Abstract
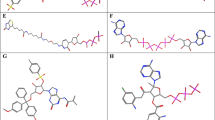
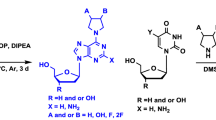
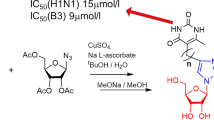
A comparative analysis of in vitro antiviral activity (in terms of the concentration of semi-maximal inhibition, IC50) against influenza virus A/PR/8/34 (H1N1) of a large series of parent 1,2,3-triazolyl nucleoside analogues (with uracil, thymine, 6-methyluracil, quinazoline-2,4-dione moieties as nucleic bases) and their prodrug forms with masked 5ʹ-phosphate groups (diethyl phosphate, diphenyl phosphate, phosphoramidate) and negatively charged H-phosphonate and monophosphate groups was carried out. Obtained structure-activity relationships were interpreted based on the assumption that the synthesized parent 1,2,3-triazolyl nucleoside analogues and their prodrug forms, by analogy with the literature data, are metabolized by cellular kinases to their active 5ʹ-triphosphate forms that inhibit the activity of viral RNA-dependent RNA polymerase (RdRp). A correlation was found between the experimental values of IC50 and the theoretical values of the binding energies of 5ʹ-triphosphate derivatives of the parent 1,2,3-triazolyl nucleoside analogues in the active site of RdRp.
Graphical Abstract







Similar content being viewed by others
References
Balzarini J. Metabolism and mechanism of antiretroviral action of purine and pyrimidine derivatives. Pharm World Sci. 1994;16:113–26. https://doi.org/10.1007/BF01880662
Pastuch-Gawołek G, Gillner D, Krol E, Walczak K, Wandzik I. Selected nucleos(t)ide-based prescribed drugs and their multi-target activity. Eur J Pharm. 2019;865:172747. https://doi.org/10.1016/j.ejphar.2019.172747
Eyer L, Nencka R, De Clercq E, Seley-Radtke K, Ruzek D. Nucleoside analogs as a rich source of antiviral agents active against arthropod-borne flaviviruses. Antiviral Chem Chemother. 2018;26:1–28. https://doi.org/10.1177/2040206618761299
Jordheim LP, Durantel D, Zoulim F, Dumontet C. Advances in the development of nucleoside and nucleotide analogues for cancer and viral diseases. Nat Rev Drug Discovery. 2013;12:447–64. https://doi.org/10.1038/nrd4010
Martin JC, Hitchcock MJM, Kaul S, Dunkle LM, Sterzycki RZ, Mansuri MM, et al. Comparative studies of 2′,3′-didehydro-2′3′-dideoxythymidine (d4T) with other pyrimidine nucleoside analogues. Ann NY Acad Sci. 1990;616:22–28. https://doi.org/10.1111/j.1749-6632.1990.tb17824.x
Gao WY, Agbaria R, Driscoll JS, Mitsuya H. Divergent anti-human immunodeficiency virus activity and anabolic phosphorylation of 2′,3′-dideoxynucleoside analogs in resting and activated human cells. J Biol Chem. 1994;269:12633–8. https://doi.org/10.1016/S0021-9258(18)99923-0
Stein DS, Moore KH. Phosphorylation of nucleoside analog antiretrovirals: a review for clinicians. Pharmacotherapy. 2001;21:11–34. https://doi.org/10.1592/phco.21.1.11.34439
Roy R, Depaix A, Périgaud C, Peyrottes S. Recent trends in nucleotide synthesis. Chem Rev. 2016;116:7854–97. https://doi.org/10.1021/acs.chemrev.6b00174
Kataev VE, Garifullin BF. Antiviral nucleoside analogs. Chem Heterocycl Comp. 2021;57:326–41. https://doi.org/10.1007/s10593-021-02912-8
