Abstract
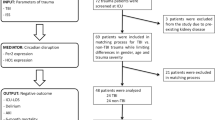
Disturbances in the circadian rhythm have been reported in patients following traumatic brain injury (TBI). However, the rhythmic expression of circadian genes in peripheral blood leukocytes (PBL) following TBI has not yet been studied. The messenger ribonucleic acid (mRNA) expression of period 1 (Per1), Per2, Per3, cryptochrome 1 (Cry1), Cry2, brain and muscle aryl hydrocarbon receptor nuclear translocator-like 1 (Bmal1), and circadian locomotor output cycles kaput (Clock) was quantified in PBLs from sham-operated rats and rats with acute subdural hematoma (ASDH) over a 48-h period. The rectal temperature of the animals was measured every 4 h over 2 days. The mesor, rhythm, amplitude, and acrophase were estimated using cosinor analysis. Cosinor analysis revealed that Per2, Cry1, and Bmal1 mRNAs were rhythmically expressed in the PBLs of sham-operated rats. In contrast, fluctuations in rhythmic expression were not observed following ASDH. The rectal temperature of sham-operated rats also exhibited rhythmicity. ASDH rats had a disrupted rectal temperature rhythm, a diminished amplitude, and an acrophase shift. TBI with ASDH results in dysregulated expression of some circadian genes and changes in body temperature rhythm. Further research is required to understand the pathophysiology of altered circadian networks following TBI.
Key messages
-
First to investigate the mRNA expression of circadian genes in PBLs of ASDH rats.
-
ASDH rats had disrupted rhythmicity of Per2, Cry1, and Bmal1 mRNA expression.
-
Cosinor analysis showed that ASDH rats had a disrupted rectal temperature rhythm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acute subdural hematoma (ASDH) is a common consequence of head injury in humans, characterized by the accumulation of blood between the dura and arachnoid membrane. Although there have been many relevant advances in emergency services, including multimodality neuromonitoring and neurointensive care, ASDH remains associated with high morbidity and mortality. Even if patients survive without major neurological deficits, they may encounter sleep–wake disturbances, particularly fatigue, hypersomnia, and insomnia [1,2,3].
Circadian rhythms are cycles lasting approximately 24 h and entail various physiological and molecular changes. Rhythms are endogenously generated in response to light, darkness, and other environmental cues [4, 5]. Circadian fluctuations play critical roles in the regulation of biological processes within the body, including the sleep–wake cycle, body temperature, eating habits, cardiovascular function, hormonal rhythms, and metabolism [6, 7]. The regulation of biological rhythms aids organisms to anticipate and adapt to environmental changes. The timings of mammalian circadian rhythms are classically inferred based on core body temperature (cBT) changes, melatonin secretion by the pineal gland, and blood cortisol levels. The primary circadian clock is in the suprachiasmatic nucleus (SCN), a bilateral structure located in the hypothalamus.
A subset of neurons within the SCN is the basis for circadian clock resetting through light entrainment and melatonin secretion. The neurons are sensitive to light signals transduced from the retina via the retinohypothalamic tract. External cues, such as the light–dark cycle, environmental temperature, and timing of food intake, can entrain these internal biological rhythms [7]. The circadian fluctuations are regulated through rhythmic gene expression within a complex neural regulatory network involving transcription and translation feedback loops, which ultimately cause oscillation [8]. In turn, output from the SCN controls the circadian rhythm throughout the body via the regulation of circadian gene expression in peripheral tissues and autonomic nervous system activity [8, 9]. Autonomous clocks exist in all peripheral tissues and are driven and synchronized by the SCN via circadian output pathways [10]. Circadian gene expression within the SCN governs the rhythms of various cellular metabolic processes, neuronal firing, and neuropeptide secretion, which ultimately manifest in physiological and behavioral rhythms [9, 11, 12]. A previous study demonstrated that 40.7% of patients with moderate-to-severe traumatic brain injury (TBI) exhibit disturbed circadian rhythms of brain temperature with a diminished amplitude in the first 72 h following operation [13]. However, the effects of TBI on the regulation of circadian rhythm-related gene expression in peripheral tissues have not yet been investigated in humans or animal models.
In this study, we analyzed the messenger ribonucleic acid (mRNA) expression dynamics of seven circadian rhythm-related genes in peripheral blood leukocytes (PBLs) from sham-operated and ASDH rats. The rhythmicity of cBT of sham-operated and ASDH rats was also analyzed. Finally, cosinor analysis was used to analyze the rhythmicity, mesor, amplitude, and acrophase of mRNA expression and cBT.
Materials and methods
Ethics statement
This study complied with the ARRIVE guidelines and was carried out in accordance with the U.K. Animals (Scientific Procedures) Act, 1986 and associated guidelines, EU Directive 2010/63/EU for animal experiments, or the National Research Council’s Guide for the Care and Use of Laboratory Animals. The study was approved by the Institutional Animal Care and Use Committee of National Taiwan University, College of Medicine (approval no. 20120485; date of approval: April 20, 2012).
General preparation
Fourteen 8-week-old male Sprague–Dawley rats, weighing 190–210 g, were used in the study. Rats were allowed to acclimate to the animal room lighting conditions for 2 weeks prior to surgical procedures. The room temperature and relative humidity were maintained at 22 ± 2 °C and 50% ± 20%, respectively. The lighting conditions in the animal room were lights on from 07:00 to 19:00, with a 12:12 h cycle. The animals received water and food ad libitum prior to the experiments. Animals were randomized to the sham-operated group (n = 7) and ASDH group (n = 7). Data were analyzed between these groups.
