Abstract
The druggable genome is limited by structural features that can be targeted by small molecules in disease-relevant proteins. While orthosteric and allosteric protein modulators have been well studied, they are limited to antagonistic/agonistic functions. This approach to protein modulation leaves many disease-relevant proteins as undruggable targets. Recently, protein-protein interaction modulation has emerged as a promising therapeutic field for previously undruggable protein targets. Molecular glues and heterobifunctional degraders such as PROTACs can facilitate protein interactions and bring the proteasome into proximity to induce targeted protein degradation. In this review, we discuss the function and rational design of molecular glues, heterobifunctional degraders, and hydrophobic tag degraders. We also review historic and novel molecular glues and targets and discuss the challenges and opportunities in this new therapeutic field.
Similar content being viewed by others
Introduction
Every major cellular process relies on a complex and dynamic network of protein-protein interactions (PPIs). The dysregulation of these PPIs often leads to disease. For example, in cancer, abnormally expressed proteins interacting with their protein-binding partners can create a network of signaling pathways, driven by molecular signatures, that promote tumorigenesis, proliferation, invasion, and metastasis [1]. As such, the disruption or modulation of these cancer PPIs has emerged as a novel and promising therapeutic field.
The “druggable genome” or the proportion of the genome that can be targeted by small molecule drugs, is limited to disease-relevant proteins that have structural features that can be targeted by small molecules [2]. For decades, research has been focused on finding, designing, and optimizing small molecules that antagonize/agonize protein function. Under this approach, most protein inhibitors are small molecules that directly bind to the interaction surface of a protein, often a hydrophobic binding pocket or enzyme active site, sterically preventing binding to its substrates or interaction partner(s). Some protein targeted drugs bind to a surface region of a protein partner outside of the protein interaction interface itself, and antagonize protein function in an allosteric fashion [3]. However, many disease-relevant signaling mechanisms are carried out by interactions between proteins.
Modulating interactions between proteins can be difficult. For example, the contact surfaces involved in PPIs are large (~1500–3000 Å2) compared to those involved in protein-small molecule interactions (~300–1000 Å2). Additionally, the contact surfaces of PPIs are often flat and lack deep pockets or grooves present at the surfaces of proteins that typically bind to small molecules. Other challenges are the presence of noncontiguous amino acid binding sites, intrinsically disordered domains, and a general lack of natural ligands [4, 5]. For these reasons, some proteins have been deemed “undruggable” by small molecules. This umbrella of “undruggable” targets includes membrane-tethered scaffold proteins, transcription factors, splicing factors, non-enzyme proteins, steroid receptors, and cancer driver genes such as the RAS protein family, MYC, Hippo/YAP, STAT3, PAX3-FOXO, etc [6,7,8].
The challenge of tackling the undruggable proteome has inspired alternative technologies to modulate PPIs by inducing protein-protein stabilization and regulating the abundance of disease-inducing proteins by targeting protein degradation via induced proximity with a ubiquitin ligase [9]. In other words, this new approach allows for the stabilization of PPIs which may lead to deactivation or activation of protein activity, or to protein degradation instead of inhibition. Given these innovative approaches, proteins that were once deemed “undruggable” may be druggable after all.
Targeted protein degradation focuses on the development of small molecule degraders, including molecular glues and proteolysis-targeting chimeras (PROTACs). In this review, we will highlight the role of molecular glues in PPI stabilization, modulation, and protein degradation as well as heterobifunctional degraders and their role in protein degradation.
The ubiquitin-proteasome system (UPS)
Protein homeostasis, also known as proteostasis, is a protein quality control process by which the cell balances the expression of newly synthesized proteins and the degradation of damaged or misfolded proteins that are beyond repair [10]. The ubiquitin-proteasome system (UPS) is a proteostasis mechanism designed to degrade polyubiquitinated proteins via the proteasome.
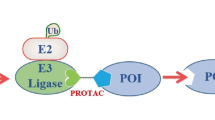
The to-be-degraded protein is targeted for proteolysis by ubiquitination, a process mediated by the E1-E2-E3 enzymatic cascade. Ubiquitin is a 76 amino acid protein that is covalently linked to lysine residues in protein substrates by an enzymatic reaction requiring ubiquitin activating (E1), ubiquitin conjugating (E2), and ubiquitin ligating (E3) enzymes acting sequentially [11]. The protein substrate becomes polyubiquitinated with at least four ubiquitin units and then is degraded by the 26S proteasome in eukaryotic cells [12]. The 26S proteasome complex is composed of a 20S catalytic subunit with 19S regulatory subunits on each end. The 20S catalytic subunit includes four ring structures and three proteolytic proteases [13]. Once a polyubiquitinated protein goes through proteolysis in the proteasome, it is cleaved into 3–25 amino acid peptides and free ubiquitin units [12, 14].
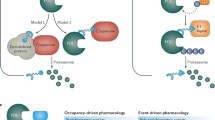
Targeted protein degradation by heterobifunctional small molecules is a strategy where the heterobifunctional molecule binds to a target protein and an E3 ligase simultaneously [15]. The induced spatial proximity can trigger a ubiquitin transfer from the E3 ligase to lysine residues on the target protein. Unlike typical small molecule inhibitors, which are controlled by occupancy for both kinetics of protein inhibition and duration of pharmacodynamics and efficacy, a heterobifunctional degrader allows for irreversible and catalytic protein degradation and oftentimes requires much lower concentrations than typical small molecule inhibitors [10].
