Abstract
Key message
A whole-genome resequencing-derived SNP dataset used for genome-wide association analysis revealed 196 loci significantly associated with drought stress based on root traits. Candidate genes identified in the regions of these loci include homologs of known drought resistance genes in A. thaliana.
Abstract
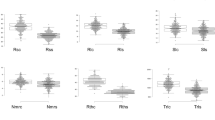
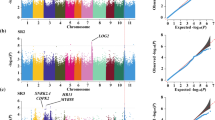
Drought is the main abiotic constraint of the production of common bean. Improved adaptation to drought environments has become a main goal of crop breeding due to the increasing scarcity of water that will occur in the future. The overall objective of our study was to identify genomic regions associated with drought resistance based on root traits using genome-wide association analysis. A natural population of 438 common bean accessions was evaluated for root traits: root surface area, root average diameter, root volume, total root length, taproot length, lateral root number, root dry weight, lateral root length, special root weight/length, using seed germination pouches under drought conditions and in well-watered environments. The coefficient of variation ranged from 11.24% (root average diameter) to 38.19% (root dry weight) in the well-watered environment and from 9.61% (root average diameter) to 39.05% (lateral root length) under drought stress. A whole-genome resequencing-derived SNP dataset revealed 196 loci containing 230 candidate SNPs associated with drought resistance. Seventeen candidate SNPs were simultaneously associated with more than two traits. Forty-one loci were simultaneously associated with more than two traits, and eleven loci were colocated with loci previously reported to be related to drought resistance. Candidate genes of the associated loci included the ABA-responsive element-binding protein family, MYB, NAC, the protein kinase superfamily, etc. These results revealed promising alleles linked to drought resistance or root traits, providing insights into the genetic basis of drought resistance and roots, which will be useful for common bean improvement.






Similar content being viewed by others
Availability of data and material
Data supporting the current study can be obtained by contacting the corresponding author (wangshumin@caas.cn).
Code availability
Not applicable.
Abbreviations
- GWAS:
-
Genome-wide association study
- SNP:
-
Single nucleotide polymorphisms
- CHR:
-
Chromosome
- LD:
-
Linkage disequilibrium
- QTL:
-
Quantitative trait loci
- ABA:
-
Abscisic acid
- RSA:
-
Root surface area
- RAD:
-
Root average diameter
- RV:
-
Root volume
- TRL:
-
Total root length
- TL:
-
Taproot length
- LRN:
-
Lateral root number
- RDW:
-
Root dry weight
- LRL:
-
Lateral root length
- SRL:
-
Special root weight/length
References
Armengaud P, Zambaux K, Hills A, Sulpice R, Pattison RJ, Blatt MR, Amtmann A (2009) EZ-Rhizo: integrated software for the fast and accurate measurement of root system architecture. Plant J 57:945–956
Asfaw A, Blair MW (2012) Quantitative trait loci for rooting pattern traits of common beans grown under drought stress versus non-stress conditions. Mol Breed 30:681–695
Asfaw A, Blair MW, Struik PC (2012) Multienvironment quantitative trait loci analysis for photosynthate acquisition accumulation and remobilization traits in common bean under drought stress. G3 2:579–595
Ashraf M (2010) Inducing drought tolerance in plant: recent advances. Biotechnol Advs 28:169–183
Beebe S (2012) Common bean breeding in the tropics. Plant Breed Rev 36:357–426
Beebe SE, Rojas-Pierce M, Yan X, Blair MW, Pedraza F, Munoz F, Tohme J, Lynch JP (2006) Quantitative trait loci for root architecture traits correlated with phosphorus acquisition in common bean. Crop Sci 46:413–423