Induction of subdural hematoma
Following general preparation, animals in the ASDH group (n = 7) were placed in a prone position. Rats were then anesthetized with 2.5% isoflurane. Thereafter, an 8-mm sagittal scalp incision was made. A 3-mm frontal burr hole was drilled into the right frontal region, 3 mm from the sagittal suture and 1 mm anterior of the coronal suture, using surgical loupes. The dura was incised with a 26-gauge needle and a Codman microsensor (Codman, Raynham, MA, USA), connected to the Codman intracranial pressure (ICP) monitor, was inserted into the subdural space together with a 26-gauge L-shaped needle. The hole was secured with bone wax. This model is a modified version of the established models [14,15,16]. Nonheparinized venous blood was obtained from the tail vein of rats, and 0.06–0.1 mL was slowly injected (> 1 min) into the subdural space through the needle via the burr hole until the ICP was 22–25 mmHg. Next, the ICP catheter was removed, and subsequently, the dural opening was sealed with gelfoam, whereas the burr hole was sealed with bone wax. The scalp incision was closed using nylon sutures. Rats were returned to housing conditions under a 12-h light–dark cycle, and ad libitum oral intake was resumed postoperatively. Considering the effects of the day-night cycle on the study of circadian rhythm, all surgical procedures were completed by 5 AM. Blood sampling for mRNA expression analysis and cBT measurements was performed from 9 AM on the same day. Blood for analysis was collected from the tail veins every 4 h for 48 h. To analyze the effects of ASDH on cBT regulation, the rectal temperature of the animals was measured at 4-h intervals using a temperature probe. In sham-operated animals, no injection into the subdural space was made after placement of the ICP catheter and needle.
At 48 h, rats were euthanized via intraperitoneal injection of a lethal dose of pentobarbital. None of the animals exhibited obvious hemiparesis or other focal neurological deficits. No seizures were observed. Considering that ICP-induced brain injury with edema may exacerbate in the first few days after ASDH, a two-day survival time was chosen for the present experiments.
Sample collection and mRNA extraction
Local anesthetic cream was applied on the surface of the tail 30 min before the experiment to minimize the effects of anesthesia on circadian gene expression. Peripheral blood samples of 100 µl each were collected from the tail veins of rats at 4 h intervals over a span of 48 h following the surgical procedure. These samples were collected to analyze the 24-h gene expression rhythms. For overnight data collection time points, tail vein blood sampling was conducted in a darkened room, with the only illumination being provided by red light. mRNA transcript measurements at 12 time points provided sufficient data for the estimation of cosinor parameters [17]. Total ribonucleic acid (RNA) was extracted from samples using the QIAamp RNA Blood Mini Kit with on-column deoxyribonuclease (DNase) treatment of RNA samples (Qiagen, Valencia, CA, USA) under strict RNAse-free conditions.
cDNA synthesis and quantitative PCR
Up to 1 µg RNA was reverse-transcribed into complementary DNA (cDNA) using the Omniscript Reverse Transcription Kit (Qiagen) and 10 µM oligo-dT primers (Applied Biosystems, Waltham, MA, USA) in accordance with the manufacturer’s instructions. The 20-µL reaction volume containing the completed first-strand cDNA synthesis reaction was diluted to 50 µL, and 1 µL of this dilution was used for each quantitative polymerase chain reaction (PCR). PCRs were performed on an Illumina Eco™ Real-Time PCR System (Illumina, San Diego, CA, USA) with the following reaction conditions: initial denaturation at 95 °C for 10 min, 40 cycles with 10 s denaturation at 94 °C, and 30 s annealing at 60 °C. mRNA expression measurements were performed in triplicate, and the average was calculated. Relative expression levels for the means of the triplicate experiments for circadian genes were normalized to those of β-actin as an internal control, and the relative threshold cycle (∆Ct) was obtained. The 2−ΔΔCt method was used to analyze mRNA expression [18, 19]. The circadian genes analyzed were period 1 (Per1), Per2, Per3, cryptochrome 1 (Cry1), Cry2, circadian locomotor output cycles kaput (Clock), and brain and muscle aryl hydrocarbon receptor nuclear translocator-like 1 (Bmal1), and the primers employed are listed in Table 1 (Per3, Cry2 [20]; Per1, Per2, Cry1, Clock [21]; Bmal1 [22]). The primers for β-actin were designed using Primer3 (version 4.0; https://bioinfo.ut.ee/primer3/) and Primer-BLAST (Basic Local Alignment Search Tool, NCBI). The mRNA expression of clock genes was quantified using β-actin as the reference gene, which has been used as a housekeeping gene for circadian rhythm studies in several tissues [23,24,25]. The circadian oscillations of mRNA expression, mesor (mean level of mRNA oscillations), amplitude of the rhythm (used to measure half of the difference between the lowest and highest levels of mRNA expression), and acrophase (the time when mRNA expression reaches its peak during the day), were determined.
cBT analysis
The rectal temperature of the animals was measured using an anal probe every 4 h over 2 days, beginning from 9 AM on the day of operation. This time point was 4 h after the operation to ensure the stabilization of body temperature after general anesthesia and potential postoperative hypothermia. Temperature measurements at eight-time points provided sufficient temperature records for a better estimation of the cosinor parameters [17]. The following characteristics of rectal temperature were analyzed: circadian oscillations of temperature, mesor, amplitude of the temperature rhythm, and acrophase.
Cosinor analysis
The circadian rhythm of mRNA expression and rectal temperature was analyzed using the cosinor method, which is the most commonly used approach for analyzing the diurnal pattern of biological rhythms [17, 26, 27]. Cosinor analysis is a nonlinear model that fits the data to a 24-h cosine curve, with estimates of rhythm, including mesor, amplitude, and acrophase [27]. An online platform (https://cosinor.online/app/cosinor.php) was used in this study [28]. If the characteristics of the data analyzed fit the cosine curve with P < 0.05, a circadian rhythm was confirmed.
Results
mRNA expression of circadian genes
To determine the effects of ASDH on circadian rhythm regulation, we measured the mRNA expression of seven circadian rhythm-related genes in PBLs. The mRNA expression patterns of Per1, Per2, Per3, Cry1, Cry2, Clock, and Bmal1 in PBLs of sham-operated rats (n = 7) and rats with ASDH (n = 7) were determined using real-time PCR (Fig. 1).
