Abstract
Nucleolin (NCL) is a multifunctional protein expressed in the nucleus, cytoplasm, and cell membrane. Overexpression of NCL has a controversial role as a poor prognostic marker in cancers. In this study, a meta-analysis was performed to evaluate the prognostic value of NCL in different subcellular localizations (cytoplasmic (CyNCL) and nuclear (NuNCL)) across a range of cancers. PubMed was searched for relevant publications. Data were extracted and analyzed from 12 studies involving 1221 patients with eight cancer types. The results revealed high total NCL was significantly associated with poor overall survival (OS) (HR = 2.85 (1.94, 4.91), p < 0.00001, I2 = 59%) and short disease-free survival (DFS) (HR = 3.57 (2.76, 4.62), p < 0.00001, I2 = 2%). High CyNCL was significantly associated with poor OS (HR = 4.32 (3.01, 6.19), p < 0.00001, I2 = 0%) and short DFS (HR = 3.00 (2.17, 4.15), p < 0.00001, I2 = 0%). In contrast, high NuNCL correlated with increased patient OS (HR = 0.42 (0.20, 0.86), p = 0.02, I2 = 66%), with no significant correlation to DFS observed (HR = 0.46 (0.19, 1.14), p = 0.09, I2 = 57%). This study supports the role of subcellular NCL as a poor prognostic cancer biomarker.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nucleolin (NCL) is a eukaryotic nucleolar phosphoprotein involved in the synthesis and maturation of ribosomes, gene silencing, senescence, cytokinesis, cell proliferation, and growth [1,2,3,4,5]. NCL is predominately located in dense fibrillar regions of the nucleolus, being observed less frequently in the cytoplasm and membrane [6,7,8,9,10]. NCL was reported to be expressed in the nuclear fraction of normal human mammary epithelial cells, with only low levels observed in the cytoplasm [11]. Moreover, nuclear NCL (NuNCL) was observed in a variety of normal human tissues, including hepatocytes and cholangiocytes in the liver, endocrine cells and exocrine glandular cells in the pancreas, respiratory epithelial cells in the nasopharynx and bronchus, squamous epithelial cells in the esophagus, and glandular cells in the stomach, according to immunohistochemistry (IHC) (data obtained from The Human Protein Atlas (https://www.proteinatlas.org/)) [12].
In cancer, NCL is reported to be overexpressed with altered subcellular localization. Immune fluorescent and cell fractionation studies demonstrated that an increase in cytoplasmic NCL and a decrease in nuclear NCL were observed in the MCF-7 breast cancer cell line compared to a normal breast cancer cell line [11]. In human cancer specimens, IHC studies report that an increase in expression of NCL (without defining the subcellular localization) was associated with poor prognosis in pediatric and adult ependymoma [13, 14], hepatocellular carcinoma [15], non-small cell lung cancer [16], esophageal squamous cell carcinomas [17], and B cell lymphoma [18]. In contrast, no significant association with NCL expression and prognosis was reported in studies of ependymoma [14] hepatocellular carcinoma [19] and pancreatic ductal adenocarcinoma [20]. High NCL expression in the cytoplasm was associated with a poor prognosis in gastric cancer patients [21] and endometrial cancer [22] whereas the high nuclear NCL expression was an independent good prognostic marker in gastric cancer, endometrial carcinoma, and pancreatic ductal adenocarcinoma [21,22,23]. However, when subcellular localization is considered, NuNCL was reported to be significantly higher in fibrosarcoma, chondrosarcoma, liposarcoma, rhabdomyosarcoma, testicular tumor, and cutaneous melanocytic lesion tissues than in normal adjacent tissues, and the mRNA expression level has been linked to poor survival in patients [24,25,26]. On balance, there is a body of evidence in the literature that supports NCL as a potential prognostic marker in cancer; however, whether its subcellular expression levels are associated with good or poor prognosis requires further clarification.
As NCL has distinct functions depending on different cellular compartments in cancer cells, it is not surprising that intracellular localization is associated with varying patient outcomes [27]. NuNCL can regulate ribosomal DNA (rDNA) and rRNA syntheses, ribosome assembly, and the transcriptional activities of RNA polymerases (RNA pol) I and II [27,28,29] leading to cell proliferation (Fig. 1). It can bind with vascular endothelial growth factor (VEGF) promotor and increases the expression of VEGF which eventually promotes angiogenesis [30]. Furthermore, NuNCL has been implicated in the regulation of miRNAs, and transcription factors such as TBX3 that are involved in tumorigenesis promote cell proliferation and cell migration [24, 31, 32]. However, NuNCL has also been reported to bind with replication protein A (RPA) and prevent DNA replication by inhibiting RPA DNA replication initiation and elongation in human bone osteosarcoma epithelial cells [33]. Moreover, NCL binds to several DNA repair proteins such as topoisomerase (Topo) in U-937 leukemic cell [34] and Rad51 in human fibrosarcoma cells [35]; Ser-139 phosphorylation of H2A histone family member X (γH2AX) in HeLa cells [36] facilitates double-stranded break DNA repair machinery which then can delay cell proliferation. In addition, NCL promoted cisplatin resistance via the YB1-MDR pathway in cervical cancer which has been reported through the NCL-mediated cell proliferation leading to the attenuation of cancer cell sensitivity to cisplatin. This overexpression of NCL was associated with increased multidrug resistance (MDR1) gene expression resulting to the increment of drug efflux via transcription factor YB1 [37].