Small molecule degraders
Molecular glues
Molecular glues are small molecule compounds (in the range of 500 Da or lower) that facilitate the interaction of proteins by bringing proteins into proximity. By inducing or “gluing” these interactions, molecular glues can stabilize and modulate PPIs [16]. In some cases, the protein-ligand-protein complexes serve to inhibit or activate the function of one or both proteins, while in others the complex stimulates degradation of one or both proteins [2]. For example, molecular glue degraders can facilitate or induce interactions between an E3 ubiquitin ligase and a target protein. Robust and compact interactions between protein-ligand and protein-protein interfaces result in the ubiquitination and subsequent degradation of the recruited protein [16]. Molecular glues can also restore or repair interactions that have become weak due to mutations or induce interactions between two proteins that are unlikely to interact on their own [17, 18]. Molecular glues, thus, can change the protein interactome by stabilizing protein interactions, repairing or restoring interactions, and inducing neo-protein interactions (Fig. 1) [19].
Molecular glues and molecular glue degraders and their mechanism of action. Molecular glues can A enhance and stabilize protein interactions, B repair protein interactions weakened by mutations or C induce de novo protein interactions. D Molecular glue degraders induce the interaction of an E3 ubiquitin ligase with a protein target. This triggers polyubiquitination of the target protein, which is degraded via the proteasome. E Molecular glues can also modulate protein function, with PKM2 an example. PKM2 is a pyruvate kinase relevant to cancer that is less active in a homodimer form. A molecular glue can stabilize the dimer and “glue” two dimers together forming a PKM2 homotetramer, which increases enzymatic activity
Molecular glues rely on the formation of ternary complexes to stabilize/induce interactions between proteins. A molecular glue-induced ternary complex usually refers to a protein complex composed of an E3 ubiquitin ligase, the small molecule degrader, and the target protein [20]. For a protein to be successfully polyubiquitinated, the ternary complex formation must be productive. This means that the ternary complex must serve as a scaffold between the target protein and a ubiquitin-conjugating enzyme (E2) for it to transfer ubiquitin to a surface lysine at the target protein. Predicting productive ternary complex formation remains a challenge when designing molecular glues [21].
Molecular glues are particularly attractive for drug discovery due to their lower molecular weight and more drug-like properties compared to large molecular entities such as PROTACs [19, 22]. An advantage in using molecular glues is that they have a unique mechanism of action that can either induce degradation of the target protein or stabilize PPIs [19]. Despite their desirable drug-like properties, molecular glues remain difficult to identify and even harder to design in a rational manner. In fact, most molecular glues have been found serendipitously [9].
General principles of rational discovery
Historically, small molecule drug discovery screening programs have relied upon initial biochemical assays to assess activity and rank order compounds. However, this has been a challenge for molecular glues because their activity results in stabilized PPIs or targeted protein degradation, which is dependent on the productive assembly of ternary complexes that can trigger a cascade of cellular events [16]. Currently, the availability to screen technologies to monitor PPIs or target protein degradation in rational and a high-throughput fashion in the context of the cellular environment is still lacking. A comprehensive review about multi-omics approaches to identify molecular glues was recently published by the Georg Winter lab [23]. Here, we present examples of successful molecular glue discovery through a variety of methods (Table 1).
Library and high-throughput screening
Recently, novel methods have been developed to identify molecular glues using library screening. For example, Li et al. developed a screening microarray for the discovery of molecular glues that trigger degradation via the autophagosome. Lie et al. screened 3375 small molecules and found four small molecules that facilitated interactions between the autophagosome LC3 protein (microtubule-associated protein A1/1B light chain 3) and the mutated form of the huntingtin protein (HTT), which causes Huntington’s disease. The four molecular glues interacted with both the wild type and mutant HTT protein. Upon further design, the optimized molecular glues were able to selectively degrade mutant HTT via the autophagosome at nanomolar concentrations in vitro and at 0.5 mg kg−1 in vivo in a Huntington’s Disease mouse model. The molecular glues did not increase the number of autophagosomes or alter autophagosome-lysosome fusion [24].
Another example of library screen for molecular glues involves a yeast two-hybrid approach. Andrea Chini described a chemogenomic screen method using yeast two-hybrid to identify molecular glues that modulate plant hormone receptor complexes [25]. Jasmonic acid (JA) and derivatives, also known as jasmonates, are hormones that modulate plant immunity and development such as root growth and male and female fertility [26]. The JA signaling pathway has three major molecular components involved in JA-responses: the coronatine insensitive 1 protein (COI1), the transcriptional repressor jasmonate-ZIM domain (JAZ) family, and transcription factors that regulate expression of JA-responsive genes. COI1 is an F-box protein that forms a functional E3 ligase that targets and ubiquitinates JAZ proteins. Once transcriptional repressor JAZ proteins have been degraded, JA-responsive genes are transcribed [26,27,28]. In order to screen compounds to disrupt this process, the plant hormonal perception complex COI1-JAZ9-JA-Ile was reconstructed in yeast to identify novel compounds with JA-Ile (a conjugate form of JA) agonist or antagonist activities. This method was applied to screen three chemical libraries containing ~22,500 molecules total. The goal was to identify compounds that could interfere with COI1 interactions with JAZ proteins [29]. To identify JA-Ile agonist compounds, yeast cells co-transformed with pGBK-COI1/pGAD-JAZ9 were grown on minimal yeast media in the presence of compounds from three chemical libraries. In theory, compounds that promote COI1-JAZ9 interactions are identified due to enhanced yeast growth and compounds that interfere with COI1-JAZ interactions inhibit yeast growth in the presence of coronatine. When they screened the libraries, they did not find any agonist hit compounds but found five hit antagonist compounds. Additionally, the antagonist hit compound were confirmed by secondary screening in an in planta assay where 35S-JAZI-GUS seeds were germinated for a few days to allow for root growth and then exposed to JA and candidate antagonist compounds. A true antagonist hit compound would prevent seed root decay in the presence of JA [25, 28]. Through this secondary screening, they confirmed that the five compounds screened through yeast two-hybrid disrupted COI1-JAZ9 interactions [29]. Although this method is meant to be used in plants, it can be used in other applications in screening for compounds that facilitate interactions involving receptor complexes, if a suitable yeast two-hybrid library is available. A limitation to this assay is that yeast two-hybrid is notorious for generating false positives so a second orthogonal assay in planta or in vitro and pull-down confirmation assays are required.