Time course-dependent changes in the relative messenger ribonucleic acid (mRNA) levels of a Per1, b Per2, c Per3, d Cry1, e Cry2, f Bmal1, and g Clock in peripheral blood leukocytes (PBL) from sham-operated (n = 7, yellow line) and acute subdural hematoma (ASDH) (n = 7, blue line) rats. mRNA levels are expressed relative to those of β-actin. Using the cosinor method analysis, the mRNA expression of Per2, Cry1, and Bmal1 exhibited significant 24-h variation in sham-operated rats (P < 0.05). The standard error is not shown to ensure clarity of presentation
The average mRNA expression at each time point was assessed using cosinor analysis. Per2, Cry1, and Bmal1 mRNAs exhibited circadian expression rhythmicity in the PBLs of sham-operated rats (P < 0.05; Fig. 2). The acrophase of Per2, Cry1, and Bmal1 mRNA expression in sham-operated rats was 4:26, 7:48, and 8:00, respectively (Table 2). No rhythmic expression was observed for Per1, Per3, Cry2, and Clock mRNAs in the PBLs of sham-operated rats. In ASDH rats, the rhythmic changes in the expression of Per2, Cry1, and Bmal1 mRNA were disrupted. No rhythmicity was observed for Per1, Per3, Cry2, and Clock mRNAs in the PBLs of the ASDH rats.
Time course-dependent changes in relative mRNA levels of a Per2, b Cry1, and c Bmal1 in the PBLs of sham-operated rats. mRNA levels are expressed relative to those of β-actin. The estimated best-fit cosine curve (continuous line) is plotted by analyzing the mean expression in seven samples at each time point. For example, the two levels of expression at hour 1 represent the data obtained at the 1st and 25th h. The mRNA expression of Per2, Cry1, and Bmal1 exhibited significant 24-h variations (P < 0.05)
cBT analysis
To determine the effects of ASDH on the regulation of core body temperature, rectal temperature was measured using an anal probe every 4 h over 2 days (Fig. 3). The average rectal temperature at each time point was analyzed using cosinor analysis (Table 3). In the sham-operated rats, the temperature changes fit the 24-h day-night rhythm (P = 0.04), with the highest values occurring at 00:16 AM (Fig. 4). In the ASDH rats, the rhythmicity was disrupted (P = 0.06), with decreased amplitude and an acrophase shift to 05:42 (Fig. 5). The mesor was 37.35 °C in the sham-operated group and 37.37 °C in the ASDH group.
Time-related patterns of rectal temperature in sham-operated rats. X–Y plots represent the fitted cosine curves (continuous line) of rectal temperature measurements at 4-h intervals over 48 h. The curve is plotted by analyzing the measurements obtained for seven rats at each time point. For example, the two measurements at hour 1 represent the data obtained at the 1st and 25th h
Time-related patterns of rectal temperature in ASDH rats. X–Y plots representing the fitted cosine curves (continuous line) of rectal temperature measurements at 4-h intervals over 48 h. The curve is plotted by analyzing the measurements for seven rats at each time point. For example, the two measurements at hour 1 represent the data obtained at the 1st and 25th h
Discussion
This study is the first to determine the mRNA expression of circadian genes in PBLs in an ASDH animal model. We observed disrupted rhythmicity of Per2, Cry1, and Bmal1 mRNA and rectal temperature after ASDH. Additionally, a depressed amplitude and acrophase shift were noted.
Oscillations in Per1 and Per2 mRNA expression were initially observed in cultured fibroblasts, suggesting the existence of peripheral circadian clocks [29, 30]. Per genes are involved in cell cycle and cancer development [31, 32]. SCN oscillations have since been detected in various cells outside the SCN as fluctuations in Per1, Per2, Per3, Cry1, Cry2, Bmal1, and Clock expression. The phase delay between SCN and peripheral expression is approximately 4–6 h [33]. Disrupted circadian function results in compromised adaptation to environmental changes, which is associated with various conditions in humans, including aging, neurological and psychiatric problems, metabolic disorders, reproductive abnormalities, and cancer development [8, 34]. SCN lesions cause dysregulation of peripheral oscillator rhythms [35] and altered gene expression in certain peripheral tissues [36]. An intrinsic transcriptional–translational feedback loop regulates rhythmic circadian gene expression [37]. The transcription factors, Clock and Bmal1, activate Per (Per1, Per2, and Per3) and Cry (Cry1 and Cry2) genes by heterodimerizing in the nucleus and binding to response elements within respective promoter sequences. Furthermore, PER and CRY repressor proteins form cytoplasmic complexes, which are translocated to the nucleus to inhibit CLOCK/BMAL-mediated transcription [32, 38]. For a new cycle to begin, CRY proteins are targeted for proteasomal degradation through association with FBXL3 (a member of the F-box protein family) E3 ubiquitin ligase complexes [39,40,41]. This transcriptional–translational feedback loop imposes activating/repressive functions in an autoregulatory cyclic manner. In another circadian feedback loop, REV-ERB proteins, the members of the nuclear receptor superfamily of intracellular transcription factors, are activated by the CLOCK/BMAL dimer, which enters the nucleus to inhibit Bmal1. Therefore, Per1 and Per2 expression in rodent SCN peaks during the day, whereas Bmal1 expression peaks at night [42,43,44]. These two feedback loops regulate circadian oscillations in gene expression.
Besides SCN, rhythmic circadian gene expression has been identified in the pineal gland, olfactory bulb, and forebrain [45,46,47,48,49,50,51]. Rhythmic circadian gene expression was detected in the lung, liver, stomach, adrenal glands, kidney, bone marrow, vasculature, adipose tissue, and peripheral blood in animals [42,43,44, 52,53,54,55,56,57,58]. Rhythmic Per1, Cry1, and Bmal1 mRNA expression in human oral mucosa and Per1 and Bmal1 in human skin is known [33]. Rhythmic changes in Per1, Per2, Per3, Cry1, Bmal1, and Clock expression occur in the PBLs and whole blood cells in healthy human subjects [59,60,61,62,63,64]. These findings suggest specific expression patterns of circadian genes in various tissues and organs. However, previous studies have yielded controversial data regarding circadian expression rhythmicity and acrophase. Moreover, consistent conclusions regarding whether all circadian genes exhibit rhythmicity in peripheral blood cells are missing. Abnormal expression levels of Per2, Clock, and Bmal1 in oral mucosa and mononuclear cells at certain time points were detected in patients with sleep disorders after TBI [65]. Protein levels of some circadian genes have been quantified in peripheral tissues using western blot analysis to clarify their roles in various physiological and pathological conditions [66,67,68,69], but not in TBI models. This highlights the need for further research on circadian gene expression in numerous healthy individuals using standardized protocols and more frequent sample collections.
