Abstract
Key message
Discovery of new germplasm sources and identification of haplotypes for the durable Soybean mosaic virus resistance gene, Rsv 4, provide novel resources for map-based cloning and genetic improvement efforts in soybean.
Abstract
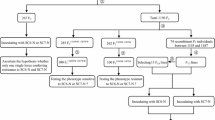
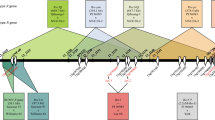
The Soybean mosaic virus (SMV) resistance locus Rsv4 is of interest because it provides a durable type of resistance in soybean [Glycine max (L.) Merr.]. To better understand its molecular basis, we used a population of 309 BC3F2 individuals to fine-map Rsv4 to a ~120 kb interval and leveraged this genetic information in a second study to identify accessions ‘Haman’ and ‘Ilpumgeomjeong’ as new sources of Rsv4. These two accessions along with three other Rsv4 and 14 rsv4 accessions were used to examine the patterns of nucleotide diversity at the Rsv4 region based on high-depth resequencing data. Through a targeted association analysis of these 19 accessions within the ~120 kb interval, a cluster of four intergenic single-nucleotide polymorphisms (SNPs) was found to perfectly associate with SMV resistance. Interestingly, this ~120 kb interval did not contain any genes similar to previously characterized dominant disease resistance genes. Therefore, a haplotype analysis was used to further resolve the association signal to a ~94 kb region, which also resulted in the identification of at least two Rsv4 haplotypes. A haplotype phylogenetic analysis of this region suggests that the Rsv4 locus in G. max is recently introgressed from G. soja. This integrated study provides a strong foundation for efforts focused on the cloning of this durable virus resistance gene and marker-assisted selection of Rsv4-mediated SMV resistance in soybean breeding programs.



Similar content being viewed by others
References
Agresti A (2013) Categorical data analysis, 3rd edn. Wiley, Hoboken
Ahlquist P, Noueiry AO, Lee W-M, Kushner DB, Dye BT (2003) Host factors in positive-strand RNA virus genome replication. J Virol 77:8181–8186. doi:10.1128/JVI.77.15.8181-8186.2003
Ashfield T, Egan AN, Pfeil BE, Chen NWG, Podicheti R, Ratnaparkhe MB, Ameline-Torregrosa C, Denny R, Cannon S, Doyle JJ, Geffroy V, Roe BA, Saghai Maroof MA, Young ND, Innes RW (2012) Evolution of a complex disease resistance gene cluster in diploid Phaseolus and tetraploid Glycine. Plant Physiol 159:336–354. doi:10.1104/pp.112.195040
Bandelt H-J, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265. doi:10.1093/bioinformatics/bth457
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol 57:289–300
Büschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, van Daelen R, van der Lee T, Diergaarde P, Groenendijk J, Töpsch S, Vos P, Salamini F, Schulze-Lefert P (1997) The barley Mlo gene: a novel control element of plant pathogen resistance. Cell 88:695–705. doi:10.1016/S0092-8674(00)81912-1
Buzzell RI, Tu JC (1989) Inheritance of a soybean stem-tip necrosis reaction to soybean mosaic virus. J Hered 80:400–401
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. doi:10.1186/1471-2105-10-421
Chen P, Buss GR, Roane CW, Tolin SA (1991) Allelism among genes for resistance to soybean mosaic virus in strain-differential soybean cultivars. Crop Sci 31:305. doi:10.2135/cropsci1991.0011183X003100020015x
Chisholm ST, Mahajan SK, Whitham SA, Yamamoto ML, Carrington JC (2000) Cloning of the Arabidopsis RTM1 gene, which controls restriction of long-distance movement of tobacco etch virus. Proc Natl Acad Sci USA 97:489–494