Subcellular NCL functions in cancer cell. NCL: nucleolin, rRNA: ribosomal ribonucleic acid, rDNA: ribosomal deoxyribonucleic acid, RNA Pol I and II: RNA polymerase I and II, VEGF: vascular endothelial growth factor, Topo: topoisomerase, RPA: replication protein A, DNA Pol: DNA polymerase, DSB: DNA double-strand break, Rad51: DNA repair protein RAD51 homolog 1, γH2AX: Ser-139 phosphorylation of H2A histone family member X, Bcl-2: B-cell lymphoma 2; Fas: Fas cell surface death receptor, Fas-L: Fas cell death ligand, Ras: Rat sarcoma, ErbB: receptor tyrosine-protein kinase erbB, MAPK: mitogen-activated protein kinase
In contrast, cytoplasmic NCL (CyNCL) and membranous NCL (MemNCL) are linked to proliferation, anti-apoptosis, and migration [9, 27]. CyNCL inhibits apoptosis by interacting with the 5′ UTR of p53 resulting in decreasing p53 translation [38] and increasing the stability of anti-apoptotic Bcl-2 mRNA in leukemic cells [39] (Fig. 1). MemNCL interacts with Fas receptor to prevent Fas-induced apoptosis activated by Fas-ligand (FAS-L) in B cell lymphoma cells [40]. CyNCL and MemNCL have been associated with an anti-apoptotic phenotype in cancer cells. Moreover, MemNCL has been reported to promote cell proliferation and tumor growth by binding to Ras and activating the Ras/MAPK cascade as the result of erythroblastic leukemia viral oncogene homolog (ErbB) receptor activation in colon cancer cells and prostate cancer cells [41]. Additionally, the MemNCL induced by VEGF via PI3K/Akt pathway in colorectal cancer [42] can act as an adhesion molecule to interact with collagen and laminin leading to cancer cell migration [43] (Fig. 1).
Based on the previously published evidence, NCL may be a potential cancer marker, and its subcellular localization may be useful to determine the prognosis of the cancer patients. Herein, the purpose of this study is to evaluate the prognostic value of NCL in varying subcellular locations in cancers using meta-analysis to propose the specific subcellular NCL as a potent prognostic marker in the patients.
Materials and methods
Literature search
An online literature search (PubMed, https://pubmed.ncbi.nlm.nih.gov/) was conducted between the 24th of October 2021 and the 27th of November 2021 to assess the level of NCL and its clinicopathological correlation in several cancers. The PubMed literature was filtered using the keywords “nucleolin” or “NCL” in combination with “expression” and “cancer”.
Inclusion and exclusion criteria
The studies were considered eligible if they met the following criteria: (1) NCL was detected in cancer tissues using immunohistochemistry (IHC) and (2) the hazard ratio (HR) for survival rate of either overall survival (OS) or disease-free survival (DFS) was calculated, and the 95% confidence intervals (CI) were provided. The following studies were excluded: (1) non-human studies; (2) non-English language studies; (3) in vitro studies; and (4) articles from which the relevant data could not be extracted.
Data extraction
Each eligible study was reviewed by two investigators (JP and SY), and the following information was extracted: the first author’s name, publication date, country, number of participants, age, gender, the percentage of NCL positivity, and the cellular localization of NCL. Furthermore, if the HR and 95% CI were reported in the text or survival table, they were collected. When it was not possible to extract HR directly from the article, Kaplan–Meier (KM) curves were used to estimate HR following the method of Tierney et al. [44]. Briefly, the percentage of survival over the interval times was extracted from the KM curve using the GetData Graph Digitizer 2.26 (http://getdata-graph-digitizer.com/download.php) and input into the established spreadsheet to generate the estimate HR and 95% CI. Disagreements between reviewers were settled through discussion.
Statistical analysis
The association of NCL expression in cancer patient survival time was evaluated by HR with an estimate of 95% CI. If the articles presented both univariate (UV) and multivariate analysis (MV), MV analysis was preferred. The heterogeneity of the data from eligible studies was evaluated by I2 statistic, which is a quantitative measure of inconsistency across studies using a random effect model. The I2 varies from 0 (no observed heterogeneity) to 100% (maximal heterogeneity). I2 value of more than 50% was considered to represent substantial heterogeneity among studies. Statistical significance was defined as p-value < 0.05. All analyses were performed using Review Manager (RevMan) version 5.4 (The Nordic Cochrane Centre, The Cochrane Collaboration, Copenhagen, Denmark).
IHC staining of NCL and scoring in breast cancer tissues
NCL was detected on the paraffin-embedded tumor microarray of 147 TNBC cases under the approval of sample collection by Siriraj Institutional Review Board (COA no. Si 580/2018). The staining protocol was as reported previously [45]. In brief, 4-µm-thick sections were incubated with anti-NCL antibody (#14574, Cell Signaling Technology, Inc) in a humidified chamber at 4 °C and then with rabbit Envision System HRP-labeled polymer IHC secondary antibody (K4003, DAKO) for 30 min at RT (room temperature). The peroxidase activity was visualized with diaminobenzidine (DAB) solution and counterstained by hematoxylin. The staining proteins were quantitatively scored by scanning the slides with 3DHistech Ltd. CaseViewer/QuantCenter software 2.4.0. (Sysmex) and scored based on: 0x% not stained + 1x% weakly stained + 2x% moderately stained + 3x% strongly stained. This gave a range of scores from 0 to 300 with nuclear, cytoplasmic, and membrane tumor-specific staining scored separately. The expression of protein at each cellular compartment was divided into low and high using the cutoff point calculated by RStudio version 2022.2.0.443 (Integrated Development for R. RStudio, PBC, Boston, MA URL http://www.rstudio.com/).