As circadian clocks are highly conserved across mammals, rodents represent valuable models for investigating the regulation of human circadian rhythms in many diseases [70, 71]. Similar mRNA expression rhythms were observed in human hepatoma cells and mouse liver, but differences in acrophase and amplitude existed [72]. Serial biopsies of human bone marrow and adipose tissue are performed for the time-series analysis of gene expression [53, 73], but it is difficult to perform for other human organs/tissues. Therefore, we adopted the ASDH rat model to explore the molecular mechanisms underlying ASDH-associated circadian clock dysregulation. PBLs were selected as source material to analyze circadian rhythm, considering the accessibility and animal welfare issues. We found that only Per2, Cry1, and Bmal1 mRNA expression was rhythmic in sham-operated rat PBLs, but it was disrupted in ASDH rats, suggesting that the accumulation of blood in the subdural space with increased ICP is responsible for this change.
ASDH also disrupted the rhythm of rectal temperature, with reduced amplitude and a shift of the acrophase. The peak cBT in sham-operated rats was at 00:16 AM, which is comparable to the findings of a previous study [74]. In a study comprising 108 patients with moderate-to-severe TBI, 40.7% presented disrupted brain temperature rhythm; however, some patients with normal brain temperature rhythm still exhibited phase shifts [13]. Only a few studies have investigated the rhythms and acrophases of body or brain temperature in patients after brain injuries. Shifts in acrophase of cBT were observed in a study comprising 28 Alzheimer’s disease patients who presented with high nocturnal activity and fragmented sleep [75]. Among 100 patients with intracerebral hemorrhage, the rhythmicity of systolic blood pressure and heart rate was lost in 43% and 52% of patients, respectively [76]. In a study of 78 patients with basal ganglia hemorrhage after surgery, brain temperature remained intact in 55.1%, with acrophase shift in 60.3% of patients [77]. Furthermore, cosinor analysis of cBT in 86 patients with ruptured cerebral aneurysms revealed elevated mesors (37.8 ± 0.4 °C) with blunted amplitudes (0.27 ± 0.14 °C), and only 27% of acrophases remained within the normative 12–6 PM quadrant [78].
Traumatic brain injury (TBI) can lead to various forms of damage in the case of ASDH, encompassing localized cortical ischemic damage related to the hematoma, secondary injury due to increased intracranial pressure (ICP), and the release of toxic substances from the hematoma itself [79,80,81]. In their study using the high-frequency head impact (HFHI) and controlled cortical impact (CCI) mouse models of TBI, Korthas et al. revealed that different forms of brain trauma can result in distinct patterns of circadian and sleep disruptions [82]. Boone et al. observed that TBI disrupts the oscillation of Per1 and Bmal1 mRNA within the SCN and hippocampus [83]. Additionally, Li et al. found that, in context of TBI in rats, Bmal1 levels in the cerebral cortex decrease, exacerbating nerve damage by increasing the phosphorylation of P38 MAPK (mitogen-activated protein kinase) [84]. The effects of TBI on mRNA expression of peripheral oscillators are under-examined. Patients with TBI of varying severity may experience impaired circadian rhythm after injury to the hypothalamus. Changes in environmental synchronizers, such as ambient temperature in the ICU and feeding schedule, have been documented [85, 86]. Hemorrhage or other injuries of the retinohypothalamic tract may lead to the disruption of retinal inputs to the SCN [87]. Repeated impacts in high-frequency head impact (HFHI) mice have been demonstrated to result in the development of chronic inflammation and damage in the optic nerve/tract. Additionally, inflammation has been documented in the hypothalamus on the same side as the controlled cortical impact (CCI) injury in mice. [82]. In an autopsy study, microhemorrhage or ischemic necrosis in the hypothalamus was found in 42.5% of patients who died after severe TBI [88].
The increased ICP after ASDH in our model underlies SCN dysregulation. We injected nonheparinized blood into the subdural space to achieve an ICP of 22–25 mmHg. A larger blood injection and higher ICP may mimic the clinical scenarios in ASDH patients, wherein ICP impairs consciousness and causes neurological deficits. However, the animals may not survive through the entire study period owing to high ICP. As several factors may contribute to the dysregulation of circadian gene expression and temperature changes, further studies are required to explore other potential mechanisms.
Our study has a few limitations. First, the frequency and duration of longitudinal peripheral blood sampling were limited. Considering the welfare of animals and the possible effects of delayed brain ischemia and edema, we obtained 12 measurements over 2 days. Additional data collection may improve the reliability of cosinor analysis but may disturb the circadian rhythms of animals. Using telemetry devices may allow researchers to monitor the body temperature of animals continuously in their home environments without disturbance [17]. Second, unilateral limb weakness after ASDH makes it difficult to observe locomotor activity in animals. Third, our study exclusively employed male rats, which may limit the generalizability of the results to female rodents. Although the question of whether male or female rats exhibit higher variability in circadian rhythm study remains inconclusive, it is important to consider the effects of androgens and estrous cycle phase on the endogenous circadian period in male and female rodents, respectively. Fourth, the model using subdural blood injection does not entirely replicate the actual clinical scenarios of SDH, which are frequently associated with extensive primary brain injury. Further research is needed to investigate gene expression oscillations in SCN and different peripheral tissues and to explore the precise mechanisms underlying dysregulation of circadian gene expression after TBI, along with its relationship with body temperature changes.