Li Y, Mao S, Hager MW, Becnel KD, Schinazi RF, Liotta DC. Synthesis and evaluation of 2ʹ-substituted cyclobutyl nucleosides and nucleotides as potential anti-HIV agents. Bioorg Med Chem Lett. 2007;17:3398–401. https://doi.org/10.1016/j.bmcl.2007.03.094
Toti KS, Derudas M, Pertusati F, Sinnaeve D, Van den Broeck F, Margamuljana L, et al. Synthesis of an apionucleoside family and discovery of a prodrug with anti-HIV activity. J Org Chem. 2014;79:5097–112. https://doi.org/10.1021/jo500659e
Chien M, Anderson TK, Jockusch D, Tao C, Li X, Kumar S, et al. Nucleotide Analogues as Inhibitors of SARS-CoV-2 Polymerase, a Key Drug Target for COVID-19. J Proteome Res. 2020;19:4690–7. https://doi.org/10.1021/acs.jproteome.0c00392
Montgomery JA, Thomas HJ, Schaeffer HJ. Synthesis of potential anticancer agents. XXVIII. Simple esters of 6-mercaptopurine ribonucleotide. J Org Chem. 1961;26:1929–33. https://doi.org/10.1021/jo01065a058
Wagner CR, Iyer VV, McIntee EJ. Pronucleotides:toward the in vivo delivery of antiviral and anticancer nucleotides. Med Res Rev. 2000;20:417–51. https://doi.org/10.1002/1098-1128(200011)20:6<417::aid-med1>3.0.co;2-z
Cahard D, McGuigan C, Balzarini J. Aryloxy phosphoramidate triesters as Pro-Tides. Mini Rev Med Chem. 2004;4:371–81. https://doi.org/10.2174/1389557043403936.
Mehellou Y, Balzarini J, McGuigan C. Aryloxy phosphoramidate triesters: a technology for delivering monophosphorylated nucleosides and sugars into cells. ChemMedChem. 2009;4:1779–91. https://doi.org/10.1002/cmdc.200900289
Singh US, Mulamoottil VA, Chu CK. 2′-Fluoro-6′-methylene carbocyclic adenosine and its phosphoramidate prodrug: A novel anti-HBV agent, active against drug-resistant HBVmutants. Med Res Rev. 2018;38:977–1002. https://doi.org/10.1002/med.21490
McGuigan C, Madela K, Aljarah M, Gilles A, Brancale A, Zonta N, et al. Design, synthesis and evaluation of a novel double pro-drug: INX-08189. A new clinical candidate for hepatitis C virus. Bioorg Med Chem Lett. 2010;20:4850–4. https://doi.org/10.1016/j.bmcl.2010.06.094
Mehellou Y, Rattan HS, Balzarini J. The ProTide prodrug technology: from the concept to the clinic. J Med Chem. 2018;61:2211–26. https://doi.org/10.1021/acs.jmedchem.7b00734
Wang G, Lim SP, Chen YL, Hunziker J, Rao R, Gu F, et al. Structure-activity relationship of uridine-based nucleoside phosphoramidate prodrugs for inhibition of dengue virus RNA-dependent RNA polymerase. Bioorg Med Chem Lett. 2018;28:2324–7. https://doi.org/10.1016/j.bmcl.2018.04.069
Schooley RT, Carlin AF, Beadle JR, Valiaeva N, Zhang XQ, Clark AE, et al. Rethinking Remdesivir: synthesis, antiviral activity, and pharmacokinetics of oral lipid prodrugs. Antimicrob Agents Chemother. 2021;65:e01155–21. https://doi.org/10.1128/AAC.01155-21
Seley-Radtke KL, Yates MK. The evolution of nucleoside analogue antivirals: A review for chemists and non-chemists. Part 1: Early structural modifications to the nucleoside scaffold. Antivir Res. 2018;154:66–86. https://doi.org/10.1016/j.antiviral.2018.04.004
Slusarczyk M, Serpi M, Pertusati F. Phosphoramidates and phosphonamidates (ProTides) with antiviral activity. Antivir Chem Chemother. 2018;26:1–31. https://doi.org/10.1177/2040206618775243
Pruijssers AJ, Denison MR. Nucleoside analogues for the treatment of coronavirus infections. Curr Opinion Virology. 2019;35:57–62. https://doi.org/10.1016/j.coviro.2019.04.002