Results
Study selection and characteristic
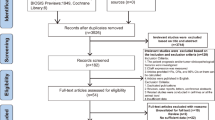
Following the initial PubMed search, 391 papers were identified. Based on their titles and content of abstracts, 265 articles were removed including 17 review articles, 79 non-human research, 10 non-English articles, 96 in vitro functional tests, and 159 irrelevant to NCL. The remaining 30 papers were identified through full text. Eighteen papers were excluded due to insufficient data for example no data of OS or DFS and did not use IHC for NCL expression measurement. Finally, 12 studies met the inclusion criteria for the meta-analysis (Fig. 2 and Table 1).
Study characteristics
The characters of the 12 papers included in the study were listed (Table 1). The included studies were published between 2008 and 2021. The OS was reported in 9 studies, and DFS was reported in 6 articles. The 12 papers include 3 studies on pediatric and adult ependymoma [13, 14, 46], two on hepatocellular carcinoma [15, 19], two on pancreatic ductal carcinoma [20, 23], two on non-small cell lung cancer [16, 47], one on endometrial cancer [22], one on gastric cancer [21], and one on B-cell lymphoma [18].
The antibodies used in IHC for NCL detection in the cancer tissues were from different clones with a variety of binding epitopes binding sites, binding to amino acids 2–17 [16] and 271–520 [18] of the 710 amino acids NCL protein or undefined NCL epitope using whole human NCL protein from Raji cell extract as the immunogen for the antibody production [13,14,15, 21,22,23, 46] or antibody clones that did not provide the antigen binding site [19, 20, 47]. The cellular localization of NCL expression was divided into three groups including expression in all cellular compartments (total NCL), CyNCL, and NuNCL (Table 1). According to these three subcellular classifications, only 8 papers reported total NCL with a total of 721 patients [13,14,15,16, 18,19,20, 46]; 1 paper reported total NCL, CyNCL, and NuNCL (225 patients) [47]; 2 papers reported CyNCL and NuNCL (206 patients) [21, 22]; and 1 paper reported only NuNCL (69 patients) [23]. Notably, no papers reported only CyNCL. For total NCL (from the articles which the authors did not provide the localization of NCL results in their work), we had 9 papers to be analyzed which the figure results confirmed the combination of both CyNCL and NuNCL [13,14,15,16, 18,19,20, 46, 47]. For CyNCL, 3 papers were provided [21, 22, 47], whereas 4 papers offered data of NuNCL [21,22,23, 47]. Three papers directly provided survival data of only DFS [13, 22, 46]. Six papers showed only patient OS data, of which two of them directly presented HR and 95% CI [21, 23], while the remaining 4 papers, the estimated HR and 95% CI were performed [16, 18,19,20]. Three papers reported both OS and DFS [14, 15, 47], of which 1 estimated HR and 95% CI [47].
The impact of total NCL, CyNCL, and NuNCL on cancer patient OS
The meta-analysis was performed on 7 studies for total NCL expression (708 patients), 2 studies for cytoplasmic NCL expression (349 patients), and 3 articles contained 418 patients for NuNCL expression assessing the association of each NCL localization with patient OS. The results from total NCL exression group showed that high NCL expression had a significant association with poor OS (HR = 2.85, 95% CI = (1.94, 4.19), p < 0.00001) with heterogeneity (I2 = 59%) (Fig. 3). High expression of CyNCL was significantly associated with poor OS in the patients (HR = 4.32, 95% CI = (3.01, 6.19), p < 0.00001) without heterogeneity (I2 = 0%). In contrast with total NCL and CyNCL, high expression of NuNCL was significantly associated with improved patient outcome (HR = 0.42, 95% CI = (0.2, 0.86), p = 0.02) with heteroginiety (I2 = 66%) (Fig. 3). The combined results of the combined total NCL, CyNCL, and NuNCL still revealed as the poor prognostic marker with HR = 1.81 (95% CI = 1.02, 3.21, p = 0.04, I2 = 90%) which notably was lower than that of total NCL alone (HR = 2.85) or of CyNCL alone (HR = 4.32). The results indicated that CyNCL had the significant impact on patient survival as a poor prognostic marker followed by the total NCL. Of note, the combined NCL (total, CyNCL, and NuNCL) showed no applicable used as it reduced the prediction of short patient OS. In contrast, NuNCL revealed the impact of a predictive marker for long patient OS with statistical sifgnificance.
High total NCL and high CyNCL were associated with poor OS in cancer patients
As both total NCL and CyNCL were associated with poor OS in the patients, their combined HR was determined. The results showed that the combined HR of the total NCL and CyNCL was associated with poor OS (HR = 3.14, 95% CI = (2.31, 4.26), p < 0.00001, I2 = 0%). This HR was lower than that of CyNCL (HR = 4.32), but higher than that of total NCL (HR = 2.85) (Fig. 4). These results may suggest the NCL in the cytoplasm as the best prognostic NCL marker for shorter OS.