In conclusion, this study provides novel insight into the rhythmicity of cBT and mRNA expression of circadian genes in PBLs in a rat model of ASDH. ASDH resulted in dysregulation of Per2, Cry1, and Bmal1 mRNA expression in PBLs and rhythmic changes in body temperature during the first 48-h post-surgery. Although further studies are needed to explore the underlying molecular mechanisms, the dysregulation of mRNA expression may be targeted for the treatment of patients with post-TBI neurological and mental disorders.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Duclos C, Dumont M, Wiseman-Hakes C, Arbour C, Mongrain V, Gaudreault PO, Khoury S, Lavigne G, Desautels A, Gosselin N (2014) Sleep and wake disturbances following traumatic brain injury. Pathol Biol (Paris) 62:252–261. https://doi.org/10.1016/j.patbio.2014.05.014
Morse AM, Kothare S (2018) Sleep disorders and concussion. Handb Clin Neurol 158:127–134. https://doi.org/10.1016/B978-0-444-63954-7.00013-6
Wolfe LF, Sahni AS, Attarian H (2018) Sleep disorders in traumatic brain injury. NeuroRehabilitation 43:257–266. https://doi.org/10.3233/NRE-182583
El Cheikh HL, Mollard P, Bonnefont X (2019) Molecular and cellular networks in the suprachiasmatic nuclei. Int J Mol Sci 20:2052. https://doi.org/10.3390/ijms20082052
Richards J, Gumz ML (2013) Mechanism of the circadian clock in physiology. Am J Physiol Regul Integr Comp Physiol 304:R1053–R1064. https://doi.org/10.1152/ajpregu.00066.2013
Green CB, Takahashi JS, Bass J (2008) The meter of metabolism. Cell 134:728–742. https://doi.org/10.1016/j.cell.2008.08.022
Paschos GK, FitzGerald GA (2010) Circadian clocks and vascular function. Circ Res 106:833–841. https://doi.org/10.1161/CIRCRESAHA.109.211706
Hastings MH, Reddy AB, Garabette M, King VM, Chahad-Ehlers S, O’Brien J, Maywood ES (2003) Expression of clock gene products in the suprachiasmatic nucleus in relation to circadian behaviour. Novartis Found Symp 253:203–217; discussion 102–209, 218–222, 281–204. https://doi.org/10.1002/0470090839.ch15
Hastings MH, Brancaccio M, Maywood ES (2014) Circadian pacemaking in cells and circuits of the suprachiasmatic nucleus. J Neuroendocrinol 26:2–10. https://doi.org/10.1111/jne.12125
Mohawk JA, Green CB, Takahashi JS (2012) Central and peripheral circadian clocks in mammals. Annu Rev Neurosci 35:445–462. https://doi.org/10.1146/annurev-neuro-060909-153128
Kalsbeek A, Perreau-Lenz S, Buijs RM (2006) A network of (autonomic) clock outputs. Chronobiol Int 23:521–535. https://doi.org/10.1080/07420520600651073
Maywood ES, O’Neill JS, Chesham JE, Hastings MH (2007) Minireview: the circadian clockwork of the suprachiasmatic nuclei–analysis of a cellular oscillator that drives endocrine rhythms. Endocrinology 148:5624–5634. https://doi.org/10.1210/en.2007-0660
Kuo LT, Lu HY, Huang AP (2021) Prognostic value of circadian rhythm of brain temperature in traumatic brain injury. J Pers Med 11:620. https://doi.org/10.3390/jpm11070620
Sasaki M, Dunn L (2001) A model of acute subdural hematoma in the mouse. J Neurotrauma 18:1241–1246. https://doi.org/10.1089/089771501317095278
Rahimi Nedjat M, Wähmann M, Bächli H, Güresir E, Vatter H, Raabe A, Heimann A, Kempski O, Alessandri B (2013) Erythropoietin neuroprotection is enhanced by direct cortical application following subdural blood evacuation in a rat model of acute subdural hematoma. Neurosci 238:125–134. https://doi.org/10.1016/j.neuroscience.2013.01.067.]
Alessandri B, Nishioka T, Heimann A, Bullock RM, Kempski O (2006) Caspase-dependent cell death involved in brain damage after acute subdural hematoma in rats. Brain Res 1111:196–202. https://doi.org/10.1016/j.brainres.2006.06.105
Cornelissen G (2014) Cosinor-based rhythmometry. Theor Biol Med Model 11:16. https://doi.org/10.1186/1742-4682-11-16
Chen YL, Chuang JH, Wang HT, Chen HC, Liu WH, Yang MY (2021) Altered expression of circadian clock genes in patients with atrial fibrillation is associated with atrial high-rate episodes and left atrial remodeling. Diagnostics (Basel) 11:90. https://doi.org/10.3390/diagnostics11010090
Hida A, Kusanagi H, Satoh K, Kato T, Matsumoto Y, Echizenya M, Shimizu T, Higuchi S, Mishima K (2009) Expression profiles of PERIOD1, 2, and 3 in peripheral blood mononuclear cells from older subjects. Life Sci 84:33–37. https://doi.org/10.1016/j.lfs.2008.10.012
Hashimoto A, Uemura R, Sawada A, Nadatani Y, Otani K, Hosomi S, Nagami Y, Tanaka F, Kamata N, Taira K, Yamagami H, Tanigawa T, Watanabe T, Fujiwara Y (2019) Changes in clock genes expression in esophagus in rat reflux esophagitis. Dig Dis Sci 64:2132–2139. https://doi.org/10.1007/s10620-019-05546-1
Sládek M, Jindráková Z, Bendová Z, Sumová A (2007) Postnatal ontogenesis of the circadian clock within the rat liver. Am J Physiol Regul Integr Comp Physiol 292:R1224–R1229. https://doi.org/10.1152/ajpregu.00184.2006
Christiansen SL, Bouzinova EV, Fahrenkrug J, Wiborg O (2016) Altered expression pattern of clock genes in a rat model of depression. Int J Neuropsychopharmacol 19:pyw061. https://doi.org/10.1093/ijnp/pyw061
Ma TJ, Zhang ZW, Lu YL, Zhang YY, Tao DC, Liu YQ, Ma YX (2018) CLOCK and BMAL1 stabilize and activate RHOA to promote F-actin formation in cancer cells. Exp Mol Med 50:1–15. https://doi.org/10.1038/s12276-018-0156-4