Beebe S, Rao IM, Cajiao C, Grajales M (2008) Selection for drought resistance in common bean also improves yield in phosphorus limited and favorable environments. Crop Sci 48:582–592
Beebe SE, Rao IM, Blair MW, Acosta-Gallegos JA (2013) Phenotyping common beans for adaptation to drought. Front Physiol 4:1–20
Berny Mier Y, Teran JC, Konzen ER, Medina V, Palkovic A, Ariani A, Tsai SM, Gilbert ME, Gepts P (2019) Root and shoot variation in relation to potential intermittent drought adaptation of Mesoamerican wild common bean (Phaseolus vulgaris L.). Ann Bot 124:917–932
Blair MW, Galeano CH, Tovar E, Torres MCM, Castrillón AV, Beebe SE (2012) Development of a Mesoamerican intra-gene pool genetic map for quantitative trait loci detection in a drought tolerant. Mol Breed 29:71–88
Boris B, Cardoso PJMK, Santa RJ, Denis B, Ribeiro GJG, Caléo A (2017) Mapping QTLs for drought tolerance in a SEA5×AND277 common bean cross with SSRs and SNP markers. Genet Mol Bio 40:813–823
Broughton WJ, Hernandez G, Blair MW, Beebe SE, Gepts P, Vanderleyden J (2003) Beans (Phaseolus spp.)—model food legumes. Plant Soil 252:55–128
Cantero A, Barthakur S, Bushart TJ, Chou S, Morgan RO, Fernandez MP, Clark GB, Roux SJ (2006) Expression profiling of the Arabidopsis annexin gene family during germination, de-etiolation and abiotic stress. Plant Physiol Biochem 44:13–24
Chapman K, Taleski M, Ogilvie HA, Imin N, Djordjevic MA (2019) CEP-CEPR1 signalling inhibits the sucrose-dependent enhancement of lateral root growth. J Exp Bot 70:3955–3967
Chen JB, Yang JW, Zhang ZY, Feng XF, Wang SM (2013) Two P5CS genes from common bean exhibiting different tolerance to salt stress in transgenic Arabidopsis. J Genet 92:461–469
Cichy KA, Snapp SS, Blair MW (2009) Plant growth habit, root architecture traits and tolerance to low soil phosphorus in an Andean bean population. Euphytica 165:257–268
Cui M, Zhang W, Zhang Q, Xu Z, Zhu Z, Duan F, Wu R (2011) Induced over-expression of the transcription factor OsDREB2A improves drought tolerance in rice. Plant Physiol Biochem 49:1384–1391
Devaiah BN, Karthikeyan AS, Raghothama KG (2007) WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol 143:1789–1801
Ding Z, Yan J, Xu X, Yu D, Li G, Zhang S, Zheng S (2014) Transcription factor WRKY46 regulates osmotic stress responses and stomatal movement independently in Arabidopsis. Plant J 79:13–27
Dwivedi SL, Sahrawat KL, Rai KN, Blair MW, Pfeiffer W (2012) Nutritionally enhanced staple food crops. Plant Breed Rev 36:169–291
Emiliano VP, González-Chavira Mario M, Patricia GC, Acosta-Gallegos JA, Juan C-P (2015) Identification of novel drought-tolerant-associated SNPs in common bean (Phaseolus vulgaris). Front Plant Sci 6:546
Fang Y, Xiong L (2015) General mechanisms of drought response and their application in drought resistance improvement in plants. Cell Mol Life Sci 72:673–689
Frahm MA, Rosas JC, Mayek-Pérez N, López-Salinas E, Acosta-Gallegos JA, Kelly JD (2004) Breeding beans for resistance to terminal drought in the lowland tropics. Euphytica 136:223–232
Fujii H, Verslues PE, Zhu JK (2011) Arabidopsis decuple mutant reveals the importance of SnRK2 kinases in osmotic stress responses in vivo. Proc Natl Acad Sci U S A 108:1717–1722
Fujita Y, Fujita M, Satoh R, Maruyama K, Parvez MM, Seki M, Hiratsu K, Ohme-Takagi M, Shinozaki K, Yamaguchi-Shinozaki K (2005) AREB1 is a transcription activator of novel ABRE-dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 17:3470–3488
Galinha C, Hofhuis H, Luijten M, Willemsen V, Blilou I, Heidstra R, Scheres B (2007) PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 449:1053–1057