Impact of total, CyNCL, and NuNCL expressions on DFS of cancer patients
Six articles, consisting of 747 patients for total NCL expression; 2 articles (307 samples) from cytoplasmic expression; and 2 articles (357 patients) from nuclear expression were included to determine the effect of total NCL, CyNCL, and NuNCL on DFS (Fig. 5). High expression of total NCL showed significant association with poor DFS (HR = 3.57, 95% CI = (2.76, 4.62), p < 0.00001) with homogeniety (I2 = 2%). In the same trend, high CyNCL was significantly associated with poor DFS (HR = 3.00, 95% CI = (2.17, 4.15), p < 0.00001) with non-heterogeneity (I2 = 0%) whereas high expression of NCL in the nucleus has no significant corelattion with DFS in the patients (HR = 0.46, 95% CI = (0.19, 1.14), p = 0.09) with heterogeniety (I2 = 57%) (Fig. 5). In the similar manner to OS, the combined HR (total NCL, CyNCL, and NuNCL) had the reduced HR compared to using either total NCL or CyNCL for DFS (HR = 2.37, 95% CI = (1.30, 4.32), p = 0.005) with heterogeneity (I2 = 90%) (Fig. 5). The results indicated that total NCL and CyNCL may predict short DFS whereas NuNCL would be a predictive marker for long DFS in patients; however, this was not statistically significant.
High total NCL and high CyNCL were associated with poor DFS in cancer patients
The combined HR and 95% CI of total NCL and CyNCL significantly correlated with poor DFS with HR of 3.34 (95% CI = 2.74, 4.08) with statistical significance (p < 0.00001) without heterogeneity (I2 = 0%) (Fig. 6). The total NCL (HR = 3.57) and CyNCL (HR = 3.00) showed almost the same HR to that of the combined HR as being the poor prognostic markers for patient DFS time.
High CyNCL was associated with poor OS in triple-negative breast cancer (TNBC) patients
We have recently reported the total NCL in clinical samples and its correlation with poor prognosis in TNBC patients [45]. From the same set of patient samples, we analyzed NuNCL and CyNCL and performed the KM analysis. A significant association between CyNCL and poor survival patient was observed (p = 0.014), while no significant correlation between NuNCL and patient survival was observed (Fig. 7). These findings support the current meta-analysis where CyNCL, but not NuNCL, was associated with poor prognosis in TNBC patients. Interestingly, no memNCL was detected in these samples.
Discussion
NCL is overexpressed in a variety of cancers. Overexpression of NCL mRNA was a marker of poor OS and DFS in triple-negative breast cancer [48], acute myeloid leukemia [49], and neuroblastoma [50]. Moreover, overexpression of total NCL protein detected by IHC was reported to be significantly associated with poor OS [15, 16, 18] and DFS [13,14,15, 47] in pediatric ependymoma, hepatocellular carcinoma, pancreatic ductal adenocarcinoma, and non-small cell lung cancer, but no significance with patient outcome was reported in the studies with regards to ependymoma, hepatocellular carcinoma, and pancreatic ductal adenocarcinoma [14, 19, 20]. Three studies reported an association with cytoplasmic NCL expression and poor prognosis in endometrial carcinoma, gastric cancer, and non-small cell lung cancer [21, 22, 47], compared to nuclear NCL being associated with good prognosis [21,22,23]. In the current study, a meta-analysis was performed to investigate the impact of different subcellular NCL localization as prognosis markers for cancer patients to provide supporting evidence for differing NCL cellular localization being associated with different patient outcome. The results using HR values as the predictive markers reveal that cytoplasmic NCL is a strong predictive marker for short patient OS, and both cytoplasmic NCL and total NCL are associated with short patient DFS. Interestingly, the nuclear NCL represents the predictive marker for patient long OS and DFS.
The different functions of subcellular NCL have been reported [27], and the potential mechanism of each subcellular NCL is summarized in Fig. 1. The depletion of NuNCL resulted in a decrease pre-rRNA and was associated with rDNA heterochromatinization [51]. NCL overexpression, on the other hand, caused an increase in pre-rRNA levels [51]. NCL has been shown to interact with rDNA chromatin [29], particularly the promoter and coding region of unmethylated rRNA genes [51]. The binding of NuNCL to rDNA inhibited the binding of thyroid transcription factor 1 (TTF-1) to the promoter-proximal terminator T0. Because TTF-1 binding is required for histone deacetylase (HDAC) recruitment, it was proposed that NCL inhibited the formation of repressive heterochromatin and required for the maintenance of a euchromatin active state, promoting active transcription of rDNA [51]. Furthermore, NuNCL forms a complex with replication protein A (RPA), a ssDNA binding protein that is required for DNA replication initiation and elongation. As a result, RPA sequestration by NCL may prevent DNA replication [52]. The cytoplasmic NCL interacted with some RNAs supporting their stability and translation. The cytoplasmic NCL recognized the AU-rich element (AUUUA) in the 3’ UTR of Bcl-xl mRNA [53]. This interaction protected Bcl-xl mRNA from nuclease degradation [53]. NCL also bound to the 3’ UTR of Bcl-2 mRNA, increasing its stability and allowing tumor cells to escape the apoptotic pathway [54]. CyNCL bound to the 5’ UTR of p53 mRNA and inhibited its translation allowing tumor cells to avoid apoptosis [27]. Membrane NCL interacted with Fas and blocked Fas-FasL interaction preventing Fas-mediated apoptosis [40]. In addition, NCL interaction with ErbB1 [55] and Ras [56] at the plasma membrane favored cell proliferation. Moreover, membrane NCL interaction with Ras-GTP increased the interaction of NCL with ErbB1 leading to an accumulation of Ras-GTP. NCL, ErbB1, and Ras acted synergistically in mediating tumor growth in nude mice [56] and in favoring cancer cell proliferation and survival in vitro in human colon cancer cells and prostate cancer cells [41].