Sakamoto A, Terui Y, Uemura T, Igarashi K, Kashiwagi K (2021) Translational regulation of clock genes BMAL1 and REV-ERBalpha by polyamines. Int J Mol Sci 22:137. https://doi.org/10.3390/ijms22031307
Satou R, Shibukawa Y, Kimura M, Sugihara N (2019) Light conditions affect rhythmic expression of aquaporin 5 and anoctamin 1 in rat submandibular glands. Heliyon 5:e02792. https://doi.org/10.1016/j.heliyon.2019.e02792
Cornélissen G, Tamura K, Tarquini B, Germanò G, Fersini C, Rostagno C, Zaslavskaya RM, Carandente O, Carandente F, Halberg F (1994) Differences in some circadian patterns of cardiac arrhythmia, myocardial infarctions and other adverse vascular events. Chronobiol 21:79–88
Halberg F (1969) Chronobiology. Annu Rev Physiol 31:675–725. https://doi.org/10.1146/annurev.ph.31.030169.003331
Molcan L (2019) Time distributed data analysis by Cosinor. Online application. BioRxiv 805960. https://doi.org/10.1101/805960
Balsalobre A, Damiola F, Schibler U (1998) A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 93:929–937. https://doi.org/10.1016/s0092-8674(00)81199-x
Yagita K, Tamanini F, van Der Horst GT, Okamura H (2001) Molecular mechanisms of the biological clock in cultured fibroblasts. Sci 292:278–281. https://doi.org/10.1126/science.1059542
Ishida N (2007) Circadian clock, cancer and lipid metabolism. Neurosci Res 57:483–490. https://doi.org/10.1016/j.neures.2006.12.012
Ko CH, Takahashi JS (2006) Molecular components of the mammalian circadian clock. Hum Mol Genet 15:271–277. https://doi.org/10.1093/hmg/ddl207
Bjarnason GA, Jordan RC, Wood PA, Li Q, Lincoln DW, Sothern RB, Hrushesky WJ, Ben-David Y (2001) Circadian expression of clock genes in human oral mucosa and skin: association with specific cell-cycle phases. Am J Pathol 158:1793–1801. https://doi.org/10.1016/S0002-9440(10)64135-1
Barnard AR, Nolan PM (2008) When clocks go bad: neurobehavioural consequences of disrupted circadian timing. PLoS Genet 4:e1000040. https://doi.org/10.1371/journal.pgen.1000040
Yoo SH, Yamazaki S, Lowrey PL, Shimomura K, Ko CH, Buhr ED, Siepka SM, Hong HK, Oh WJ, Yoo OJ, Menaker M, Takahashi JS (2004) PERIOD2: LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc Natl Acad Sci USA 101:5339–5346. https://doi.org/10.1073/pnas.0308709101
Guo H, Brewer JM, Lehman MN, Bittman EL (2006) Suprachiasmatic regulation of circadian rhythms of gene expression in hamster peripheral organs: effects of transplanting the pacemaker. J Neurosci 26:6406–6412. https://doi.org/10.1523/JNEUROSCI.4676-05.2006
Dunlap JC, Loros JJ, Liu Y, Crosthwaite SK (1999) Eukaryotic circadian systems: cycles in common. Genes Cells 4:1–10. https://doi.org/10.1046/j.1365-2443.1999.00239.x
Sassone-Corsi P (2010) Commentary: the year in circadian rhythms. Mol Endocrinol 24:2081–2087. https://doi.org/10.1210/me.2010-0359
Busino L, Bassermann F, Maiolica A, Lee C, Nolan PM, Godinho SI, Draetta GF, Pagano M (2007) SCFFbxl3 controls the oscillation of the circadian clock by directing the degradation of cryptochrome proteins. Sci 316:900–904. https://doi.org/10.1126/science.1141194
Godinho SI, Maywood ES, Shaw L, Tucci V, Barnard AR, Busino L, Pagano M, Kendall R, Quwailid MM, Romero MR, O’Neill J, Chesham JE, Brooker D, Lalanne Z, Hastings MH, Nolan PM (2007) The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Sci 316:897–900. https://doi.org/10.1126/science.1141138
Yoo SH, Mohawk JA, Siepka SM, Shan Y, Huh SK, Hong HK, Kornblum I, Kumar V, Koike N, Xu M, Nussbaum J, Liu X, Chen Z, Chen ZJ, Green CB, Takahashi JS (2013) Competing E3 ubiquitin ligases govern circadian periodicity by degradation of CRY in nucleus and cytoplasm. Cell 152:1091–1105. https://doi.org/10.1016/j.cell.2013.01.055
Chen YG, Mantalaris A, Bourne P, Keng P, Wu JH (2000) Expression of mPer1 and mPer2, two mammalian clock genes, in murine bone marrow. Biochem Biophys Res Commun 276:724–728. https://doi.org/10.1006/bbrc.2000.3536
Yamamoto T, Nakahata Y, Soma H, Akashi M, Mamine T, Takumi T (2004) Transcriptional oscillation of canonical clock genes in mouse peripheral tissues. BMC Mol Biol 5:18. https://doi.org/10.1186/1471-2199-5-18
Yamazaki S, Numano R, Abe M, Hida A, Takahashi R, Ueda M, Block GD, Sakaki Y, Menaker M, Tei H (2000) Resetting central and peripheral circadian oscillators in transgenic rats. Sci 288:682–685. https://doi.org/10.1126/science.288.5466.682
Abe M, Herzog ED, Yamazaki S, Straume M, Tei H, Sakaki Y, Menaker M, Block GD (2002) Circadian rhythms in isolated brain regions. J Neurosci 22:350–356. https://doi.org/10.1523/JNEUROSCI.22-01-00350.2002
Abraham U, Prior JL, Granados-Fuentes D, Piwnica-Worms DR, Herzog ED (2005) Independent circadian oscillations of Period1 in specific brain areas in vivo and in vitro. J Neurosci 25:8620–8626. https://doi.org/10.1523/JNEUROSCI.2225-05.2005
Amir S, Lamont EW, Robinson B, Stewart J (2004) A circadian rhythm in the expression of PERIOD2 protein reveals a novel SCN-controlled oscillator in the oval nucleus of the bed nucleus of the stria terminalis. J Neurosci 24:781–790. https://doi.org/10.1523/JNEUROSCI.4488-03.2004
Granados-Fuentes D, Prolo LM, Abraham U, Herzog ED (2004) The suprachiasmatic nucleus entrains, but does not sustain, circadian rhythmicity in the olfactory bulb. J Neurosci 24:615–619. https://doi.org/10.1523/JNEUROSCI.4002-03.2004
