidh F, Santhakumaran S, Brett SJ, Hinds CJ et al (2019) Transcriptomic signatures in sepsis and a differential response to steroids from the VANISH randomized trial. Am J Respir Crit Care Med 199(8):980–986
Sinha P, Delucchi KL, McAuley DF, O’Kane CM, Matthay MA, Calfee CS (2020) Development and validation of parsimonious algorithms to classify acute respiratory distress syndrome phenotypes: a secondary analysis of randomised controlled trials. Lancet Respir Med 8(3):247–257
Cano-Gamez E, Burnham KL, Goh C, Allcock A, Malick ZH, Overend L et al (2022) An immune dysfunction score for stratification of patients with acute infection based on whole-blood gene expression. Sci Transl Med 14(669):eabq4433
Davenport EE, Burnham KL, Radhakrishnan J, Humburg P, Hutton P, Mills TC et al (2016) Genomic landscape of the individual host response and outcomes in sepsis: a prospective cohort study. Lancet Respir Med 4(4):259–271
Kim HY (2017) Statistical notes for clinical researchers: Chi-squared test and Fisher’s exact test. Restor Dent Endod 42(2):152–155
Yu G, He QY (2016) ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol Biosyst 12(2):477–479
Gillespie M, Jassal B, Stephan R, Milacic M, Rothfels K, Senff-Ribeiro A et al (2022) The reactome pathway knowledgebase 2022. Nucleic Acids Res 50(D1):D687–D692
Zhao H, Kennedy JN, Wang S, Brant EB, Bernard GR, DeMerle K et al (2021) Revising host phenotypes of sepsis using microbiology. Front Med (Lausanne) 8:775511
Heijnen NFL, Hagens LA, Smit MR, Cremer OL, Ong DSY, van der Poll T et al (2021) Biological subphenotypes of ARDS show prognostic enrichment in mechanically ventilated patients without ARDS. Am J Respir Crit Care Med 203:1503
van Amstel RBE, van Vught LA, Bos LDJ (2023) Integrating biology into clinical trial design. Curr Opin Crit Care 29(1):26–33
López de Maturana E, Alonso L, Alarcón P, Martín-Antoniano IA, Pineda S, Piorno L et al (2019) Challenges in the integration of omics and non-omics data. Genes (Basel) 10(3):238
Wu J, Vodovotz Y, Abdelhamid S, Guyette FX, Yaffe MB, Gruen DS et al (2021) Multi-omic analysis in injured humans: patterns align with outcomes and treatment responses. Cell Rep Med 2(12):100478
Sinha P, Spicer A, Delucchi KL, McAuley DF, Calfee CS, Churpek MM (2021) Comparison of machine learning clustering algorithms for detecting heterogeneity of treatment effect in acute respiratory distress syndrome: a secondary analysis of three randomised controlled trials. EBioMedicine 74:103697
Maslove DM, Tang B, Shankar-Hari M, Lawler PR, Angus DC, Baillie JK et al (2022) Redefining critical illness. Nat Med 28(6):1141–1148
Acknowledgements
We thank the MARS consortium for their extensive role in the original study. (MARS consortium Amsterdam UMC location University of Amsterdam : Friso M. de Beer, Lieuwe D. J. Bos, Gerie J. Glas, Arie J. Hoogendijk, Roosmarijn T. M. van Hooijdonk, Janneke Horn, Mischa A. Huson, Laura R. A. Schouten, Marcus J. Schultz, Brendon P. Scicluna, Marleen Straat, Lonneke A. van Vught, Luuk Wieske, Maryse A. Wiewel, Esther Witteveen. University Medical Center Utrecht: Marc J.M. Bonten, Olaf M. Cremer, David S.Y. Ong, Jos F. Frencken, Peter M.C. Klein Klouwenberg, Maria E. Koster‐Brouwer, Kirsten van de Groep, Diana M. Verboom).
Funding
Supported by the Center for Translational Molecular Medicine (http://www.ctmm.nl), project MARS (grant 04I-201). Lonneke A van Vught was supported by the Netherlands Organisation for Health Research and Development ZonMW (Nederlandse Organisatie voor Wetenschappelijk Onderzoek NWO) VENI grant 09150161910033 and the European Society of Clinical Microbiology and Infectious Diseases (ESCMID) Research Grant. Christopher W Seymour was funded in part by grants from the National Institutes of Health (R35GM119519, R21GM144851). The funders of the study had no role in design and conduct of the study; collection, management, analysis interpretation of the data or preparation, review, approval of the manuscript or the decision to submit.
Author information
Authors and Affiliations
Consortia
Contributions
All authors conceived and designed the study. RA, JK, BS, LB, CS, TP, OC, and LV participated in data collection. RA, JK and LV accessed and verified all the data. RA, JK, BS, HPS, JB, and LV performed the data analysis. RA, CS, TP and LV drafted the article. All authors participated in revision of the manuscript. All authors gave final approval for the version to be submitted. The corresponding author attests that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted.
Corresponding author
Ethics declarations
Conflicts of interest
The authors disclose that they do not have any potential conflicts of interest.
Ethical approval
Patients were included via an opt-out consent method approved by the institutional review boards of both hospitals (IRB No. 10-056C).
Consent to participate
Patients were included via an opt-out consent method approved by the institutional review boards of both hospitals (IRB No. 10-056C).
Consent to publish
Not applicable/all data used for the present study have been anonymized and the submission does not include information that may identify individual persons.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The members of the MARS consortium are listed in the Acknowledgements.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
van Amstel, R.B.E., Kennedy, J.N., Scicluna, B.P. et al. Uncovering heterogeneity in sepsis: a comparative analysis of subphenotypes. Intensive Care Med 49, 1360–1369 (2023). https://doi.org/10.1007/s00134-023-07239-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00134-023-07239-w