Hence, the papers included in the current study were divided into three groups based on the results of the full-text screening: total, cytoplasmic, and nuclear NCL expression. Interestingly, membrane NCL was reported to be associated with cancer progression and explored as a target for cancer treatment in the clinical trials using various molecules such as AS1411 [57]. The membrane NCL data cannot be separately analyzed for its prognostic value as the result of no membrane NCL reported in the recruited articles in this meta-analysis study.
When compared to total NCL expression, high cytoplasmic NCL expression had a strong prognostic value for OS, whereas total and cytoplasmic NCL expressions had the same prognostic value for DFS. These findings can be supported by the previous report of the cytoplasmic NCL function to promote cell proliferation, anti-apoptosis, tumor migration, and metastasis leading to the aggressive phenotypes of cancer cells [9]. Hence, it is strongly suggested that cytoplasmic NCL is a high-impact marker for short survival time in cancer patients. Using cytoplasmic NCL in combination with total NCL or cytoplasmic + nuclear + total NCL can predict bad prognosis. Interestingly, only cytoplasmic NCL is the most potent marker for aggressive cancer which may require aggressive treatment and intensive follow-up. Our result from the TNBC cases confirms the prognostic value of CyNCL as was associated with poor prognosis in TNBC patients.
High expression of NuNCL exhibited the statistically significant marker for long OS, even though no significant correlation with patient long DFS. This could be explained by NuNCL regulating gene transcription, DNA replication, and DNA repair [33,34,35,36], resulting in low capability of cancer cell to grow and survive. Though the proposed functions of NCL in the nucleus to control angiogenesis which can then induce aggressive cancer resulting to shot survival time, several mechanisms involved in new vessels formation [30]. Therefore, NuNCL may have major function in controlling gene expression involved in inhibiting cell proliferation. A future study involving target genes controlled by NuNCL is required to explore and better understand how NuNCL contributes to good prognosis in cancer patients.
Total NCL has been reported, which does not specify which cellular compartment NCL is predominantly expressed in. Therefore, the representative images of NCL protein expression were carefully examined and on examining the images of the published papers recruited in this study, only half of the papers displayed NuNCL; the remaining were observed to have both NuNCL and CyNCL expression, with the exception of one paper where only CyNCL was observed. As NuNCL and CyNCL, supported by our findings and functional studies, demonstrate distinct function and prognosis value, the subcellular NCL score should be considered separately for use as a cancer prognosis marker rather than using total scoring.
The overall results emphasize the prognostic value of subcellular NCL as a potential cancer prognosis marker. However, there are two main limitations to this study: (1) There was a high heterogeneity among the studies, which could be attributed to the pooling of data sets from various cancer types due to the limited number of NCL studies in each cancer and (2) the lower precision of HR due to the extraction method rather than directly acquired from the original data. To validate these findings, a larger and well-characterized patient cohort from the same cancer type, employing the same antibody, is required.
Conclusion
The finding from this meta-analysis supports the observation that high NCL expression is associated with poor prognosis in cancer patients including ependymoma, hepatocellular carcinoma, non-small cell lung cancer, pancreatic ductal carcinoma, endometrial carcinoma, gastric cancer, and B cell lymphoma. We propose herein, using subcellular localization of NCL is more suitable as a prognosis marker in cancers than the total NCL. High CyNCL expression is a prognosis marker for short survival time, whereas high NuNCL expression is a potential prediction marker for prolong survival in cancer patient. However, further cohort studies are required to support this conclusion; this information highlights the importance of different cellular localization of NCL with different proposed functions leading to the potential of being predictive marker for bad or good prognosis in cancer patients.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Tajrishi MM, Tuteja R, Tuteja N (2011) Nucleolin: the most abundant multifunctional phosphoprotein of nucleolus. Commun Integr Biol 4:267–275. https://doi.org/10.4161/cib.4.3.14884
Ginisty H, Sicard H, Roger B, Bouvet P (1999) Structure and functions of nucleolin. J Cell Sci 112(Pt 6):761–772