Multicovalent natural products
A rational design to identify molecular glues from electrophilic natural products was proposed by Isobe et al. [30]. Based on previous data describing natural products as a robust source of many known molecular glue interactions such as rapamycin, auxin, and brefeldin A, Isobe et al. explored electrophilic natural products since they represent an underexplored subset of natural products where molecular glues could be found. One of the unique characteristics about electrophilic natural products is their potential for covalent bond formation due to their multiple reactive sites that could produce ternary complexes for PPIs and protein degradation. Two fungal polyketide natural products were explored: asukamycin and manumycin A, which exhibit antibiotic and antiproliferative properties. Both natural products possess three potentially electrophilic functional groups with the additional possibility of regioisomeric nucleophilic attack (Fig. 2A). Targets of the covalent ligand asukamycin were explored in 231MPF cells by isotopic tandem orthogonal proteolysis activity-based protein profiling. Briefly, vehicle or asukamycin-treated samples were labeled with either IA-alkyne or N-hex-5-ynyl-2-iodoacetamide. Samples were then sequentially digested with trypsin and TEV and prepared for liquid chromatography-mass spectrometry. The validated method can be found here [31, 32]. Amino acid C374 of UBR7, a putative E3 ubiquitin ligase, was identified as the primary target of asukamycin and was validated by gel based ABPP. The UBR7-asukamycin interaction led to a gain of function as opposed to simple inhibition. Using proteomic analysis of pull-down eluates of UBR7-expressing 231MPF cells, 13 interacting proteins were identified, including DNA protein kinase (PRKDC) and TP53. Furthermore, the UBR7-asukamycin complex thermodynamically stabilized TP53 and increased its transcriptional activity resulting in antiproliferative effects in 231MPF cells [30, 33]. However, the authors were not able to identify the binding mode of UBR7-asukamycin to TP53 and proposed this could be clarified by solving the crystal structure of the ternary of UBR7-asukamycin-TP53 complex. This method could be used to explore new molecular glues from natural products with multiple, distal electrophilic sites such as isariotins and abikoviromycin [34,35,36]. Given their multicovalent nature, screening similar natural products could lead to the discovery of multiple molecular glue targets.
Chemogenomic screening
Chemogenomics is the study of gene products using small molecule pharmacological modulators [37]. Chemogenomic libraries, comprised of selective annotated small molecules, are used in phenotypic screens, where target deconvolution is driven by the known pharmacological targets of the hits. Chemogenomic screens have demonstrated a significant impact in uncovering new biology. However, further experiments to validate the target modulation and mechanism of action are almost always necessary [38].
One recent example of successful chemogenomic screening is the target identification of molecular glue indisulam, RBM39. Indisulam facilitates the interaction of RBM39 with the DCAF15-CUL4-CRL E3 complex, which causes RBM39 polyubiquitination and subsequent degradation [39, 40]. The increased understanding of E3 ligase biology through the story of indisulam inspired chemogenomic screening for previously undruggable targets. For example, Simonetta et al. conducted a screen aimed at identifying small molecules with glue activity that could enhance the interaction between β-catenin and the natural E3 ligase β-TrCP, subsequently mediating the degradation of mutant β-catenin present in cancers [17].
Another example are the multiple chemogenomic screening approaches to discover cyclin K molecular glue degraders Słabicki et al. used a systemic data mining screening approach analyzing the correlations between an antitumor drug sensitivity database of 4518 compounds against 578 cancer cell lines and the mRNA expression level of 499 E3 ligase compounds [41]. From this screening, they discovered that the mRNA level of DDB1, a CUL4 adapter protein, correlated with the cytotoxicity of the CDK inhibitor (R)-CR8 (Fig. 3). Subsequent quantitative proteome-wide mass spectrometry to evaluate protein abundance after treating cells with (R)-CR8 showed that (R)-CR8 had targeted degradation activity toward cyclin K. The crystal structure revealed that CR8 binds to the ATP pocket site of CDK12 inducing an interaction between CDK12 and DDB1 through its phenylpyridine group. The ternary DDB1-(R)-CR8-CDK12 complex acts as a recruiter of cyclin K to DDB1, which leads to its polyubiquitination and degradation [41].
Mayor-Ruiz et al. used a scalable strategy toward glue degrader discovery by comparative chemical screening in hyponeddylated cellular models with broadly abrogated ligase activity [42]. They based this approach on the knowledge that cullin-RING ligase activity is dependent on the reversible attachment of the ubiquitin-like protein NEDD8 to the respective cullin scaffold, a process that is managed by NAE (an E1 enzyme), UBE2M and UBE2F (E2 enzymes), and several E3 enzymes. Their hypothesis was that comparing the chemical profiles in hyponeddylated vs. neddylation-proficient cells would allow to screen for small molecules that are functionally linked to cullin-RING ligases. They screened a library of ~2000 small molecules in wild type and UB2M mutant cells that yielded four chemical scaffolds dCeMM1, dCeMM2, dCeMM3, and dCeMM4 that were functionally dependent on uninterrupted neddylation levels (Fig. 3). They identified dCeMM1 as a new molecular glue that rewired CRL4 DCAF15 ligase, capable of degrading RBM39. dCeMM2, dCeMM3, and dCeMM4 were found to be cyclin K degraders and mild CDK13 and CDK13 degraders mediated by the CRL4B ligase complex [42]. The authors did not address the therapeutic value of these compounds or their pharmacological activities.