Tatarinov DA, Garifullin BF, Belenok MG, Andreeva OV, Strobykina IYU, Shepelina AV, et al. The first 5′-phosphorylated 1,2,3-triazolyl nucleoside analogues with uracil and quinazoline-2,4-dione moieties. Synthesis and antiviral evaluation. Molecules. 2022;27:6214. https://doi.org/10.3390/molecules27196214
Andreeva OV, Garifullin BF, Zarybaev VV, Slita AV, Yesaulkova IL, Saifina LF, et al. Synthesis and biological evaluation of 1,2,3-triazolyl nucleoside analogues as potential antiviral agents against influenza virus A (H1N1) and coxsackievirus B4. Mol Diversity. 2021;25:473–90. https://doi.org/10.1007/s11030-020-10141-y
Fan WQ, Katritzky AR 1,2,3-Triazoles. In: Katritzky AR, Rees CW, Scriven EFV, editors. Comprehensive Heterocyclic Chemistry. Oxford, UK; Pergamon; 1997. Vol. 4. p. 2-129.
Alexandrova LA, Efremenkova OV, Andronova VL, Galegov GA, Solyev PN, Karpenko IL, et al. 5-(4-Alkyl-1,2,3-triazol-1-yl)methyl derivatives of 2’-deoxyuridine as inhibitors of viral and bacterial growth. Russ J Bioorg Chem. 2016;42:677–84. https://doi.org/10.1134/S1068162016050022
Chatzileontiadou DSM, Parmenopoulou V, Manta S, Kantsadi AL, Kylindri P, Griniezaki M, et al. Triazole double-headed ribonucleosides as inhibitors of eosinophil derived neurotoxin. Bioorg Chem. 2015;63:152–65. https://doi.org/10.1016/j.bioorg.2015.10.007
Malin AA, Ostrovskii VA. Synthesis of thymidine derivatives as potential pharmaceuticals against HIV/AIDS infection. Russ J Org Chem. 2001;37:759–80. https://doi.org/10.1023/A:1012441026853
Kowalinski E, Zubieta C, Wolkerstorfer A, Szolar OHJ, Ruigrok RWH, Cusack S. Structural analysis of specific metal chelating inhibitor binding to the endonuclease domain of influenza pH1N1 (2009) polymerase. PloS Pathog. 2012;8:e1002831. https://doi.org/10.1371/journal.ppat.1002831
Jockusch S, Tao C, Li X, Anderson TK, Chien M, Kumar S, et al. A library of nucleotide analogues terminate RNA synthesis catalyzed by polymerases of coronaviruses that cause SARS and COVID-19. Antiviral Res. 2020;180:104857. https://doi.org/10.1016/j.antiviral.2020.104857
Jácome R, Campillo-Balderas JA, Ponce de León S, Becerra A, Lazcano A. Sofosbuvir as a potential alternative to treat the SARS-CoV-2 epidemic. Sci. Rep. 2020;10:9294. https://doi.org/10.1038/s41598-020-66440-9
Das K, Aramini JM, Ma LC, Krug RM, Arnold E. Structures of influenza A proteins and insights into antiviral drug targets. Nature SMB. 2010;17:530–8. https://doi.org/10.1038/nsmb.1779
Fudo S, Yamamoto N, Nukaga M, Odagiri T, Tashiro M, Hoshino T. Two distinctive binding modes of endonuclease inhibitors to the N‑terminal region of influenza virus polymerase acidic subunit. Biochemistry. 2016;55:2646–60. https://doi.org/10.1021/acs.biochem.5b01087
Lao J, Vanet A. A new strategy to reduce influenza escape: detecting therapeutic targets constituted of invariance groups. Viruses. 2017;9:38. https://doi.org/10.3390/v9030038
Luganini A, Terlizzi ME, Catucci G, Gilardi G, Maffei ME, Gribaudo G. The cranberry extract Oximacro® exerts in vitro virucidal activity against influenza virus by interfering with hemagglutinin. Front Microbiol. 2018;9:1826. https://doi.org/10.3389/fmicb.2018.01826.