Cho E-K, Goodman RM (1979) Strains of soybean mosaic virus: classification based on virulence in resistant soybean cultivars. Phytopathology 69:467–470
Cho EK, Goodman RM (1982) Evaluation of resistance in soybeans to soybean mosaic virus strains. Crop Sci 22:1133. doi:10.2135/cropsci1982.0011183X002200060012x
Choi BK, Koo JM, Ahn HJ, Yum HJ, Choi CW, Ryu KH, Chen P, Tolin SA (2005) Emergence of Rsv-resistance breaking soybean mosaic virus isolates from Korean soybean cultivars. Virus Res 112:42–51. doi:10.1016/j.virusres.2005.03.020
Chung W-H, Jeong N, Kim J, Lee WK, Lee Y-G, Lee S-H, Yoon W, Kim J-H, Choi I-Y, Choi H-K, Moon J-K, Kim N, Jeong S-C (2014) Population structure and domestication revealed by high-depth resequencing of Korean cultivated and wild soybean genomes. DNA Res 21:153–167. doi:10.1093/dnares/dst047
Cook DE, Lee TG, Guo X, Melito S, Wang K, Bayless AM, Wang J, Hughes TJ, Willis DK, Clemente TE, Diers BW, Jiang J, Hudson ME, Bent AF (2012) Copy number variation of multiple genes at Rhg1 mediates nematode resistance in soybean. Science 338:1206–1209. doi:10.1126/science.1228746
de Ronde D, Butterbach P, Kormelink R (2014) Dominant resistance against plant viruses. Front Plant Sci 5:307. doi:10.3389/fpls.2014.00307
Firth D (1993) Bias reduction of maximum likelihood estimates. Biometrika 80:27–38. doi:10.1093/biomet/80.1.27
Fu D, Uauy C, Distelfeld A, Blechl A, Epstein L, Chen X, Sela H, Fahima T, Dubcovsky J (2009) A kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323:1357–1360. doi:10.1126/science.1166289
Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B, Higgins J, DeFelice M, Lochner A, Faggart M, Liu-Cordero SN, Rotimi C, Adeyemo A, Cooper R, Ward R, Lander ES, Daly MJ, Altshuler D (2002) The structure of haplotype blocks in the human genome. Science 296:2225–2229. doi:10.1126/science.1069424
Gagarinova AG, Babu M, Poysa V, Hill JH, Wang A (2008) Identification and molecular characterization of two naturally occurring soybean mosaic virus isolates that are closely related but differ in their ability to overcome Rsv4 resistance. Virus Res 138:50–56. doi:10.1016/j.virusres.2008.08.010
Gascuel O (1997) BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol Biol Evol 14:685–695
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186. doi:10.1093/nar/gkr944
Gore MA, Hayes AJ, Jeong S-C, Yue YG, Buss GR, Saghai Maroof MA (2002) Mapping tightly linked genes controlling potyvirus infection at the Rsv1 and Rpv1 region in soybean. Genome 45:592–599. doi:10.1139/g02-009
Grant D, Nelson RT, Cannon SB, Shoemaker RC (2010) SoyBase, the USDA-ARS soybean genetics and genomics database. Nucl Acids Res 38(suppl 1):D843–D846. doi:10.1093/nar/gkp798
Gunduz I, Buss GR, Chen P, Tolin SA (2002) Characterization of SMV resistance genes in Tousan 140 and Hourei soybean. Crop Sci 42:90–95
Gunduz I, Buss GR, Chen P, Tolin SA (2004) Genetic and phenotypic analysis of soybean mosaic virus resistance in PI 88788 soybean. Phytopathology 94:687–692. doi:10.1094/PHYTO.2004.94.7.687
Guo D, Spetz C, Saarma M, Valkonen JPT (2003) Two potato proteins, including a novel RING Finger Protein (HIP1), interact with the potyviral multifunctional protein HCpro. Mol Plant Microbe Interact 16:405–410. doi:10.1094/MPMI.2003.16.5.405
Hayes AJ, Ma G, Buss GR, Saghai Maroof MA (2000) Molecular marker mapping of Rsv4, a gene conferring resistance to all known strains of soybean mosaic virus. Crop Sci 40:1434. doi:10.2135/cropsci2000.4051434x