Lv et al. also discovered a molecular glue, HQ-461 (Fig. 3), that binds to CDK12’s kinase domain, creating a modified CDK12 version that becomes a substrate specific receptor for DDB1-CUL4-RBX1 E3 ubiquitin ligase and triggers polyubiquitination of cyclin K via the proteasome [43]. HQ-461 was discovered by phenotype-based high-throughput screening of small molecules looking for NRF2 activity suppression but its role as molecular glue was elucidated by chemical genetics and biochemical reconstitution [43]. For the chemical genetics approach, a gain of function screening was performed in HCT-116 cells, which are defective in mismatch repair and therefore have a high rate of random point mutation. HQ-461 resistant HCT-116 cells were generated and whole-exon sequencing in both the HQ-461 sensitive and resistant HCT-116 cells identified a variant G731E CDK12 gene in the resistant cell line. Further experiments confirmed CDK12 as the target of HQ-461 [43].
Phenotypic screens and chemical proteomics
Another CDK12 molecular glue was recently discovered by phenotypic screening in 3D patient-derived tumor models [44]. The authors cultured patient-derived colorectal cancer tumor spheroids and performed high-throughput screening of a ~80,000 non-characterized small molecule compound library. From the library hits, compounds of interest were narrowed to those that showed inhibitory activity in only a subset of patient-derived tumor spheroid cultures to minimize the risk of non-specific toxicity. From that criterion and due to a pronounced activity in tumor spheroid cultures and no activity in primary fibroblasts, NCT02 was selected (Fig. 3). NCT02 induced apoptosis and arrested the cell cycle in tumor spheroid cultures. To identify a potential target of NCT02, the authors performed thermal proteome profiling, which revealed CDK12 had the strongest and most significant shift in thermal stability. NCT02 destabilized the CDK12-cyclin K complex and mass spectrometry showed that cyclin K was only identified in cells treated with inactive NCT02 or DMSO but not NCT02. Further chemical proteomic analysis showed that NCT02 triggers degradation of both cyclin K and CDK12 via the proteasome and that CDK12 degradation occurs after cyclin K degradation. To test whether NCT02 acted as a molecular glue, co-immunoprecipitation experiments were performed with DDB1 in the presence of cyclin K, the kinase domain of CDK12, and multiple CDK12 inhibitors such as THZ-531 (non-degrading), SR-485 (cyclin K degrading), CR8 (molecular glue to CDK12 and cyclin K degrader), and NCT02 acted in a similar way to CR8. Structure-based docking revealed that NCT02 acts a molecular glue by mediating the interaction between cyclin K and DDB1 [44].
Heterobifunctional degraders
PROTACs
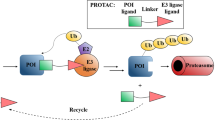
Proteolysis-targeting chimeras or PROTACs are heterobifunctional small molecules whereby one end of the molecule recruits an E3 ubiquitin ligase while the other end engages the target protein (Fig. 4A). These molecules were developed under different names and are sometimes referred as Specific and Non-genetic inhibitors of apoptosis protein [IAP]-dependent Protein Erasers or SNIPERs [45]. A PROTAC consists of three elements: an E3 ubiquitin ligase ligand, a protein of interest (POI) ligand, and a linker [8]. The E3 ubiquitin ligase ligand recruits a E3 ubiquitin ligase, the POI ligand targets and hijacks the POI, and the linker conjugates these two ligands. Once a stable ternary complex forms between the POI and the E3 ubiquitin ligase, ubiquitin-mediated proteolysis is triggered, where the E3 ligase recognizes the POI and covalently attaches ubiquitin onto it, leading to subsequent degradation by the proteasome [46]. In contrast to inhibitor-based pharmacology, PROTACs require only transient drug-target binding to rewire the ubiquitin-proteasome pathway to induce ubiquitination and degradation of target proteins [47]. As such, PROTACs have emerged as a novel therapeutic approach to the challenging multidomain proteins or “undruggable” proteins.
Structure and mechanism of action of heterobifunctional degraders. A PROTAC. Proteolysis-targeting chimeras (PROTACs) are comprised of a protein ligand domain and an E3 ligand domain joined by a linker. Once the PROTAC links the protein of interest to an E3 ligase, the protein of interest is ubiquitinated and degraded via the proteasome. B LYTAC. Lysosomal targeted chimeras (LYTACs) have an antibody domain and an oligoglycopeptide domain. The antibody binds to extracellular or transmembrane proteins and the oligoglycopeptide domain binds to cell surface receptors that trigger endocytosis. The proteins are degraded via the lysosome. C AUTAC. Autophagy-targeting chimeras (AUTACs) are composed of a protein ligand domain and a guanine tag joined by a linker. The AUTAC binds to the protein domain and the guanine tag mimics S-guanylation, a post translational modification that triggers autophagy. The protein is then degraded via the autophagosome. D ATTEC. Autophagosome-tethering compounds (ATTECs) have a protein or lipid ligand domain and an L3 ligand domain joined by a linker. The protein or lipid ligand domain binds to the protein or lipid of interest and the L3 domain binds to the L3 protein in the phagophore (autophagosome precursor). The protein or lipid is then enclosed into the autophagosome and degraded. E HaloPROTAC. HaloPROTACs are small molecules that induce ubiquitination of proteins tagged with HaloTag7 fusion proteins. The HaloPROTAC has a chloroalkane reside that can covalently conjugate to the HaloTag and an E3 ligand domain that can bring VHL or cIAP E3 ligases into proximity. F AbTAC. Antibody-based PROTACs are fully recombinant bispecific antibodies (IgG) that can simultaneously bind a transmembrane E3 ligase and a membrane-bound protein. The E3-AbTEC-protein complex triggers endocytosis and the protein is degraded via the lysosome
Crews and Deshaies reported the first PROTAC in 2001. This PROTAC was peptide based and recruited MetAP2 to SCF, a ubiquitin ligase complex [48]. In 2008, Schneekloth et al. reported the first small molecule PROTAC in 2008, which degraded the androgen receptor through the recruitment of the E3 ligase MDM2 [49]. Recently, PROTACs have been optimized for lower molecular weights to increase cell permeability. Some PROTACs have been designed to degrade undruggable targets such as estrogen-related receptor alpha, androgen receptor, cellular retinoic acid binding proteins, bromodomain proteins and multiple tyrosine kinases [50,51,52,53,54,55]. Currently there are three clinical trials in phase 1 or 1/2 involving PROTACs targeting the androgen receptor (drug ARV-110, NCT03888612), the estrogen receptor (drug ARV-471, NCT04072952), and Bruton Tyrosine Kinase (BGB-16673, NCT05006716).