Bouvet P, Diaz JJ, Kindbeiter K, Madjar JJ, Amalric F (1998) Nucleolin interacts with several ribosomal proteins through its RGG domain. J Biol Chem 273:19025–19029. https://doi.org/10.1074/jbc.273.30.19025
Pederson T (1998) The plurifunctional nucleolus. Nucleic Acids Res 26:3871–3876. https://doi.org/10.1093/nar/26.17.3871
Mongelard F, Bouvet P (2007) Nucleolin: a multiFACeTed protein. Trends Cell Biol 17:80–86. https://doi.org/10.1016/j.tcb.2006.11.010
Biggiogera M, Burki K, Kaufmann SH, Shaper JH, Gas N, Amalric F, Fakan S (1990) Nucleolar distribution of proteins B23 and nucleolin in mouse preimplantation embryos as visualized by immunoelectron microscopy. Development 110:1263–1270
Bugler B, Caizergues-Ferrer M, Bouche G, Bourbon H, Amalric F (1982) Detection and localization of a class of proteins immunologically related to a 100-kDa nucleolar protein. Eur J Biochem 128:475–480. https://doi.org/10.1111/j.1432-1033.1982.tb06989.x
Lischwe MA, Richards RL, Busch RK, Busch H (1981) Localization of phosphoprotein C23 to nucleolar structures and to the nucleolus organizer regions. Exp Cell Res 136:101–109. https://doi.org/10.1016/0014-4827(81)90041-0
Jia W, Yao Z, Zhao J, Guan Q, Gao L (2017) New perspectives of physiological and pathological functions of nucleolin (NCL). Life Sci 186:1–10. https://doi.org/10.1016/j.lfs.2017.07.025
Escande ML, Gas N, Stevens BJ (1985) Immunolocalization of the 100 K nucleolar protein in CHO cells. Biol Cell 53:99–109. https://doi.org/10.1111/j.1768-322x.1985.tb00359.x
Soundararajan S, Chen W, Spicer EK, Courtenay-Luck N, Fernandes DJ (2008) The nucleolin targeting aptamer AS1411 destabilizes Bcl-2 messenger RNA in human breast cancer cells. Cancer Res 68:2358–2365. https://doi.org/10.1158/0008-5472.CAN-07-5723
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson A, Kampf C, Sjostedt E, Asplund A et al (2015) Proteomics. Tissue-based map of the human proteome. Science 347:1260419. https://doi.org/10.1126/science.1260419
Ridley L, Rahman R, Brundler MA, Ellison D, Lowe J, Robson K, Prebble E, Luckett I, Gilbertson RJ, Parkes S et al (2008) Multifactorial analysis of predictors of outcome in pediatric intracranial ependymoma. Neuro Oncol 10:675–689. https://doi.org/10.1215/15228517-2008-036
Modena P, Buttarelli FR, Miceli R, Piccinin E, Baldi C, Antonelli M, Morra I, Lauriola L, Di Rocco C, Garre ML et al (2012) Predictors of outcome in an AIEOP series of childhood ependymomas: a multifactorial analysis. Neuro Oncol 14:1346–1356. https://doi.org/10.1093/neuonc/nos245
Guo X, Xiong L, Yu L, Li R, Wang Z, Ren B, Dong J, Li B, Wang D (2014) Increased level of nucleolin confers to aggressive tumor progression and poor prognosis in patients with hepatocellular carcinoma after hepatectomy. Diagn Pathol 9:175. https://doi.org/10.1186/s13000-014-0175-y
Huang F, Wu Y, Tan H, Guo T, Zhang K, Li D, Tong Z (2019) Phosphorylation of nucleolin is indispensable to its involvement in the proliferation and migration of non-small cell lung cancer cells. Oncol Rep 41:590–598. https://doi.org/10.3892/or.2018.6787
Qi J, Li H, Liu N, Xing Y, Zhou G, Wu Y, Liu Y, Chen W, Yue J, Han B et al (2015) The implications and mechanisms of the extra-nuclear nucleolin in the esophageal squamous cell carcinomas. Med Oncol 32:45. https://doi.org/10.1007/s12032-015-0484-3
Jain N, Zhu H, Khashab T, Ye Q, George B, Mathur R, Singh RK, Berkova Z, Wise JF, Braun FK et al (2018) Targeting nucleolin for better survival in diffuse large B-cell lymphoma. Leukemia 32:663–674. https://doi.org/10.1038/leu.2017.215
Qiu W, Wang G, Sun X, Ye J, Wei F, Shi X, Lv G (2015) The involvement of cell surface nucleolin in the initiation of CCR6 signaling in human hepatocellular carcinoma. Med Oncol 32:75. https://doi.org/10.1007/s12032-015-0530-1
Gilles ME, Maione F, Cossutta M, Carpentier G, Caruana L, Di Maria S, Houppe C, Destouches D, Shchors K, Prochasson C et al (2016) Nucleolin targeting impairs the progression of pancreatic cancer and promotes the normalization of tumor vasculature. Cancer Res 76:7181–7193. https://doi.org/10.1158/0008-5472.CAN-16-0300
Qiu W, Zhou F, Zhang Q, Sun X, Shi X, Liang Y, Wang X, Yue L (2013) Overexpression of nucleolin and different expression sites both related to the prognosis of gastric cancer. APMIS 121:919–925. https://doi.org/10.1111/apm.12131
Lin Q, Ma X, Hu S, Li R, Wei X, Han B, Ma Y, Liu P, Pang Y (2021) Overexpression of nucleolin is a potential prognostic marker in endometrial carcinoma. Cancer Manag Res 13:1955–1965. https://doi.org/10.2147/CMAR.S294035
Peng L, Liang J, Wang H, Song X, Rashid A, Gomez HF, Corley LJ, Abbruzzese JL, Fleming JB, Evans DB et al (2010) High levels of nucleolar expression of nucleolin are associated with better prognosis in patients with stage II pancreatic ductal adenocarcinoma. Clin Cancer Res 16:3734–3742. https://doi.org/10.1158/1078-0432.CCR-09-3411