German MA, Burdman S, Okon Y, Kigel J (2000) Effects of Azospirillum brasilense on root morphology of common bean (Phaseolus vulgaris L.) under different water regimes. Biol Fert Soils 32:259–264
Gupta A, Rico-Medina A, Caño-Delgado AI (2020) The physiology of plant responses to drought. Science 368:266–269
Hagerty CH, Cuesta-Marcos A, Cregan PB, Song Q, McClean P, Noffsinger S, Myers JR (2015) Mapping and root rot resistance and root architecture quantitative trait loci in common bean. Crop Sci 55:1969–1977
He G, Xu J, Wang Y, Liu J, Li P, Chen M, Ma Y, Xu Z (2016) Drought-responsive WRKY transcription factor genes TaWRKY1 and TaWRKY33 from wheat confer drought and/or heat resistance in Arabidopsis. BMC Plant Biol 16:116
Hoyos-Villegas V, Song Q, Kelly JD (2017) Genome-wide association analysis for drought tolerance and associated traits in common bean. Plant Genome. https://doi.org/10.3835/plantgenome2015.12.0122
Hu Z, Wang R, Zheng M, Liu X, Sun Q (2018) TaWRKY51 promotes lateral root formation through negatively regulating ethylene biosynthesis in wheat (Triticum aestivum L.). Plant J 96:372–388
Huang B, Gao H (2000) Root physiological characteristics associated with drought resistance in tall fescue cultivars. Crop Sci 40:196–203
Jin X, Xue Y, Wang R, Xu R, Bian L, Zhu B, Han H, Peng R, Yao Q (2012) Transcription factor OsAP21 gene increases salt/drought tolerance in transgenic Arabidopsis thaliana. Mol Biol Rep 40:1743–1752
Kamoshita A, Babu RC, Manikanda Boopathi N, Fukai S (2008) Phenotypic and genotypic analysis of drought-resistance traits for development of rice cultivars adapted to rainfed environments. Field Crops Res 109:1–23
Kim JH, Nguyen NH, Jeong CY, Nguyen NT, Hong SW, Lee H (2013) Loss of the R2R3 MYB, AtMyb73, causes hyper-induction of the SOS1 and SOS3 genes in response to high salinity in Arabidopsis. J Plant Physiol 170:1461–1465
Lata C, Mishra AK, Muthamilarasan M, Bonthala VS, Khan Y, Prasad M (2014) Genome-wide investigation and expression profiling of AP2/ERF transcription factor superfamily in foxtail millet (Setaria italica L.). PLoS ONE 9:e113092
Lee SB, Suh MC (2015) Cuticular wax biosynthesis is up-regulated by the MYB94 transcription factor in Arabidopsis. Plant Cell Physiol 56:48–60
Li J, Kinoshita T, Pandey S, Ng KY, Gygi SP, Shimazaki KI (2002) Modulation of an RNA-binding protein by abscisic-acid-activated protein kinase. Nature 418:793–797
Li F, Asami T, Wu X, Tsang EW, Cutler AJ (2007) A putative hydroxysteroid dehydrogenase involved in regulating plant growth and development. Plant Physiol 145:87–97
Li L, Wang L, Wu J, Jing R, Wang S (2013) Drought tolerance in common bean germplasm at bud stage. J Plant Genet Res 14:600–605
Li X, Guo Z, Lv Y, Cen X, Ding X, Wu H, Li X, Huang J, Xiong L (2017) Genetic control of the root system in rice under normal and drought stress conditions by genome-wide association study. PLoS Genet 13:e1006889
Li L, Mao X, Wang J, Chang X, Matthew R, Jing R (2019a) Genetic dissection of drought and heat-responsive agronomic traits in wheat. Plant Cell Environ 42:1–14
Li L, Peng Z, Mao X, Wang J, Chang X, Matthew R, Jing R (2019b) Genome-wide association study reveals genomic regions controlling root and shoot traits at late growth stages in wheat. Ann Bot 124:993–1006
Li Z, Liu C, Zhang Y, Wang B, Ran Q, Zhang J (2019c) The bHLH family member ZmPTF1 regulates drought tolerance in maize by promoting root development and abscisic acid synthesis. J Exp Bot 70:5471–5486
Liao H, Rubio G, Yan X, Cao A, Brown KM, Lync JP (2001) Effect of phosphorus availability on basal root shallowness in common bean. Plant Soil 232:69–79
Liao H, Yan X, Rubio G, Beebe S, Blair M, Lynch J (2004) Genetic mapping of basal root gravitropism and phosphorus acquisition efficiency in common bean. Funct Plant Biol 33:959–970