Advantages to using PROTACs vs. traditional small molecule inhibitors are the ability to target undruggable targets,, avoid drug resistance by point mutations in the POI, gain target specificity, increased potency through the catalytic MOA, remove all functions of the target protein through degradation, and prolonged pharmacodynamic effects [56]. Some limitations of PROTACs are higher molecular weights that can restrict cellular permeability, the need for a stable ternary complex formation, and the “hook effect” that can inhibit ternary complex formation at high concentrations.
Rational design of PROTACs
Unlike molecular glues, PROTACs are, in theory, much easier to design rationally due to their modular structure. Theoretically, a new PROTAC can be developed by replacing its “warhead” or target ligand domain [16, 45]. Since the target ligand does not need to be a potent inhibitor of the target protein, a nonselective or promiscuous ligand can have the potential to be converted to a potent degrader when incorporated into a PROTAC [57, 58]. Bondeson et al. found proof of concept of this by designing CRBN- and VHL-recruiting PROTACs using a promiscuous kinase inhibitor with the potential to degrade over 100 substrate proteins. They found that the warhead affinity to the target ligand did not necessarily predict successful degradation. The factor that best correlated with degradation potency was the ability of that protein to induce a stable ternary complex with the PROTAC and the E3 ligase [59].
The choice of which E3 ligase to recruit matters in PROTAC design. There are hundreds of E3 ligases in cells but only a handful have been used for targeted protein degradation [45]. Many groups have shown that using a different E3 ligases to the same target protein results in different degradation potencies [54, 59,60,61,62,63]. This again suggests that stability of the ternary complex is important for targeted degradation. Exploring E3 ligases for recruitment may be worthwhile for a variety of reasons. For example, some E3 ligases are tissue specific or tumor specific [64,65,66,67] which could restrict degradation of target proteins to a tissue type or a tumor site and increase selective toxicity. Moreover, resistance to PROTAC induced targeted degradation has been recently reported [68]. The resistance mechanism lies in an alteration of the UPS pathway rather than the target proteins. Recruiting novel E3 ligases could restore degradation of the target proteins [68].
The PROTAC linker length and attachment point can also affect stability of the ternary complex formation and selectivity and degradation profiles can be affected [69]. In 2017, the structure solution of the BET bromodomain degrader MZ1 in complex with BRD4BD2 and VBC (VHL:ElonginB:ElonginC) showed that the linker coiled around itself, aiding in the formation of PPIs that resulted in a stable ternary complex [57]. Another study demonstrated that PROTACs with the same protein ligand and E3 recruiting ligand offer different selectivity profiles depending on the linker chemistry. Furthermore, an increase in linker length of 3 atoms changed the degradation profile of a lapatinib-based PROTAC from targeting both EGFR and HER2 into one that only degrades EGFR [70].
Screening for PROTAC mediated degradation in a high-throughput fashion has been a challenge. Traditional approaches for quantifying protein level changes, such as Western blots, are typically low throughput with limited quantification. Recently, several approaches for high-throughput screening of PROTACs have been published that are noteworthy. For example, the utilization of native mass spectrometry to study ternary structures. Sternicki et al. used native mass spectrometry at high resolution to measure ternary complex formation as a function of PROTAC concentration using the PROTAC GNE‐987 between Brd4 bromodomains 1 and 2 and VHL [71]. This method was used to measure complex stability and affinity in both the ternary complex and other intermediate protein species in a high-throughput fashion [71]. Beveridge et al. also utilized native mass spectrometry to study binary and ternary complex formation that specifically revealed preferentially formed E3-PROTAC-protein of interest (POI) combinations and this method could also be used for high-throughput screening for the development of PROTACs [72]. Simard et al. recently published a review of high-throughput screening of PROTACs that included high-throughput flow cytometry, in-cell based Western blotting, AlphaLISA, TR-FRET assays, and Nano-Glo HiBiT technology, all of which are promising to speed up the development and optimization of PROTACs [73].
Designing PROTACs with desirable pharmacokinetics remains a big challenge. Additionally, PROTACs often do not conform to Lipinski’s “rule of five” due to their high molecular weight, which restricts their cellular absorbance and other drug-like properties [74]. For these reasons, many doubted that PROTACs could ever be anything other than tool compounds. Against the odds, in 2019, ARV-110 and ARV-471, PROTACs targeting the androgen and estrogen receptor, respectively, were the first PROTACs to enter clinical trials (NCT03888612, NCT04072952). Both PROTACs are now in phase II clinical trials. Since then, multiple other PROTACs have entered the clinic for multiple indications (NCT04886622, NCT04830137, NCT05006716, NCT04772885, NCT04428788). Moreover, a database with the aim to promote PROTAC rational design is now available with open source structural and experimental data related to warheads, linkers, and E3 ligands, along with their biological activity and physiochemical properties [74].
