Abstract
Key message
BSA-seq combined with whole-genome resequencing map-based cloning delimited the cucumber det-novel locus into a 44.5 kb region in chromosome 6 harboring a putative candidate gene encoding a phosphatidylethanolamine-binding protein (CsCEN).
Abstract
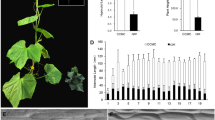
Determinate and indeterminate growth habits of cucumber can affect plant architecture and crop yield. The TERMINAL FLOWER 1 (TFL1) controls determinate/indeterminate growth in Arabidopsis. In this study, a novel mutation in cucumber TFL1 homolog (CsCEN) has shown to regulate determinate growth and product of terminal flowers in cucumber (Cucumis sativus L.), which is similar to the function of CsTFL1 as previously reported. Genetic analysis in two determinate genotypes (D226 and D082) and indeterminate genotype (CCMC) revealed that a single recessive gene is responsible for this determinate growth trait. With the combination of BSA-seq and whole-genome resequencing, the locus of determinate-novel (det-novel) trait was mapped to a 44.5 kb genomic region in chromosome 6. Sequence alignment identified one non-synonymous SNP mutation (A to T) in the third exon of CsCEN, resulting in an amino acid substitution (Thr to Pro), suggesting that determinate growth might be controlled by a novel gene CsCEN (Csa6G152360) which differed from the reported CsTFL1 gene. The CsCEN expression level in shoot apexes and axillary buds was significantly lower in D226 compared to CCMC, suggesting its essential role in sustaining indeterminate growth habit. Identification and characterization of the CsCEN in the present study provide a new insight into plant architecture modification and development of cucumber cultivars suited to mechanized production system.




Similar content being viewed by others
References
Birnbaum KD, Alvarado AS (2008) Slicing across kingdoms: regeneration in plants and animals. Cell 132:697–710
Bradley D, Carpenter R, Copsey L, Vincent C, Rothstein S, Coen E (1996) Control of inflorescence architecture in Antirrhinum. Nature 379:791–797
Bradley D, Ratcliffe O, Vincent C, Carpenter R, Coen E (1997) Inflorescence commitment and architecture in Arabidopsis. Science 275:80–83
Çagirgan M (2006) Selection and morphological characterization of induced determinate mutants in sesame. Field Crop Res 96:13–18
Carmel-Goren L, Liu YS, Lifschitz E, Zamir D (2003) The SELF-PRUNING gene family in tomato. Plant Mol Biol 52:1215–1222
Fazio G, Staub J, Stevens M (2003) Genetic mapping and QTL analysis of horticultural traits in cucumber (Cucumis sativus L.) using recombinant inbred lines. Theor Appl Genet 107:864–874
Foucher F, Morin J, Courtiade J, Cadioux S, Ellis N, Banfield MJ, Rameau C (2003) DETERMINATE and LATE FLOWERING are two TERMINAL FLOWER1/CENTRORADIALIS homologs that control two distinct phases of flowering initiation and development in pea. Plant Cell 15:2742–2754
Hanano S, Goto K (2011) Arabidopsis TERMINAL FLOWER1 is involved in the regulation of flowering time and inflorescence development through transcriptional repression. Plant Cell 23:3172–3184
Hanzawa Y, Money T, Bradley D (2005) A single amino acid converts a repressor to an activator of flowering. Proc Natl Acad Sci USA 102:7748–7753
Huang SW, Li RQ, Zhang ZH, Li L, Gu XF et al (2009) The genome of the cucumber, Cucumis sativus L. Nat Genet 41:1275–1281
Koinange EM, Singh SP, Gepts P (1996) Genetic control of the domestication syndrome in common bean. Crop Sci 36:1037–1045
Landrein B, Refahi Y, Besnard F, Hervieux N, Mirabet V, Boudaoud A, Vernoux T, Hamant O (2015) Meristem size contributes to the robustness of phyllotaxis in Arabidopsis. J Exp Bot 66:1317–1324
Liu B, Watanabe S, Uchiyama T, Kong FJ, Kanazawa A et al (2010) The soybean stem growth habit gene Dt1 is an ortholog of Arabidopsis TERMINAL FLOWER1. Plant Physiol 153(1):198–210
Nakagawa M, Shimamoto K, Kyozuka J (2002) Overexpression of RCN1 and RCN2, rice TERMINAL FLOWER 1/CENTRORADIALIS homologs, confers delay of phase transition and altered panicle morphology in rice. Plant J 29:743–750