Guillaumond F, Dardente H, Giguère V, Cermakian N (2005) Differential control of Bmal1 circadian transcription by REV-ERB and ROR nuclear receptors. J Biol Rhythms 20:391–403. https://doi.org/10.1177/0748730405277232
Preitner N, Damiola F, Lopez-Molina L, Zakany J, Duboule D, Albrecht U, Schibler U (2002) The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 110:251–260. https://doi.org/10.1016/s0092-8674(02)00825-5
Sato TK, Panda S, Miraglia LJ, Reyes TM, Rudic RD, McNamara P, Naik KA, FitzGerald GA, Kay SA, Hogenesch JB (2004) A functional genomics strategy reveals Rora as a component of the mammalian circadian clock. Neuron 43:527–537. https://doi.org/10.1016/j.neuron.2004.07.018
Granda TG, Liu XH, Smaaland R, Cermakian N, Filipski E, Sassone-Corsi P, Lévi F (2005) Circadian regulation of cell cycle and apoptosis proteins in mouse bone marrow and tumor. FASEB J 19:304–306. https://doi.org/10.1096/fj.04-2665fje
Loboda A, Kraft WK, Fine B, Joseph J, Nebozhyn M, Zhang C, He Y, Yang X, Wright C, Morris M, Chalikonda I, Ferguson M, Emilsson V, Leonardson A, Lamb J, Dai H, Schadt E, Greenberg HE, Lum PY (2009) Diurnal variation of the human adipose transcriptome and the link to metabolic disease. BMC Med Genomics 2:7. https://doi.org/10.1186/1755-8794-2-7
McNamara P, Seo SB, Rudic RD, Sehgal A, Chakravarti D, FitzGerald GA (2001) Regulation of CLOCK and MOP4 by nuclear hormone receptors in the vasculature: a humoral mechanism to reset a peripheral clock. Cell 105:877–889. https://doi.org/10.1016/s0092-8674(01)00401-9
Nonaka H, Emoto N, Ikeda K, Fukuya H, Rohman MS, Raharjo SB, Yagita K, Okamura H, Yokoyama M (2001) Angiotensin II induces circadian gene expression of clock genes in cultured vascular smooth muscle cells. Circ 104:1746–1748. https://doi.org/10.1161/hc4001.098048
Oishi K, Miyazaki K, Kadota K, Kikuno R, Nagase T, Atsumi G, Ohkura N, Azama T, Mesaki M, Yukimasa S, Kobayashi H, Iitaka C, Umehara T, Horikoshi M, Kudo T, Shimizu Y, Yano M, Monden M, Machida K, Matsuda J, Horie S, Todo T, Ishida N (2003) Genome-wide expression analysis of mouse liver reveals CLOCK-regulated circadian output genes. J Biol Chem 278:41519–41527. https://doi.org/10.1074/jbc.M304564200
Pizarro A, Hayer K, Lahens NF, Hogenesch JB (2013) CircaDB: a database of mammalian circadian gene expression profiles. Nucleic Acids Res 41:D1009–D1013. https://doi.org/10.1093/nar/gks1161
Tokonami N, Mordasini D, Pradervand S, Centeno G, Jouffe C, Maillard M, Bonny O, Gachon F, Gomez RA, Sequeira-Lopez ML, Firsov D (2014) Local renal circadian clocks control fluid-electrolyte homeostasis and BP. J Am Soc Nephrol 25:1430–1439. https://doi.org/10.1681/ASN.2013060641
Boivin DB, James FO, Wu A, Cho-Park PF, Xiong H, Sun ZS (2003) Circadian clock genes oscillate in human peripheral blood mononuclear cells. Blood 102:4143–4145. https://doi.org/10.1182/blood-2003-03-0779
James FO, Cermakian N, Boivin DB (2007) Circadian rhythms of melatonin, cortisol, and clock gene expression during simulated night shift work. Sleep 30:1427–1436. https://doi.org/10.1093/sleep/30.11.1427
Kusanagi H, Mishima K, Satoh K, Echizenya M, Katoh T, Shimizu T (2004) Similar profiles in human period1 gene expression in peripheral mononuclear and polymorphonuclear cells. Neurosci Lett 365:124–127. https://doi.org/10.1016/j.neulet.2004.04.065
Soares VR, Silva Martins C, Martinez EZ, Araujo LD, Roa SLR, Silva LR, Moreira AC, De Castro M (2020) Peripheral clock system circadian abnormalities in Cushing’s disease. Chronobiol Int 37:867–876. https://doi.org/10.1080/07420528.2020.1758126
Takimoto M, Hamada A, Tomoda A, Ohdo S, Ohmura T, Sakato H, Kawatani J, Jodoi T, Nakagawa H, Terazono H, Koyanagi S, Higuchi S, Kimura M, Tukikawa H, Irie S, Saito H, Miike T (2005) Daily expression of clock genes in whole blood cells in healthy subjects and a patient with circadian rhythm sleep disorder. Am J Physiol Regul Integr Comp Physiol 289:R1273–R1279. https://doi.org/10.1152/ajpregu.00126.2005
Teboul M, Barrat-Petit MA, Li XM, Claustrat B, Formento JL, Delaunay F, Lévi F, Milano G (2005) Atypical patterns of circadian clock gene expression in human peripheral blood mononuclear cells. J Mol Med (Berl) 83:693–699. https://doi.org/10.1007/s00109-005-0697-6
Zhanfeng N, Hechun X, Zhijun Z, Hongyu X, Zhou F (2019) Regulation of circadian clock genes on sleep disorders in traumatic brain injury patients. World Neurosurg 130:e475–e486. https://doi.org/10.1016/j.wneu.2019.06.122
Chen X, Yu F, Guo X, Su C, Li SS, Wu B (2021) Clock gene Bmal1 controls diurnal rhythms in expression and activity of intestinal carboxylesterase 1. J Pharm Pharmacol 73:52–59. https://doi.org/10.1093/jpp/rgaa035
Lagishetty V, Parthasarathy PT, Phillips O, Fukumoto J, Cho Y, Fukumoto I, Bao H, Cox R Jr, Galam L, Lockey RF, Kolliputi N (2014) Dysregulation of CLOCK gene expression in hyperoxia-induced lung injury. Am J Physiol Cell Physiol 306:C999–C1007. https://doi.org/10.1152/ajpcell.00064.2013
Li H, Sun NL, Wang J, Liu AJ, Su DF (2007) Circadian expression of clock genes and angiotensin II type 1 receptors in suprachiasmatic nuclei of sinoaortic-denervated rats. Acta Pharmacol Sin 28:484–492. https://doi.org/10.1111/j.1745-7254.2007.00543.x
Taniguchi H, Fernández AF, Setién F, Ropero S, Ballestar E, Villanueva A, Yamamoto H, Imai K, Shinomura Y, Esteller M (2009) Epigenetic inactivation of the circadian clock gene BMAL1 in hematologic malignancies. Cancer Res 69:8447–8454. https://doi.org/10.1158/0008-5472.CAN-09-0551