LYTAC
Lysosome targeting chimeras (LYTACs) are bifunctional degraders that carry an oligoglycopeptide on one end, an antibody or small molecule on the other end, with both regions joined by a chemical linker (Fig. 4B) [75, 76]. The purpose of a LYTAC is to deliver a target protein to the lysosome, where it will be degraded, while the LYTAC is recycled. Unlike the ubiquitin-proteasomal pathway, the lysosomal pathway for protein degradation is not limited to proteins with intracellular domains, therefore, LYTACs can induce degradation of secreted, extracellular, and membrane-bound proteins [77].
Banik et al. reported the first LYTAC in 2020 with an oligoglycopeptide group that binds to the cell surface receptor CI-M6PR and an antibody that binds to a transmembrane or extracellular protein. The resulting complex is engulfed by the cell membrane forming a transport vesicle that carries the complex to the lysosome. The protein is then degraded, and the receptor is recycled. The authors could not determine if the LYTAC was also degraded [78].
AUTAC
Autophagy-targeting chimeras (AUTACs) are small molecule protein degraders that induce autophagy. Autophagy is an important cellular process that can degrade not only proteins but dysfunctional organelles and intracellular pathogens [79]. AUTACs are designed to link a specific ligand to a guanine tag (Fig. 4C). This event signals for selective autophagy of the substrate. AUTACs have a therapeutic component in the sense that they can degrade both cytosolic and membrane-bound proteins, which can mediate clearance of disease-related proteins and debris. In 2019, Takahashi et al. described AUTACs as cargo-specific degraders that used autophagy as the degradation pathway. The authors found that S-guanylation of a protein was sufficient to designate it for autophagy. Furthermore, AUTACs can degrade fragmented mitochondria and promote mitochondria turnover, which is beneficial as it enhances mitochondria quality [80].
ATTECs
ATTECs, autophagy-tethering compounds, are a novel class of bifunctional molecules that can hijack the autophagosomal pathway for the potential degradation of other cellular components in addition to proteins. The mechanism of action of ATTECs is to recruit LC3, a lipidated protein found on the membranes of autophagosomes, and direct a target of interest for autophagy without a guanine tag, like with AUTACs (Fig. 4D) [81]. Although ATTECs seem an attractive technology for targeted degradation, ATTECs are limited in the breadth of targets and are selective only for proteins with polyarginine groups. Multiple biophysical/structural studies would be required to establish the LC3-binding moiety to create ATTECs that induce degradation of other proteins [79].
Fu et al. developed a proof-of-concept molecule from ATTECs called LD-ATTECS, designed to clear lipid droplets (LD), which could be helpful in fighting chronic diseases such as obesity, cardiovascular disease, cancer, etc. This is one of the few examples of targeted degradation of non-protein targets [82]. LD-ATTECs were designed by linking LC3-binding molecules with LD detection probes and tested in mouse embryonic fibroblasts, where near complete clearance of oleic acid was observed. Degradation of large LD was also observed in differentiated adipocytes treated with LD-ATTECs. In mouse models, treatment with LD-ATTECs reduced whole-body weight, body fat:lean ratio, liver weight, liver LDs, and serum TAG and cholesterol levels [82]. Further studies are warranted to confirm the potential therapeutic benefits of these degraders.
HaloPROTAC
HaloPROTAC are PROTACs molecules designed to degrade HaloTag7 fusion proteins [83]. HaloTag is a modified bacterial dehalogenase that reacts covalently with hexyl chloride tags. HaloTag fusion proteins have been used to bioorthogonally label proteins in vivo. HaloPROTACs have two motifs and a linker: one motif binds to the to the HaloTag7 fusion protein and the other motif binds to an E3 ligase such as VHL or cIAP1 (Fig. 4E) [83, 84]. The linker length is crucial for the HaloPROTAC to be able to induce degradation, and it was reporter that lack of long peptidic chains in linker lengths deem HaloPROTACs more effective [83]. This provides a higher advantage for the usage of HaloPROTACs because in general they are smaller in size and possess drug-like properties. A potential limitation of HaloPROTACs is that stoichiometric occupancy of the tagged protein is needed to achieve full degradation of the target protein [85].
AbTACs
AbTACs are fully recombinant bispecific antibodies that recruit membrane-bound E3 ligases for the degradation of cell-surface proteins [86]. Bispecific antibodies refer to a family of immunoglobulins that bind to two different epitopes co-localizing their proteins [87]. By designing one of the binding-epitopes to be one of a protein of interest and the other to be a membrane-bound E3 ligase, an AbTAC could target protein degradation for membrane-bound proteins (Fig. 4F) [88].
Proof of concept for AbTACs was recently reported by the Wells lab in the development of an AbTAC with bifunctionality for PD-L1 (atezolizumab) and RNF43 (transmembrane E3 ligase) [86]. The AbTAC, AC-1, induced PD-L1 degradation in MDA-MB-24 cells via the lysosome, although the endocytosis mechanism is unclear. AC-1 demonstrated partial degradation of PD-L1 in cell lines derived from clinically relevant indications for atezolizumab [86].
Hydrophobic tag-mediated degraders
Hydrophobic tagging is another approach for targeted protein degradation. Hydrophobic tag-mediated degraders link a protein of interest ligand with a highly lipophilic moiety that triggers the unfolded protein response, a cellular stress response that is activated by high levels of misfolded or unfolded proteins in the endoplasmic reticulum. This response will then trigger degradation of the protein of interest [22].
This concept of hydrophobic tagging was first grasped with fulvestrant, also known as Faslodex, 7α-alkylsulphinyl analog of 17β-estradiol (Fig. 5A) [89]. Fulvestrant binds to the ER in a conformation that exposes the hydrophobic tag region. This conformation inhibits receptor dimerization, blocks nuclear localization, and renders the receptor inactive. Moreover, the fulvestrant-ER complex is unstable, which triggers the unfolded protein response and subsequent degradation via the proteasome [90]. This results in full anti-estrogen effects due to halting all transcriptional ER activity (Fig. 5B) [91]. Fulvestrant is currently approved for use alone and in combination with palbociclib, ribociclib, and abemaciclib to treat hormone receptor positive advanced breast cancer and is undergoing additional clinical trials to be used in combination with other drugs for breast cancer treatment [92,93,94,95,96].