Willmer T, Damerell V, Smyly S, Sims D, Du Toit M, Ncube S, Sinkala M, Govender D, Sturrock E, Blackburn JM et al (2021) Targeting the oncogenic TBX3:nucleolin complex to treat multiple sarcoma subtypes. Am J Cancer Res 11:5680–5700
Mourmouras V, Cevenini G, Cosci E, Epistolato MC, Biagioli M, Barbagli L, Luzi P, Mannucci S, Miracco C (2009) Nucleolin protein expression in cutaneous melanocytic lesions. J Cutan Pathol 36:637–646. https://doi.org/10.1111/j.1600-0560.2008.01126.x
Masiuk M, Lewandowska M, Teresinski L, Dobak E, Urasinska E (2019) Nucleolin and nucleophosmin expression in seminomas and non-seminomatous testicular tumors. Folia Histochem Cytobiol 57:139–145. https://doi.org/10.5603/FHC.a2019.0015
Berger CM, Gaume X, Bouvet P (2015) The roles of nucleolin subcellular localization in cancer. Biochimie 113:78–85. https://doi.org/10.1016/j.biochi.2015.03.023
Ugrinova I, Monier K, Ivaldi C, Thiry M, Storck S, Mongelard F, Bouvet P (2007) Inactivation of nucleolin leads to nucleolar disruption, cell cycle arrest and defects in centrosome duplication. BMC Mol Biol 8:66. https://doi.org/10.1186/1471-2199-8-66
Rickards B, Flint SJ, Cole MD, LeRoy G (2007) Nucleolin is required for RNA polymerase I transcription in vivo. Mol Cell Biol 27:937–948. https://doi.org/10.1128/MCB.01584-06
Uribe DJ, Guo K, Shin YJ, Sun D (2011) Heterogeneous nuclear ribonucleoprotein K and nucleolin as transcriptional activators of the vascular endothelial growth factor promoter through interaction with secondary DNA structures. Biochemistry 50:3796–3806. https://doi.org/10.1021/bi101633b
Tominaga K, Srikantan S, Lee EK, Subaran SS, Martindale JL, Abdelmohsen K, Gorospe M (2011) Competitive regulation of nucleolin expression by HuR and miR-494. Mol Cell Biol 31:4219–4231. https://doi.org/10.1128/MCB.05955-11
Pichiorri F, Palmieri D, De Luca L, Consiglio J, You J, Rocci A, Talabere T, Piovan C, Lagana A, Cascione L et al (2013) In vivo NCL targeting affects breast cancer aggressiveness through miRNA regulation. J Exp Med 210:951–968. https://doi.org/10.1084/jem.20120950
Kim K, Dimitrova DD, Carta KM, Saxena A, Daras M, Borowiec JA (2005) Novel checkpoint response to genotoxic stress mediated by nucleolin-replication protein a complex formation. Mol Cell Biol 25:2463–2474. https://doi.org/10.1128/MCB.25.6.2463-2474.2005
Bharti AK, Olson MO, Kufe DW, Rubin EH (1996) Identification of a nucleolin binding site in human topoisomerase I. J Biol Chem 271:1993–1997. https://doi.org/10.1074/jbc.271.4.1993
De A, Donahue SL, Tabah A, Castro NE, Mraz N, Cruise JL, Campbell C (2006) A novel interaction [corrected] of nucleolin with Rad51. Biochem Biophys Res Commun 344:206–213. https://doi.org/10.1016/j.bbrc.2006.03.113
Kobayashi J, Fujimoto H, Sato J, Hayashi I, Burma S, Matsuura S, Chen DJ, Komatsu K (2012) Nucleolin participates in DNA double-strand break-induced damage response through MDC1-dependent pathway. PLoS ONE 7:e49245. https://doi.org/10.1371/journal.pone.0049245
Ke J, Gu C, Zhang H, Liu Y, Zhang W, Rao H, Li S, Wu F (2021) Nucleolin promotes cisplatin resistance in cervical cancer by the YB1-MDR1 pathway. J Oncol 2021:9992218. https://doi.org/10.1155/2021/9992218
Chen J, Guo K, Kastan MB (2012) Interactions of nucleolin and ribosomal protein L26 (RPL26) in translational control of human p53 mRNA. J Biol Chem 287:16467–16476. https://doi.org/10.1074/jbc.M112.349274
Otake Y, Soundararajan S, Sengupta TK, Kio EA, Smith JC, Pineda-Roman M, Stuart RK, Spicer EK, Fernandes DJ (2007) Overexpression of nucleolin in chronic lymphocytic leukemia cells induces stabilization of bcl2 mRNA. Blood 109:3069–3075. https://doi.org/10.1182/blood-2006-08-043257
Wise JF, Berkova Z, Mathur R, Zhu H, Braun FK, Tao RH, Sabichi AL, Ao X, Maeng H, Samaniego F (2013) Nucleolin inhibits Fas ligand binding and suppresses Fas-mediated apoptosis in vivo via a surface nucleolin-Fas complex. Blood 121:4729–4739. https://doi.org/10.1182/blood-2012-12-471094
Schokoroy S, Juster D, Kloog Y, Pinkas-Kramarski R (2013) Disrupting the oncogenic synergism between nucleolin and Ras results in cell growth inhibition and cell death. PLoS ONE 8:e75269. https://doi.org/10.1371/journal.pone.0075269
Wu DM, Zhang P, Liu RY, Sang YX, Zhou C, Xu GC, Yang JL, Tong AP, Wang CT (2014) Phosphorylation and changes in the distribution of nucleolin promote tumor metastasis via the PI3K/Akt pathway in colorectal carcinoma. FEBS Lett 588:1921–1929. https://doi.org/10.1016/j.febslet.2014.03.047
Huang Y, Shi H, Zhou H, Song X, Yuan S, Luo Y (2006) The angiogenic function of nucleolin is mediated by vascular endothelial growth factor and nonmuscle myosin. Blood 107:3564–3571. https://doi.org/10.1182/blood-2005-07-2961
Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR (2007) Practical methods for incorporating summary time-to-event data into meta-analysis. Trials 8:16. https://doi.org/10.1186/1745-6215-8-16
Thongchot S, Jirapongwattana N, Luangwattananun P, Chiraphapphaiboon W, Chuangchot N, Sa-Nguanraksa D, P OC, Thuwajit P, Yenchitsomanus PT, Thuwajit C, (2022) Adoptive transfer of anti-nucleolin T cells combined with PD-L1 inhibition against triple-negative breast cancer. Mol Cancer Ther 21:727–739. https://doi.org/10.1158/1535-7163.MCT-21-0823
Chen C, Chen L, Yao Y, Qin Z, Chen H (2016) Nucleolin overexpression is associated with an unfavorable outcome for ependymoma: a multifactorial analysis of 176 patients. J Neurooncol 127:43–52. https://doi.org/10.1007/s11060-015-2007-7
Xu JY, Lu S, Xu XY, Hu SL, Li B, Li WX, Chang JY (2016) Prognostic significance of nuclear or cytoplasmic nucleolin expression in human non-small cell lung cancer and its relationship with DNA-PKcs. Tumour Biol 37:10349–10356. https://doi.org/10.1007/s13277-016-4920-6
Nguyen Van Long F, Lardy-Cleaud A, Bray S, Chabaud S, Dubois T, Diot A, Thompson AM, Bourdon JC, Perol D, Bouvet P et al (2018) Druggable nucleolin identifies breast tumours associated with poor prognosis that exhibit different biological processes. Cancers (Basel). https://doi.org/10.3390/cancers10100390
Marcel V, Catez F, Berger CM, Perrial E, Plesa A, Thomas X, Mattei E, Hayette S, Saintigny P, Bouvet P et al (2017) Expression profiling of ribosome biogenesis factors reveals nucleolin as a novel potential marker to predict outcome in AML patients. PLoS ONE 12:e0170160. https://doi.org/10.1371/journal.pone.0170160
Wang F, Zhou S, Qi D, Xiang SH, Wong ET, Wang X, Fonkem E, Hsieh TC, Yang J, Kirmani B et al (2019) Nucleolin is a functional binding protein for salinomycin in neuroblastoma stem cells. J Am Chem Soc 141:3613–3622. https://doi.org/10.1021/jacs.8b12872
Cong R, Das S, Ugrinova I, Kumar S, Mongelard F, Wong J, Bouvet P (2012) Interaction of nucleolin with ribosomal RNA genes and its role in RNA polymerase I transcription. Nucleic Acids Res 40:9441–9454. https://doi.org/10.1093/nar/gks720
Daniely Y, Borowiec JA (2000) Formation of a complex between nucleolin and replication protein A after cell stress prevents initiation of DNA replication. J Cell Biol 149:799–810. https://doi.org/10.1083/jcb.149.4.799
Zhang J, Tsaprailis G, Bowden GT (2008) Nucleolin stabilizes Bcl-X L messenger RNA in response to UVA irradiation. Cancer Res 68:1046–1054. https://doi.org/10.1158/0008-5472.CAN-07-1927
Ishimaru D, Zuraw L, Ramalingam S, Sengupta TK, Bandyopadhyay S, Reuben A, Fernandes DJ, Spicer EK (2010) Mechanism of regulation of bcl-2 mRNA by nucleolin and A+U-rich element-binding factor 1 (AUF1). J Biol Chem 285:27182–27191. https://doi.org/10.1074/jbc.M109.098830
Di Segni A, Farin K, Pinkas-Kramarski R (2008) Identification of nucleolin as new ErbB receptors- interacting protein. PLoS ONE 3:e2310. https://doi.org/10.1371/journal.pone.0002310
Farin K, Schokoroy S, Haklai R, Cohen-Or I, Elad-Sfadia G, Reyes-Reyes ME, Bates PJ, Cox AD, Kloog Y, Pinkas-Kramarski R (2011) Oncogenic synergism between ErbB1, nucleolin, and mutant Ras. Cancer Res 71:2140–2151. https://doi.org/10.1158/0008-5472.CAN-10-2887
Rosenberg JE, Bambury RM, Van Allen EM, Drabkin HA, Lara PN Jr, Harzstark AL, Wagle N, Figlin RA, Smith GW, Garraway LA et al (2014) A phase II trial of AS1411 (a novel nucleolin-targeted DNA aptamer) in metastatic renal cell carcinoma. Invest New Drugs 32:178–187. https://doi.org/10.1007/s10637-013-0045-6
Funding
The authors would like to thank Mid-Career Research Grant, The National Research Council of Thailand (grant no. RSA6280091), for financial support to CT.
Author information
Authors and Affiliations
Contributions
CT, JP, and SY contributed to the framework and overall perspective of the study design. The literature search was carried out by SY, JP, and ST. SY and JP extracted the data and assisted with quality control. JP carried out the statistical analysis. SY wrote the manuscript and created the tables and figures. The statistical analysis was supervised and verified by PH, JE, and CT. SY, JP, PT, PY, JE, and CT contributed to the study’s quality assessment and manuscript revision. The final manuscript was read and approved by all authors.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supaporn Yangngam and Jaturawit Prasopsiri are co-first authors.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yangngam, S., Prasopsiri, J., Hatthakarnkul, P. et al. Cellular localization of nucleolin determines the prognosis in cancers: a meta-analysis. J Mol Med 100, 1145–1157 (2022). https://doi.org/10.1007/s00109-022-02228-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-022-02228-w