Andreani TS, Itoh TQ, Yildirim E, Hwangbo DS, Allada R (2015) Genetics of circadian rhythms. Sleep Med Clin 10:413–421. https://doi.org/10.1016/j.jsmc.2015.08.007
Rosbash M (2009) The implications of multiple circadian clock origins. PLoS Biol 7:e62. https://doi.org/10.1371/journal.pbio.1000062
Mazzoccoli G, Rubino R, Tiberio C, Giuliani F, Vinciguerra M, Oben J, De Cata A, Tarquini R, De Cosmo S, Liu S, Cai Y (2016) Clock gene expression in human and mouse hepatic models shows similar periodicity but different dynamics of variation. Chronobiol Int 33:181–190. https://doi.org/10.3109/07420528.2015.1132722
Tsinkalovsky O, Smaaland R, Rosenlund B, Sothern RB, Hirt A, Steine S, Badiee A, Abrahamsen JF, Eiken HG, Laerum OD (2007) Circadian variations in clock gene expression of human bone marrow CD34+ cells. J Biol Rhythms 22:140–150. https://doi.org/10.1177/0748730406299078
Briese E (1998) Normal body temperature of rats: the setpoint controversy. Neurosci Biobehav Rev 22:427–436. https://doi.org/10.1016/s0149-7634(97)00051-1
Satlin A, Volicer L, Stopa EG, Harper D (1995) Circadian locomotor activity and core-body temperature rhythms in Alzheimer’s disease. Neurobiol Aging 16:765–771. https://doi.org/10.1016/0197-4580(95)00059-n
Guan J, Ding Y, Liu Y, Li Y, Liu Y, Wang Z (2009) Circadian effects on outcome following surgery for intracerebral hemorrhage in humans? Brain Res 1258:78–85. https://doi.org/10.1016/j.brainres.2008.11.106
Lu HY, Huang AP, Kuo LT (2021) Prognostic value of circadian brain temperature rhythm in basal ganglia hemorrhage after surgery. Neurol Ther 10:1045–1059. https://doi.org/10.1007/s40120-021-00283-y
Kirkness CJ, Burr RL, Thompson HJ, Mitchell PH (2008) Temperature rhythm in aneurysmal subarachnoid hemorrhage. Neurocrit Care 8:380–390. https://doi.org/10.1007/s12028-007-9034-y
Duhaime AC, Gennarelli LM, Boardman C (1996) Neuroprotection by dextromethorphan in acute experimental subdural hematoma in the rat. J Neurotrauma 13:79–84. https://doi.org/10.1089/neu.1996.13.79
Macpherson P, Graham DI (1978) Correlation between angiographic findings and the ischaemia of head injury. J Neurol Neurosurg Psychiatry 41:122–127. https://doi.org/10.1136/jnnp.41.2.122
Miller JD, Bullock R, Graham DI, Chen MH, Teasdale GM (1990) Ischemic brain damage in a model of acute subdural hematoma. Neurosurg 27:433–439. https://doi.org/10.1097/00006123-199009000-00016
Korthas HT, Main BS, Harvey AC, Buenaventura RG, Wicker E, Forcelli PA, Burns MP (2022) The effect of Traumatic Brain Injury on Sleep architecture and circadian rhythms in mice-a comparison of high-frequency head impact and controlled cortical injury. Biology (Basel) 8(11):1031. https://doi.org/10.3390/biology11071031
Boone DR, Sell SL, Micci MA, Crookshanks JM, Parsley M, Uchida T, Prough DS, DeWitt DS, Hellmich HL (2012) Traumatic brain injury-induced dysregulation of the circadian clock. PLoS ONE 7:e46204. https://doi.org/10.1371/journal.pone.0046204
Li B, Li D, Ni H, Liu c, Xiong J, Liu H, Gao R, Zhang L, Chen G, (2022) The circadian clock regulator Bmal1 affects traumatic brain injury in rats through the p38 MAPK signalling pathway. Brain Res Bull 178:17–28. https://doi.org/10.1016/j.brainresbull.2021.11.003
Refinetti R (2010) Entrainment of circadian rhythm by ambient temperature cycles in mice. J Biol Rhythms 25:247–256. https://doi.org/10.1177/0748730410372074
Refinetti R (2015) Comparison of light, food, and temperature as environmental synchronizers of the circadian rhythm of activity in mice. J Physiol Sci 65:359–366. https://doi.org/10.1007/s12576-015-0374-7
Ohi K, Takashima M, Nishikawa T, Takahashi K (1991) N-methyl-D-aspartate receptor participates in neuronal transmission of photic information through the retinohypothalamic tract. Neuroendocrinol 53:344–348. https://doi.org/10.1159/000125740
Crompton MR (1971) Hypothalamic lesions following closed head injury. Brain 94:165–172. https://doi.org/10.1093/brain/94.1.165
Acknowledgements
We thank Editage (www.editage.com) for the English language editing.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Lu-Ting Kuo: conceptualization, methodology, formal analysis, writing—original draft, writing—review and editing, supervision, project administration. Hsueh-Yi Lu: methodology, formal analysis, investigation. Yi-Hsing Chen: data curation, formal analysis, investigation. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This study complied with the ARRIVE guidelines and was carried out in accordance with the UK Animals (Scientific Procedures) Act, 1986 and associated guidelines, EU Directive 2010/63/EU for animal experiments, or the National Research Council's Guide for the Care and Use of Laboratory Animals. The study was approved by the Institutional Animal Care and Use Committee of National Taiwan University, College of Medicine (approval no. 20120485; date of approval: April 20, 2012).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kuo, LT., Lu, HY. & Chen, YH. Traumatic brain injury-induced disruption of the circadian clock. J Mol Med 102, 403–414 (2024). https://doi.org/10.1007/s00109-024-02416-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-024-02416-w