Lin C, Choi HS, Cho HT (2011) Root hair-specific EXPANSIN A7 is required for root hair elongation in Arabidopsis. Mol Cells 3:393–397
Liu Y, Wu Y, Huang X, Sun J, Xie Q (2011) AtPUB19, a U-box E3 ubiquitin ligase, negatively regulates abscisic acid and drought responses in Arabidopsis thaliana. Mol Plant 4:938–946
Liu W, Tai H, Li S, Gao W, Zhao M, Xie C, Li W (2014) bHLH122 is important for drought and osmotic stress resistance in Arabidopsis and in the repression of ABA catabolism. New Phytol 201:1192–1204
Mabuchi K, Maki H, Itaya T, Suzuki T, Nomoto M, Sakaoka S, Morikami A, Higashiyama T, Tada Y, Busch W, Tsukagoshi H (2018) MYB30 links ROS signaling, root cell elongation, and plant immune responses. Proc Natl Acad Sci U S A 115:E4710–E4719
Mao X, Chen S, Li A, Zhai C, Jing R (2014) Novel NAC transcription factor TaNAC67 confers enhanced multi-abiotic stress tolerances in Arabidopsis. PLoS ONE 9:e84359
Mao H, Wang H, Liu S, Li Z, Yang X, Yan J, Li J, Tran LS, Qin F (2015) A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat Commun 6:8326
Mukeshimana G, Butare G, Cregan P, Blair MW, Kelly JD (2014) Identification of quantitative trait loci associated with drought tolerance in common bean using SNP markers. Crop Sci 54:923–938
Mun˜oz-Perea CG, Teran H, Allen RG, Wright JL, Westermann DT, Singh SP (2006) Selection for drought resistance in dry bean landraces and cultivars. Crop Sci 46:2111–2120
Ochoa I, Blair MW, Lynch J (2006) QTL analysis of adventitious root formation in common bean under contrasting phosphorus availability. Crop Sci 46:1609–1621
Ortiz-Masia D, Perez-Amador MA, Carbonell J, Marcote MJ (2007) Diverse stress signals activate the C1 subgroup MAP kinases of Arabidopsis. FEBS Lett 2581:1834–1840
Park MY, Kim SY (2014) The Arabidopsis J protein AtJ1 is essential for seedling growth, flowering time control and ABA response. Plant Cell Physiol 55:2152–2163
Park HY, Seok HY, Park BK, Kim SH, Goh CH, Lee BH, Lee CH, Moon YH (2008) Overexpression of Arabidopsis ZEP enhances tolerance to osmotic stress. Biochem Biophys Res Commun 375:80–85
Piao S, Ciais P, Huang Y, Shen Z, Peng S, Li J (2010) The impacts of climate change on water resources and agriculture in china. Nature 467:43–51
Podia V, Milioni D, Martzikou M, Haralampidis K (2018) The role of Arabidopsis thaliana RASD1 gene in ABA-dependent abiotic stress response. Plant Biol (Stuttg) 20:307–317
Priest DM, Ambrose SJ, Vaistij Fabián E, Elias L, Bowles DJ (2010) Use of the glucosyltransferase UGT71B6 to disturb abscisic acid homeostasis in Arabidopsis thaliana. Plant J 46:492–502
Pruthvi V, Narasimhan R, Nataraja KN (2014) Simultaneous expression of abiotic stress responsive transcription factors, AtDREB2A, AtHB7 and AtABF3 improves salinity and drought tolerance in peanut (Arachis hypogaea L.). PLoS One 9:e111152
Qin Y, Tian Y, Liu X (2015) A wheat salinity-induced WRKY transcription factor TaWRKY93 confers multiple abiotic stress tolerance in Arabidopsis thaliana. Biochem Biophys Res Commun 464:428–433
Ramegowda V, Gill US, Sivalingam PN, Gupta A, Gupta C, Govind G, Nataraja KN, Pereira A, Udayakumar M, Mysore KS, Senthil-Kumar M (2017) GBF3 transcription factor imparts drought tolerance in Arabidopsis thaliana. Sci Rep 7:9148
Rao IM (2001) Role of physiology in improving crop adaptation to abiotic stresses in the tropics: the case of common bean and tropical forages. In: Pessarakli M (ed) Handbook of plant and crop physiology. Marcel Dekker Inc, New York, pp 583–613
Ré DA, Capella M, Bonaventure G (2014) Arabidopsis AtHB7 and AtHB12 evolved divergently to fine tune processes associated with growth and responses to water stress. BMC Plant Biol 14:1–14