An example the mechanism of action of hydrophobic tags. A Estradiol and analog fulvestrant structures. Fulvestrant functions as a hydrophobic tag to the estrogen receptor (ER). B Fulvestrant competitively binds to the ER inhibiting its dimerization and nuclear localization. The fulvestrant-ER complex is unstable, inducing unfolding and exposure of hydrophobic residues which triggers the unfolded protein response. The fulvestrant-ER complex is then degraded via the proteasome
Historical examples of molecular glues
Molecular glue origins
The field of molecular glues began when a series of natural products were found to have immunosuppressive and cell growth inhibitory effects and yet an unknown mechanism of action. This was the case for cyclosporin A and FK506, also known as tacrolimus. Both drugs are macrocyclic molecules used in the clinic for their ability to prevent rejection following organ transplant by inhibiting T cell activation [97]. Schreiber and Crabtree described their unknown mechanism of action in 1992, where they pointed out that cyclosporin A and FK506 bound to endogenous intracellular receptors, a termed they coined as “the immunophilins”, and the resulting complex targets the protein phosphatase, calcineurin, to exert an immunosuppressive effect [98]. In other words, these “molecular glues” simultaneously bind two proteins and inhibit the function of one of the proteins.
Another example this mechanism of action was demonstrated with another macrolide called rapamycin, which was shown to have potent immunosuppressive activity in IL-2 stimulated T cells by cell cycle arrest. Rapamycin binds to immunophilin FKBP12 forming a unique effector molecular complex, which then interacts with mammalian target of rapamycin or mTOR [99, 100]. The downstream effects of this interaction include inhibition of translational pathways and ribosomal biogenesis, which leads to overall translation blockade and cell cycle arrest [101]. Later it was discovered that mTOR was a kinase involved in many cellular pathways such as immune cell differentiation, apoptosis, autophagy, and tumor metabolism [102]. As such, rapamycin and derivatives, acting as molecular glues, modulate mTOR’s multiple functions.
One of the first biological indications that molecular glues could induce protein degradation came from plant hormones. Auxin and methyl jasmonate, both phytohormones, were found to bind the plant F-box CRL receptors (RIR1 and COI1, respectively) and facilitate interaction and degradation of two transcription factors [16]. Auxin-RIP1 and methyl jasmonate-COI1 crystal structures revealed that these phytohormones facilitate nanomolar target-ligase interactions through a small protein-ligand interface [17, 103].
Thalidomide and analogs (IMiDs)
In 1957, thalidomide was introduced to the West German market as a completely non-toxic sedative that could be used safely, even during pregnancy. After many babies were born with phocomelia or abnormally short limb outgrowth, thalidomide was withdrawn from market in 1962 [104]. Not long after its market withdrawal, a single case report was published describing thalidomide’s anti-inflammatory properties in which a patient suffering from Hansen’s disease, commonly known as leprosy, had complete resolution of their inflammatory skin lessons [105]. Since then, and through a series of unexpected discoveries, thalidomide and derivatives lenalidomide and pomalidomide have been repurposed for treating multiple myeloma and myelodysplastic syndrome (Fig. 6A) [106]. They also show promise in the treatment of autoimmune disorders such as systemic lupus erythematosus and inflammatory bowel disease [107].
The mechanism of action of thalidomide was discovered by the Handa laboratory where they identified cereblon (CRBN) as a thalidomide-binding protein [108]. They also showed CRBN forms an E3 ubiquitin ligase complex with damage-specific DNA binding protein 1 (DDB1) and CUL4A, inhibiting its autoubiquitination activity. Later research suggested thalidomide and its derivatives IMiDs (Immunomodulatory imide Drug derivatives) acted as molecular glues by stabilizing CRBN and recruited several neosubstrates for ubiquitination leading to the proteasomal degradation of the transcription factors (IKZF1) and Aiolos (IKZF3) (Fig. 6B) [109]. The direct outcome of IKZF1 and IKZF3 is the downregulation of the IRF4/MYC pathway immunomodulating the function of T cells and B cells, which leads to the observed cytotoxicity in multiple myeloma cells [110].
This new mechanism of action in which small molecules could stabilize PPIs and selectively target proteins for ubiquitination and further degradation presented a novel class of therapeutics for certain proteins that were thought to be undruggable. A recent example of this was the development of arylsulfonamides, which bind and stabilize DCAF15, the substrate receptor for the CRL4-DCAF15 E3 ubiquitin ligase and promote targeted protein degradation of mRNA splicing factor RBM39 [39, 111]. This in turn inhibits cell cycle progression and has potential use as an anticancer therapeutic.
Novel molecular glue compounds and targets
PKM2 molecular glues
Pyruvate kinase is an enzyme involved in the final and rate-limiting step of glycolysis: the conversion of phosphoenolpyruvate (PEP) to pyruvate. As such, pyruvate kinase plays an important role in regulating cell metabolism. In mammals, pyruvate kinase has four isomeric, tissue specific forms: PKL, PKR, PKM1, and PKM2 [112]. PKL is mainly found in kidney, liver, and red blood cells; PKR is found in red blood cells; PKM1 is found in myocardium, skeletal muscle, and brain tissues; PKM2 is found in brain and liver tissues [113, 114].