Hayes AJ, Jeong S-C, Gore MA, Yu YG, Buss GR, Tolin SA, Saghai Maroof MA (2004) Recombination within a nucleotide-binding-site/leucine-rich-repeat gene cluster produces new variants conditioning resistance to soybean mosaic virus in soybeans. Genetics 166:493–503. doi:10.1534/genetics.166.1.493
Heinze G, Ploner M, Beyea J (2013) Confidence intervals after multiple imputation: combining profile likelihood information from logistic regressions. Stat Med 32:5062–5076. doi:10.1002/sim.5899
Hill JH, Bailey TB, Benner HI, Tachibana H, Durand DP (1987) Soybean mosaic virus: effects of primary disease incidence on yield and seed quality. Plant Dis 71:237. doi:10.1094/PD-71-0237
Hofius D, Maier AT, Dietrich C, Jungkunz I, Börnke F, Maiss E, Sonnewald U (2007) Capsid protein-mediated Recruitment of host DnaJ-like proteins Is required for Potato Virus Y infection in tobacco plants. J Virol 81:11870–11880. doi:10.1128/JVI.01525-07
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267. doi:10.1093/molbev/msj030
Hwang T-Y, Moon J-K, Yu S, Yang K, Mohankumar S, Yu YH, Lee YH, Kim HS, Kim HM, Saghai Maroof MA, Jeong S-C (2006) Application of comparative genomics in developing molecular markers tightly linked to the virus resistance gene Rsv4 in soybean. Genome 49:380–388. doi:10.1139/g05-111
Innes RW, Ameline-Torregrosa C, Ashfield T, Cannon E, Cannon SB, Chacko B, Chen NWG, Couloux A, Dalwani A, Denny R, Deshpande S, Egan AN, Glover N, Hans CS, Howell S, Ilut DC, Jackson S, Lai H, Mammadov J, del Campo SM, Metcalf M, Nguyen A, O’Bleness M, Pfeil BE, Podicheti R, Ratnaparkhe MB, Samain S, Sanders I, Ségurens B, Sévignac M, Sherman-Broyles S, Thareau V, Tucker DM, Walling J, Wawrzynski A, Yi J, Doyle JJ, Geffroy V, Roe BA, Saghai Maroof MA, Young ND (2008) Differential accumulation of retroelements and diversification of NB-LRR disease resistance genes in duplicated regions following polyploidy in the ancestor of soybean. Plant Physiol 148:1740–1759. doi:10.1104/pp.108.127902
Jeong N, Jeong S-C (2014) Multiple genes confer resistance to soybean mosaic virus in the soybean cultivar Hwangkeum. Plant Genet Resour 12:S41–S44. doi:10.1017/S1479262114000227
Jeong S-C, Kristipati S, Hayes AJ, Maughan PJ, Noffsinger SL, Gunduz I, Buss GR, Saghai Maroof MA (2002) Genetic and sequence analysis of markers tightly linked to the resistance gene, Rsv3. Crop Sci 42:265–270. doi:10.2135/cropsci2002.2650
Jeong N, Moon J-K, Kim HS, Kim C-G, Jeong S-C (2010) Fine genetic mapping of the genomic region controlling leaflet shape and number of seeds per pod in the soybean. Theor Appl Genet 122:865–874. doi:10.1007/s00122-010-1492-5
Jørgensen IH (1992) Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 63:141–152. doi:10.1007/BF00023919
Kanaya S (2009) Ribonuclease H. FEBS J 276:1481. doi:10.1111/j.1742-4658.2009.06906.x
Kiihl RAS, Hartwig EE (1979) Inheritance of reaction to soybean mosaic virus in soybeans. Crop Sci 19:372. doi:10.2135/cropsci1979.0011183X001900030024x
Kim SD, Hong EH, Seong YK, Lee SH, Kim HS, Ryu YH, Kim YS (1996) A new soybean variety for sprouting ‘Pureunkong’ with green seed coat and cotyledon, good seed quality. RDA J Agric Sci 38:238–241
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363. doi:10.1126/science.1166453
Krattinger SG, Jordan DR, Mace ES, Raghavan C, Luo M-C, Keller B, Lagudah ES (2012) Recent emergence of the wheat Lr34 multi-pathogen resistance: insights from haplotype analysis in wheat, rice, sorghum and Aegilops tauschii. Theor Appl Genet 126:663–672. doi:10.1007/s00122-012-2009-1
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181. doi:10.1016/0888-7543(87)90010-3