Remans R, Beebe S, Blair MW, Manrique G, Tovar E, Rao IM, Croonenborghs A, Torres-Gutierrez R, El-Howeity M, Michiels J, Vanderleyden J (2008) Physiological and genetic analyses of root responsiveness to auxin-producing plant growth-promoting bacteria in common bean (Phaseolus vulgaris L.). Plant Soil 302:149–161
Schmutz J, McClean PE, Mamidi S, Wu GA, Cannon SB, Grimwood J, Jenkins J, Shu S, Song Q, Chavarro C, Torres-Torres M, Geffroy V, Moghaddam SM, Gao D, Abernathy B, Barry K, Blair M, Brick MA, Chovatia M, Gepts P, Goodstein DM, Gonzales M, Hellsten U, Hyten DL, Jia G, Kelly JD, Kudrna D, Lee R, Richard MM, Miklas PN, Osorno JM, Rodrigues J, Thareau V, Urrea CA, Wang M, Yu Y, Zhang M, Wing RA, Cregan PB, Rokhsar DS, Jackson SA (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713
Schneider KA, Brothers ME, Kelly JD (1997) Marker-assisted selection to improve drought resistance in common bean. Crop Sci 37:51
Söderman E, Mattsson J, Engstrm P (1996) The Arabidopsis homeobox gene ATHB-7 is induced by water deficit and by abscisic acid. Plant J 10:375–381
Shin R, Burch AY, Huppert KA, Tiwari SB, Murphy AS, Guilfoyle TJ, Schachtman DP (2007) The Arabidopsis transcription factor MYB77 modulates auxin signal transduction. Plant Cell 19:2440–2453
Smucker AJM, Nunez-Barrios A, Ritchie JT (1991) Root dynamics in drying soil environments. Below-ground Ecol 1:1–5
Sponchiado BN, White JW, Castillo JA, Jones PG (1989) Root growth of four common bean cultivars in relation to drought tolerance in environments with contrasting soil types. Exp Agr 25:249–257
Strizhov N, Abrahám E, Okrész L, Blickling S, Zilberstein A, Schell J, Koncz C, Szabados L (1997) Differential expression of two P5CS genes controlling proline accumulation during salt-stress requires ABA and is regulated by ABA1, ABI1 and AXR2 in Arabidopsis. Plant J 12:557–569
Sun Z, Wang X, Liu Z, Gu Q, Zhang Y, Li Z, Ke H, Yang J, Wu J, Wu L (2017) Genome-wide association study discovered genetic variation and candidate genes of fibre quality traits in Gossypium hirsutum L. Plant Biotechnol J 15:982–996
Sura W, Kabza M, Karlowski WM, Bieluszewski T, Kus-Slowinska M, Pawełoszek Ł, Sadowski ŁJ, Ziolkowski PA (2017) Dual role of the histone variant H2A.Z in transcriptional regulation of stress-response genes. Plant Cell 29:791–807
Thung M, Rao IM (1999) Integrated management of abiotic stresses. In: Singh SP (ed) Common bean improvement in the twenty-first century. Kluwer, Dordrecht, pp 331–370
Trapp JJ, Urrea CA, Cregan PB, Miklas PN (2015) Quantitative trait loci for yield under multiple stress and drought conditions in a dry bean population. Crop Sci 55:1596
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci U S A 97:11632–11637
Wang R, Wu H, Zhang M, Ni Z, Sun Q (2013) Cloning, characterization and transgenic function analysis of wheat (Triticum aestivum L.) TaWRKY51 gene. JAB 21:1019–1027
Wang X, Jia N, Zhao C, Fang Y, Lv T, Zhou W, Sun Y, Li B (2014) Knockout of AtDjB1, a J-domain protein from Arabidopsis thaliana, alters plant responses to osmotic stress and abscisic acid. Physiol Plant 152:286–300
Wang X, Wang H, Liu S, Ferjani A, Li J, Yan J, Yang X, Qin F (2016) Genetic variation in ZmVPP1 contributes to drought tolerance in maize seedlings. Nat Genet 48:1233–1241
Wang Y, Wang Q, Liu M, Bo C, Cai R (2017a) Overexpression of a maize MYB48 gene confers drought tolerance in transgenic Arabidopsis plants. J Plant Biol 60:612–621
Wang N, Zhang W, Qin M, Li S, Qiao M, Liu Z, Xiang F (2017b) Drought tolerance conferred in soybean (Glycine max. L) by GmMYB84, a novel R2R3-MYB transcription factor. Plant Cell Physiol 58:1764–1776