Zhang W, Chen ST, He QY, Huang LQ, Li X, Lai XP, et al. Asprellcosides B of Ilex asprella Inhibits Influenza A virus infection by blocking the hemagglutinin- mediated membrane fusion. Front Microbiol. 2019;9:3325. https://doi.org/10.3389/fmicb.2018.03325
Gamblin SJ, Haire LF, Russell RJ, Stevens DJ, Xiao B, Ha Y, et al. The structure and receptor binding properties of the 1918 influenza hemagglutinin. Science. 2004;303:1838–43. https://doi.org/10.1126/science.1093155
Berman H, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, et al. The protein data bank. Nucleic Acids Res. 2000;28:235–42. https://doi.org/10.1093/nar/28.1.235
Benton DJ, Wharton SA, Martin SR, McCauley JW. Role of neuraminidase in influenza A(H7N9) virus receptor binding. J Virol. 2017;91:e02293–16. https://doi.org/10.1128/JVI.02293-16
Dou D, Revol R, Östbye H, Wang H, Daniels R. Influenza A virus cell entry, replication, virion assembly and movement. Front Immunol. 2018;9:1581. https://doi.org/10.3389/fimmu.2018.01581
D’Souza C, Kanyalkar M, Joshi M, Coutinho E, Srivastava S. Search for novel neuraminidase inhibitors: design, synthesis and interaction of oseltamivir derivatives with model membrane using docking, NMR and DSC methods. Biochim Biophys Acta Biomembr. 2009;1740:1740–51. https://doi.org/10.1016/j.bbamem.2009.04.014
Herlambang SJ, Saleh R. Molecular docking investigation for Indonesian H274Y mutant neuraminidase type 1 with neuraminidase inhibitors. Am J Mol Biol. 2012;2:49–59. https://doi.org/10.4236/ajmb.2012.21006
Le K, Tran D, Nguyen A, Le L. A screening of neuraminidase inhibition activities of isoquinolone alkaloids in Coptis chinensis using molecular docking and pharmacophore analysis. ACS Omega. 2020;5:30315–22. https://doi.org/10.1021/acsomega.0c04847
van der Vries E, Collins PJ, Vachieri SG, Xiong X, Liu J, Walker PA, et al. H1N1 2009 pandemic influenza virus: resistance of the I223R neuraminidase mutant explained by kinetic and structural analysis. PLoS Pathog. 2012;8:e1002914. https://doi.org/10.1371/journal.ppat.1002914
Trott O, Olson FJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem. 2010;31:455–61. https://doi.org/10.1002/jcc.21334
HyperChem Professional 8.0 (2007). Hypercube, Inc. http://www.hyper .com/?tabid=360. Accessed 26 November 2022.
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR. Open babel: an open chemical toolbox. J Cheminform. 2011;3:33. https://doi.org/10.1186/1758-2946-3-33
Acknowledgements
We are grateful to the Assigned Spectral-Analytical Centre of FRC Kazan Scientific Centre of RAS for technical assistance in research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Garifullin, B.F., Tatarinov, D.A., Andreeva, O.V. et al. Synthesis, antiviral evaluation, molecular docking study and cytotoxicity of 5′-phosphorylated 1,2,3-triazolyl nucleoside analogues with thymine and 6-methyl uracil moieties. Med Chem Res 32, 1770–1803 (2023). https://doi.org/10.1007/s00044-023-03112-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-023-03112-z