Layer RM, Chiang C, Quinlan AR, Hall IM (2014) LUMPY: a probabilistic framework for structural variant discovery. Genome Biol 15:R84. doi:10.1186/gb-2014-15-6-r84
Lee WK, Kim N, Kim J, Moon J-K, Jeong N, Choi I-Y, Kim SC, Chung W-H, Kim HS, Lee S-H, Jeong S-C (2013) Dynamic genetic features of chromosomes revealed by comparison of soybean genetic and sequence-based physical maps. Theor Appl Genet 126:1103–1119. doi:10.1007/s00122-012-2039-8
Lee TG, Kumar I, Diers BW, Hudson ME (2015) Evolution and selection of Rhg1, a copy-number variant nematode-resistance locus. Mol Ecol 24:1774–1791. doi:10.1111/mec.13138
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol. doi:10.1111/2041-210X.12410
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26:589–595. doi:10.1093/bioinformatics/btp698
Li D, Chen P, Alloatti J, Shi A, Chen YF (2010) Identification of new alleles for resistance to soybean mosaic virus in soybean. Crop Sci 50:649. doi:10.2135/cropsci2009.06.0302
Libault M, Farmer A, Joshi T, Takahashi K, Langley RJ, Franklin LD, He J, Xu D, May G, Stacey G (2010) An integrated transcriptome atlas of the crop model Glycine max, and its use in comparative analyses in plants. Plant J 63:86–99. doi:10.1111/j.1365-313X.2010.04222.x
Lipka AE (2009) Associating single nucleotide polymorphisms (SNPs) with binary traits. Dissertation, Purdue University
Ma G, Chen P, Buss GR, Tolin SA (1995) Genetic characteristics of two genes for resistance to soybean mosaic virus in PI486355 soybean. Theor Appl Genet 91:907–914. doi:10.1007/BF00223899
Malik HS (2005) Ribonuclease H evolution in retrotransposable elements. Cytogenet Genome Res 110:392–401. doi:10.1159/000084971
Marchler-Bauer A, Bryant SH (2004) CD-Search: protein domain annotations on the fly. Nucleic Acids Res 32:W327–W331. doi:10.1093/nar/gkh454
Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Bryant SH (2015) CDD: NCBI’s conserved domain database. Nucleic Acids Res 43:D222–D226. doi:10.1093/nar/gku1221
McHale L, Tan X, Koehl P, Michelmore RW (2006) Plant NBS-LRR proteins: adaptable guards. Genome Biol 7:212. doi:10.1186/gb-2006-7-4-212
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. doi:10.1101/gr.107524.110
Michelmore RW, Christopoulou M, Caldwell KS (2013) Impacts of resistance gene genetics, function, and evolution on a durable future. Annu Rev Phytopathol 51:291–319. doi:10.1146/annurev-phyto-082712-102334
Moon J-K, Jeong S-C, Van K, Saghai Maroof MA, Lee S-H (2009) Marker-assisted identification of resistance genes to soybean mosaic virus in soybean lines. Euphytica 169:375–385. doi:10.1007/s10681-009-9970-z
Rausch T, Zichner T, Schlattl A, Stütz AM, Benes V, Korbel JO (2012) DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28:i333–i339. doi:10.1093/bioinformatics/bts378
R Core Team (2015) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Saghai Maroof MA, Jeong S-C, Gunduz I, Tucker DM, Buss GR, Tolin SA (2008) Pyramiding of soybean mosaic virus resistance genes by marker-assisted selection. Crop Sci 48:517. doi:10.2135/cropsci2007.08.0479
Saghai Maroof MA, Tucker DM, Skoneczka JA, Bowman BC, Tripathy S, Tolin SA (2010) Fine mapping and candidate gene discovery of the soybean mosaic virus resistance gene, Rsv4. Plant Genome 3:14. doi:10.3835/plantgenome2009.07.0020
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang X-C, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183. doi:10.1038/nature08670
Severin AJ, Woody JL, Bolon Y-T, Joseph B, Diers BW, Farmer AD, Muehlbauer GJ, Nelson RT, Grant D, Specht JE, Graham MA, Cannon SB, May GD, Vance CP, Shoemaker RC (2010) RNA-Seq atlas of Glycine max: a guide to the soybean transcriptome. BMC Plant Biol 10:160. doi:10.1186/1471-2229-10-160
