Abstract
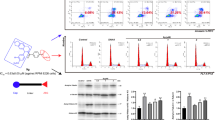
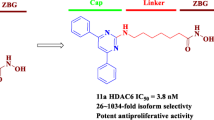
In recent years, histone deacetylase (HDAC) has become one of the hottest and most effective targets for the treatment of cancer. In this work, we designed and synthesized a series of novel o-aminobenzamide based HDAC inhibitors and evaluated their antitumor properties in vitro. All 23 compounds obtained showed micromolar IC50 values against A549 cells proliferation, and the most effective compound was 8 u (IC50 = 0.165 μM). In vitro, 8 u showed potent antiproliferative activity against another three cancer cell lines, outperforming the approved drug Chidamide. Enzyme inhibition and western blot assays confirmed that 8 u was a selective inhibitor of HDAC1-3 isoforms. 8 u was able to induce apoptosis of A549 cells and arrest the tumor cells in G2/M phase. Moreover, 8 u significantly mitigated the migration A549 cells. All these results suggest that 8 u deserves further biological studies.











Similar content being viewed by others
References
Ansari J, Shackelford RE, El-Osta H. Epigenetics in non-small cell lung cancer: from basics to therapeutics. Transl Lung Cancer Res. 2016;5:155–71. https://doi.org/10.21037/tlcr.2016.02.02
Bondarev AD, Attwood MM, Jonsson J, Chub-arev VN, Tarasov VV, Schiöth HB. Recent developments of HDAC inhibitors: emerging indications and novel molecules. Br J Clin Pharmacol. 2021;87:4577–97. https://doi.org/10.1111/bcp.14889
Ramaiah MJ, Tangutur AD, Manyam RR. Epigenetic modulation and understanding of HDAC inhibitors in cancer therapy. Life Sci. 2021;277:119504–22. https://doi.org/10.1016/j.lfs.2021.119504
Peserico A, Simone C. Physical and functional HAT/HDAC interplay regulates protein acetylation balance. J Biomed and Biotechnol. 2011;2011:1–10. https://doi.org/10.1155/2011/371832
Cheng B, Pan W, Xiao Y, Ding Z, Zhou Y, Fei X, et al. HDAC-targeting epigenetic modulators for cancer immunotherapy. Eur J Med Chem. 2024;265:116129–45. https://doi.org/10.1016/j.ejmech.2024.116129
Botrugno OA, Santoro F, Minucci S. Histone deacetylase inhibitors as a new weapon in the arsenal of differentiation therapies of cancer. Cancer Lett. 2009;280:134–44. https://doi.org/10.1016/j.canlet.2009.02.027
Eckschlager T, Plch J, Stiborova M, Hrabeta J. Histone deacetylase inhibitors as anticancer D-rugs. Int J Mol Sci. 2017;18:1414–38. https://doi.org/10.3390/ijms18071414
Shetty MG, Pai P, Deaver RE, Satyamoorthy K, Babitha KS. Histone deacetylase 2 selective inhibitors: a versatile therapeutic strategy as next generation drug target in cancer therapy. Pharmacol Res. 2021;170:105695–7. https://doi.org/10.1016/j.phrs.2021.105695
Sangwan R, Rajan R, Mandal PK. HDAC as onco target: Reviewing the synthetic approaches with SAR study of their inhibitors. Eur J Med Chem. 2018;158:620–706. https://doi.org/10.1016/j.ejmech.2018.08.073
Yuan H, Marmorstein R. Structural basis for sirtuin activity and inhibition. J Biolog Chem. 2012;287:42428–35. https://doi.org/10.1074/jbc.R112.372300
Smalley JP, Baker IM, Pytel WA, Lin L-Y, Bowman KJ, Schwabe JWR, et al. Optimization of class I histone deacetylase PROTACs reveals that HDAC1/2 degradation is critical to induce apoptosis and cell arrest in cancer cells. J Med Chem. 2022;65:5642–59. https://doi.org/10.1021/acs.jmedchem.1c02179
Schübeler D, Hess L, Moos V, Lauber AA, Reiter W, Schuster M, et al. A toolbox for class I HDACs reveals isoform specific roles in gene regulation and protein acetylation. PLOS Genet. 2022;18:1010376–92. https://doi.org/10.1371/journal.pgen.1010376
Li J, Lu L, Liu L, Ren X, Chen J, Yin X, et al. HDAC1/2/3 are major histone desuccinylases critical for promoter desuccinylation. Cell Discov. 2023;9:85–101. https://doi.org/10.1038/s41421-023-00573-9
Federico M, Bagella L. Histone deacetylase inhibitors in the treatment of hematological malignancies and solid tumors. J Biomed Biotechnol. 2011;2011:1–12. https://doi.org/10.1155/2011/475641
Guerrant W, Mwakwari SC, Chen PC, Khan SI, Tekwani BL, Oyelere AKJC. A structure–activity relationship study of the antimalarial and antileishmanial activities of nonpeptide macrocyclic histone deacetylase inhibitors. 2010;5:1232-5. https://doi.org/10.1002/cmdc.201000087.