Wen Z, Tan R, Zhang S, Collins P, Yuan J, Du W, Gu C, Ou S, Song Q, An Y (2018) Integrating GWAS and gene expression data for functional characterization of resistance to white mould in soya bean. Plant Biotechnol J 16:1825–1835
Wu J, Wang L, Li L, Wang S (2014) De novo assembly of the common bean transcriptome using short reads for the discovery of drought-responsive genes. PLoS ONE 9:e109262
Wu J, Wang L, Fu J, Chen J, Wei S, Zhang S, Zhang J, Tang Y, Chen M, Zhu J, Lei L, Geng Q, Liu C, Wu L, Li X, Wang Q, Wang Z, Xing S, Zhang H, Blair M, Wang S (2020) Resequencing of 683 common bean genotypes identifies yield component trait associations across a north-south cline. Nat Genet 52:118–125
Xi J, Qiu Y, Du L, Poovaiah BW (2012) Plant-specific trihelix transcription factor AtGT2l interacts with calcium/calmodulin and responds to cold and salt stresses. Plant Sci 185–186:274–280
Xing DH, Lai ZB, Zheng ZY, Vinod KM, Fan BF, Chen ZX (2008) Stress- and pathogen-induced Arabidopsis WRKY48 is a transcriptional activator that represses plant basal defense. Mol Plant 1:459–470
Xiong L, Lee H, Ishitani M, Zhu JK (2002) Regulation of osmotic stress-responsive gene expression by the LOS6/ABA1 locus in Arabidopsis. J Biol Chem 277:8588–8596
Xiong H, Li J, Liu P, Duan J, Zhao Y, Guo X, Li Y, Zhang H, Ali J, Li Z (2014) Overexpression of OsMYB48-1, a novel MYB-related transcription factor, enhances drought and salinity tolerance in rice. PLoS ONE 9:e92913
Yan X, Liao H, Beebe SE, Blair MW, Lynch JP (2004) QTL mapping of root hair and acid exudation traits and their relationship to phosphorus uptake in common bean. Plant Soil 265:17–29
Yang SD, Seo PJ, Yoon HK, Park CM (2011) The Arabidopsis NAC transcription factor VNI2 integrates abscisic acid signals into leaf senescence via the COR/RD genes. Plant Cell 23:2155–2168
Yordanov I, Velikova V, Tsonev T (2000) Plant responses to drought, acclimation, and stress tolerance. Photosyntheetica 38:171–186
Zhang J, Guo X, Li X, Xiang F, Zhou B, Yu D, Tang D, Liu X (2012) The genetic and physiological analysis of late-flowering phenotype of T-DNA insertion mutants of AtCAL1 and AtCAL2 in Arabidopsis. Mol Biol Rep 39:1527–1535
Zhao Y, Xing L, Wang X, Hou YJ, Gao J, Wang P, Duan CG, Zhu X, Zhu JK (2014) The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes. Sci Signal 7:ra53
Zhao K, Zhang D, Lv K, Zhang X, Cheng Z, Li R, Zhou B, Jiang T (2019) Functional characterization of poplar WRKY75 in salt and osmotic tolerance. Plant Sci 289:110259
Zheng Y, Schumaker KS, Guo Y (2012) Sumoylation of transcription factor MYB30 by the small ubiquitin-like modifier E3 ligase SIZ1 mediates abscisic acid response in Arabidopsis thaliana. Proc Natl Acad Sci U S A 109:12822–12827
Funding
This work was supported by grants from the National Key R & D Program of China (2018YFD1000700/2018YFD1000704), the Ministry of Agriculture of China [the earmarked fund for the China Agriculture Research System (CARS-08)], the Agricultural Science and Technology Innovation Program of CAAS.
Author information
Authors and Affiliations
Contributions
L W, YJ C performed QTL analysis and characterized candidate genes. L W, LF W, J W and SM W designed and conducted the field experiments and generated phenotype data. L W, YJ C and J W wrote and finalized the manuscript. SM W planned and organized this study, and co-wrote the manuscript. All authors contributed to and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Communicated by Janila Pasupuleti.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, L., Chang, Y., Wang, L. et al. Genetic dissection of drought resistance based on root traits at the bud stage in common bean. Theor Appl Genet 134, 1047–1061 (2021). https://doi.org/10.1007/s00122-020-03750-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03750-6