Song QJ, Marek LF, Shoemaker RC, Lark KG, Concibido VC, Delannay X, Specht JE, Cregan PB (2004) A new integrated genetic linkage map of the soybean. Theor Appl Genet 109:122–128. doi:10.1007/s00122-004-1602-3
Suh SJ, Bowman BC, Jeong N, Yang K, Kastl C, Tolin SA, Saghai Maroof MA, Jeong S-C (2011) The Rsv3 locus conferring resistance to soybean mosaic virus is associated with a cluster of coiled-coil nucleotide-binding leucine-rich repeat genes. Plant Genome J 4:55. doi:10.3835/plantgenome2010.11.0024
Tucker DM, Saghai Maroof MA, Jeong S-C, Buss GR, Tolin SA (2009) Validation and interaction of the soybean mosaic virus lethal necrosis allele, Rsv1-n, in PI 507389. Crop Sci 49:1277. doi:10.2135/cropsci2008.08.0477
Untergasser A, Cutcutache I, Koressaar T, Ye J, Faircloth BC, Remm M, Rozen SG (2012) Primer3—new capabilities and interfaces. Nucleic Acids Res 40:e115. doi:10.1093/nar/gks596
Wang D, Ma Y, Yang Y, Liu N, Li C, Song Y, Zhi H (2010) Fine mapping and analyses of RSC8 resistance candidate genes to soybean mosaic virus in soybean. Theor Appl Genet 122:555–565. doi:10.1007/s00122-010-1469-4
Wen R-H, Khatabi B, Ashfield T, Saghai Maroof MA, Hajimorad MR (2012) The HC-Pro and P3 cistrons of an avirulent soybean mosaic virus are recognized by different resistance genes at the complex Rsv1 locus. Mol Plant Microbe Interact 26:203–215. doi:10.1094/MPMI-06-12-0156-R
Whitham SA, Anderberg RJ, Chisholm ST, Carrington JC (2000) Arabidopsis RTM2 gene is necessary for specific restriction of tobacco etch virus and encodes an unusual small heat shock-like protein. Plant Cell 12:569–582
Wrather JA, Anderson TR, Arsyad DM, Tan Y, Ploper LD, Porta-Puglia A, Ram HH, Yorinori JT (2001a) Soybean disease loss estimates for the top ten soybean-producing countries in 1998. Can J Plant Pathol 23:115–121. doi:10.1080/07060660109506918
Wrather JA, Stienstra WC, Koenning SR (2001b) Soybean disease loss estimates for the United States from 1996 to 1998. Can J Plant Pathol 23:122–131. doi:10.1080/07060660109506919
Yu Y, Saghai Maroof MA, Buss GR, Maughan PJ, Tolin SA (1994) RFLP and microsatellite mapping of a gene for soybean mosaic virus resistance. Phytopathology 84:60–64
Acknowledgments
This study in the SCJ lab was supported principally by a grant from the Next-Generation BioGreen 21 Program (PJ01101503), by the Rural Development Administration, and partly supported by the National Research Foundation of Korea grant funded by the Korea government (MSIP) (No. 2015R1A2A2A01002969) and by the Korea Research Institute of Bioscience and Biotechnology Research Initiative Program. MASM acknowledges funding from the Virginia Soybean Board, the Virginia Agricultural Experiment Station and the Hatch Program of the National Institute of Food and Agriculture, US Department of Agriculture. This work was also partly funded by USDA-NIFA/DOE Biomass Research and Development Initiative (BRDI) Grant No. 2012-10006 (MAG), Cornell University startup funds (MAG), and University of Illinois startup funds (AEL).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by H. T. Nguyen.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ilut, D.C., Lipka, A.E., Jeong, N. et al. Identification of haplotypes at the Rsv4 genomic region in soybean associated with durable resistance to soybean mosaic virus. Theor Appl Genet 129, 453–468 (2016). https://doi.org/10.1007/s00122-015-2640-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-015-2640-8