Cheshmazar N, Hamzeh-Mivehroud M, Nozad Charoudeh H, Hemmati S, Melesina J, Dastmal-chi S. Current trends in development of HDAC-based chemotherapeutics. Life Sci. 2022;308. https://doi.org/10.1016/j.lfs.2022.120946.
Marks PA, Breslow R. Dimethyl sulfoxide to vorinostat: development of this histone deacetylase inhibitor as an anticancer drug. Nat Biotechnol. 2007;25:84–90. https://doi.org/10.1038/nbt1272
Sawas A, Radeski D, O’Connor OA. Belinostat in patients with refractory or relapsed peripheral T-cell lymphoma: a perspective review. Ther Adv Hematol. 2015;6:202–8. https://doi.org/10.1177/2040620715592567
Sharma S, Bailey H, Stenehjem D. Panobinostat for the treatment of multiple myeloma: the evidence to date. J Blood Med. 2015;6:269–76. https://doi.org/10.2147/jbm.S69140
Reddy SA. Romidepsin for the treatment of relapsed/refractory cutaneous T-cell lymphoma (mycosis fungoides/Sézary syndrome): Use in a community setting. Crit Rev Oncol Hematol. 2016;106:99–107. https://doi.org/10.1016/j.critrevonc.2016.07.001
Ning ZQ, Li ZB, Newman MJ, Shan S, Wang XH, Pan DS, et al. Chidamide (CS055/HBI-8000): a new histone deacetylase inhibitor of the benzamide class with antitumor activity and the ability to enhance immune cell-mediated tumor cell cytotoxicity. Cancer Chemother Pharmacol. 2011;69:901–9. https://doi.org/10.1007/s00280-011-1766-x
Han H, Feng X, He T, Wu Y, He T, Yue Z, et al. Discussion on structure classification and regulation function of histone deacetylase and their inhibitor. Chem Biol Drug Des. 2023;103:14366–84. https://doi.org/10.1111/cbdd.14366
Yu F, Ran J, Zhou J. Ciliopathies: does HDAC6 represent a new therapeutic target? Trends Pharmacol Sci. 2016;37:114–9. https://doi.org/10.1016/j.tips.2015.11.002
San JE, Gimenez C, Agirre X, Prosper F. HDAC inhibitors in acute myeloid leukemia. Cancers. 2019;11:1794–17. https://doi.org/10.3390/cancers11111794
Thomas M, Clarhaut J, Tranoy-Opalinski I, Gesson JP, Roche J, Papot S. Synthesis and biological evaluation of glucuronide prodrugs of the histone deacetylase inhibitor CI-994 for application in selective cancer chemotherapy. Bioorg Med Chem. 2008;16:8109–16. https://doi.org/10.1016/j.bmc.2008.07.048
Jespersen H, Olofsson Bagge R, Ullenhag G, Carneiro A, Helgadottir H, Ljuslinder I, et al. Phase II multicenter open label study of pembrolizumab and entinostat in adult patients with metastatic uveal melanoma (PEMDAC study). Ann Oncol. 2019;19:415–21. https://doi.org/10.1093/annonc/mdz394.068
Huang R, Zhang X, Min Z, Shadia AS, Yang S, Liu X. MGCD0103 induces apoptosis and simultaneously increases the expression of NF‑κB and PD‑L1 in classical Hodgkin’s lymphoma. Exp Ther Med. 2018;16:3827–34. https://doi.org/10.3892/etm.2018.6677