Pañeda A, Rodríguez-Suárez C, Campa A, Ferreira JJ, Giraldez R (2008) Molecular markers linked to the fin gene controlling determinate growth habit in common bean. Euphytica 162:241–248
Périlleux C, Bouché F, Randoux M, Orman-Ligeza B (2019) Turning meristems into fortresses. Trends Plant Sci 24:431–442
Pierce LK, Wehner TC (1990) Review of genes and linkage groups in cucumber. HortScience 25:605–615
Pnueli L, Carmel-Goren L, Hareven D, Gutfinger T, Alvarez J, Ganal M, Zamir D, Lifschitz E (1998) The SELF-PRUNING gene of tomato regulates vegetative to reproductive switching of sympodial meristems and is the ortholog of CEN and TFL1. Development 125:1979–1989
Pnueli L, Gutfinger T, Hareven D, Ben-Naim O, Ron N, Adir N, Lifschitz E (2001) Tomato SP-interacting proteins define a conserved signaling system that regulates shoot architecture and flowering. Plant Cell 13:2687–2702
Sato H, Heang D, Sassa H, Koba T (2009) Identification and characterization of FT/TFL1 gene family in cucumber. Breed Sci 59:3–11
Shannon S, Meeks-Wagner DR (1991) A mutation in the Arabidopsis TFL1 gene affects inflorescence meristem development. Plant Cell 3:877–892
Soyk S, Müller NA, Park SJ, Schmalenbach I, Jiang K, Hayama R, Zhang L, Van EJ, Jiménez-Gómez JM, Lippman Z (2017) Variation in the flowering gene SELF PRUNING 5G promotes day-neutrality and early yield in tomato. Nat Genet 49:162–168
Tel-Zur N, Abbo S, Myslabodski D, Mizrahi Y (1999) Modified CTAB procedure for DNA isolation from epiphytic cacti of the genera Hylocereus and Selenicereus (Cactaceae). Plant Mol Biol Rep 17:249–254
Wang YH, Li JY (2008) Molecular basis of plant architecture. Annu Rev Plant Biol 59(1):253–279
Wen CL, Zhao WS, Liu WL, Yang LM, Wang YH et al (2019) CsTFL1 inhibits determinate growth and terminal flower formation through interaction with CsNOT2a in cucumber. Development 146:dev180166
Weng Y, Johnson S, Staub JE, Huang S (2010) An extended intervarietal microsatellite linkage map of cucumber, Cucumis sativus L. Am Soc Hortic Sci 45:882–886
Wicart G, Mouras A, Lutz A (1984) Histological study of organogenesis and embryogenesis in Cyclamen persicum Mill. tissue cultures: evidence for a single organogenetic pattern. Protoplasma 119:159–167
Zhang D, Yuan Z (2014) Molecular control of grass inflorescence development. Annu Rev Plant Biol 65:553–578
Zhang Y, Wang L, Gao Y, Li D, Yu J, Zhou R, Zhang X (2018) Genetic dissection and fine mapping of a novel dt gene associated with determinate growth habit in sesame. BMC Genet 19:38–48
Zhao WS, Gu R, Che G, Cheng Z, Zhang X et al (2018) CsTFL1b may regulate the flowering time and inflorescence architecture in cucumber (Cucumis sativas L.). Biochem Biophys Res Commun 499:307–313
Acknowledgements
A Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions and the Belt—Road project by Jiangsu Provincial Government; Also partly funded by National Key Research and Development Program of China (2018YFD1000804), Jiangsu Agricultural Innovation of New Cultivars ‘Innovation and application of cucumber cultivars with high quality and multi disease resistances (No. PZCZ201719),’ National Natural Science Foundation of China (No. 31672168) and Natural Science Foundation of Jiangsu Province (BK20191312).
Author information
Authors and Affiliations
Contributions
JFC and JOO conceived and supervised the projects. MKN, FY and XYW collected phenotypic data from field. MKN and FY performed the experiments. MKN, FY and JL analyzed the data and wrote manuscript. JL, JFC and JOO contributed to revising the manuscript. All authors reviewed and contributed in drafting the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical standards
The experiments were performed in accordance with all relevant Chinese laws.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Njogu, M.K., Yang, F., Li, J. et al. A novel mutation in TFL1 homolog sustaining determinate growth in cucumber (Cucumis sativus L.). Theor Appl Genet 133, 3323–3332 (2020). https://doi.org/10.1007/s00122-020-03671-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03671-4