Nakagawa-Saito Y, Saitoh S, Mitobe Y, Sugai A, Togashi K, Suzuki S, et al. HDAC Class I inhibitor domatinostat preferentially targets glioma stem cells over their differentiated progeny. Int J Mol Sci. 2022;23:8084–95. https://doi.org/10.3390/ijms23158084
Shinke G, Yamada D, Eguchi H, Iwagami Y, Asaoka T, Noda T, et al. Role of histone deacetylase 1 in distant metastasis of pancreatic ductal cancer. Cancer Sci. 2018;109:2520–31. https://doi.org/10.1111/cas.13700
Zhang RH, Guo HY, Deng H, Li J, Quan ZS. Piperazine skeleton in the structural modification of natural products: a review. J Enzyme Inhib Med Chem. 2021;36:1165–97. https://doi.org/10.1080/14756366.2021.1931861
Hatnapure GD, Keche AP, Rodge AH, Birajd-ar SS, Tale RH, Kamble VM. Synthesis and biological evaluation of novel piperazine derivatives of flavone as potent anti-inflammatory and antimicrobial agent. Bioorg Med Chem Lett. 2012;22:6385–90. https://doi.org/10.1016/j.bmcl.2012.08.071
Li R, Wu J, He Y, Hai L, Wu Y. Synthesis and in vitro evaluation of 12-(substituted aminomethyl) berberrubine derivatives as anti-diabetics. Bioorg Med Chem Lett. 2014;24:1762–5. https://doi.org/10.1016/j.bmcl.2014.02.032
Cao F, Zwinderman MRH, van Merkerk R, Ettema PE, Quax WJ, Dekker FJ. Inhibitory selectivity among class I HDACs has a major impact on inflammatory gene expression in macrophages. Eur J Med Chem. 2019;177:457–66. https://doi.org/10.1016/j.ejmech.2019.05.038
Ruzic D, Ellinger B, Djokovic N, Santibanez JF, Gul S, Beljkas M, et al. Discovery of 1-Benzhydryl-Piperazine-Based HDAC inhibitors with anti-breast cancer activity: synthesis, molecular modeling, in vitro and in vivo biological evaluation. Pharmaceutics. 2022;14:2600–22. https://doi.org/10.3390/pharmaceutics14122600
Trivedi P, Adhikari N, Amin SA, Bobde Y, Ganesh R, Jha T, et al. Design, synthesis, biological evaluation and molecular docking study of arylcarboxamido piperidine and piperazine-based hydroxamates as potential HDAC8 inhibitors with promising anticancer activity. Eur J Pharm Sci. 2019;138:105046–60. https://doi.org/10.1016/j.ejps.2019.105046
Li L, Mei DT, Zeng Y. HDAC2 promotes the migration and invasion of non-small cell lung cancer cells via upregulation of fibronectin. Biomed Pharmacother. 2016;84:284–90. https://doi.org/10.1016/j.biopha.2016.09.030
Acknowledgements
This work was supported by the National Science Foundation for Young Scientists of China to Yepeng Luan (NSFC No.81602947).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cai, C., Luan, Y. Design, synthesis and antitumor activity evaluation of novel benzamide HDAC inhibitors. Med Chem Res (2024). https://doi.org/10.1007/s00044-024-03210-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00044-024-03210-6