PKM1 and PKM2 are formed by alternative splicing from a single PKM mRNA transcript [115]. PKM1 and PKM2 differ by 22 amino acids and have distinct regulatory properties. PKM1 is a constitutively active enzyme that shows increased affinity for its substrate PEP, and thus, PKM1 expression is predominantly found in differentiated adult tissues with high ATP requirements. PKM2 enzyme activity is dependent on allosteric regulation by fructose-1,6-bisphosphate (FBP) and posttranslational modifications [116]. PKM2 is usually expressed during fetal development and upregulated in all cancers and cancer cell lines, suggesting that PKM2 has a role in tumorigenesis and tumor growth [117]. Due to increased metabolic requirements in tumorigenesis, we observe cancer cells favor glycolysis over oxidative phosphorylation for their energy production, a phenomenon known as the Warburg effect. One would think that during cancer formation, there would be a switch from PKM1 to PKM2 isoform expression. However, the data suggests that there is no exchange in PKM1 to PKM2 isoform expression during cancer formation. Additionally, PKM2 is not specific to proliferating tissue and PKM1 is not specific for non-proliferation tissue [118].
PKM2 has high and low-activity oligomers. PKM2’s high activity oligomer is its tetrameric conformation. PKM2 is activated in this conformation by binding to FBP, which causes PKM2 to adopt a stable, active conformation similar to that of PKM1. PKM2 tetramer activation by FBP can be overridden by interaction of PKM2 with tyrosine-phosphorylated proteins in response to growth factor signaling, by post translational modifications, and by interactions with other metabolites, such as serine, rendering PKM2 into a dimeric conformation. The low-activity dimer PKM2 has low affinity for its substrate (PEP) and regulates the rate-limiting step of glycolysis that shifts the glucose metabolism from the normal respiratory chain to lactate production in tumor cells, thus allowing proliferating cells to accumulate glycolytic intermediates needed for biosynthesis [119]. Additionally, the dimeric form of PKM2 has been shown to translocate to the nucleus in response to epidermal growth factor receptor (EGFR) signaling, where it can regulate gene transcription by acting as a protein kinase [120]. From here, PKM2 can promote EMT and stemness in cancer cells [121]. Multiple studies have shown that activating PKM2 (tetramer conformation) correlates with reduced tumorigenicity in both in vitro and in vivo studies [122,123,124,125].
In the last 10 years, academic groups and pharmaceutical companies have been working on developing small molecule PKM2 activators [113, 122, 124, 126]. A recently published article by the Kumar lab offers a comprehensive review on PKM2 activators [127]. Well-known PKM2 activators such as Michelolide (MCL), DASA-58, and TEPP-46 (Fig. 7A) act by allosterically binding to PKM2 (Fig. 7B). Tolero Pharmaceuticals, now known as Sumitomo Dainippon Pharma Oncology, recently developed a small molecule PKM2 activator: TP-1454. TP-1454 was derived from SGI-9380, compounds designed by Astex Pharmaceuticals in 2013 [128]. One of the main differences between SGI-9380 and other PKM2 activators is that each SGI-9380 molecule makes contact with at least one residue in a second PKM2 monomer. This in turns bridges two PKM2 monomer forming a homodimer. The newly formed homodimer then binds to another homodimer forming a tetramer containing four SGI-9380 molecules (Fig. 7B). TP-1454, optimized from SGI-9380 for improved metabolic activity and pharmacokinetics, is thought to bind in a similar fashion at the dimer-dimer interface of PKM2, acting as a glue between each monomer of PKM2, which then form a tetramer with four bound TP-1454 molecules.
A PKM2 molecular glues that function as activators. MCL (Micheliolide), DASA-58, TEPP-46, SGI-9380 bind to PKM2 in 1:1, one molecule per monomer. B Binding sites for PKM2 substrate fructose 1,6-bisphosphatase (FBP) and activators on the crystal structure of a PKM2 tetramer. Note the diverse binding sites for each of the activators that bind allosterically to PKM2 monomers inducing the formation of a tetramer, which is the more active form of PKM2. C Structure of PKR activator Mitapivat
TP-1454 has shown success at modulating cellular metabolism and enhancing response to checkpoint inhibitors in preclinical solid tumor models [123]. Additionally, TP-1454 is a potent PKM2 activator in biochemical assays (AC50 = 10 nM) and cellular assays (AC < 50 nM). It is also reported to have low toxicity and be well tolerated in mice, rats, and dogs, even in repeat doses as high as 1000 mg/kg/day [129]. In 2020, TP-1454 entered human clinical trials as the first oral PKM2 activator for the treatment of advanced solid tumors alone and in combination with ipilimumab and nivolumab (NCT04328740).
PKR glues
PKR is the constitutively expressed pyruvate kinase isoform found in red blood cells with an important role in erythropoiesis. Mutations in PKR cause autosomal recessive enzymopathy that leads to deficiencies in pyruvate kinase activity in red blood cells [130]. These deficiencies result in compromised red-cell membrane homeostasis and hemolysis, which causes hemolytic anemia. In addition, pyruvate kinase deficiencies in red blood cells are associated with gallstones, pulmonary hypertension, extramedullary hematopoiesis, and iron overload [131]. Studies in mouse models genetically lacking PKR have shown extramedullary erythropoiesis associated with ineffective erythropoiesis, further supporting the role of PKR in erythroid maturation[132].
Mitapivat (AG-348) (Fig. 7F) is a PKR activator that functions as a molecular glue, currently in clinical trials and developed by Agios. Mitapivat specifically binds to a PKR allosteric pocket to stabilize/glue the active tetrameric form and enhance its affinity for its substrate, PEP, successfully modulating PKR activity [133, 134]. A phase 2 clinical trial of mitapivat in adult patients with pyruvate kinase deficiency who were not receiving regular transfusions showed that the molecule elicited a rapid and sustained increase in hemoglobin levels in ~50% of treated patients. Mitapivat was well tolerated, with a few transient adverse effects [135].
Another molecular glue that stabilizes the tetrameric form of PKR similarly to mitapivat is FT-4202, a compound developed by FORMA Therapeutics. FT-4202 works as an allosteric activator of PKR that also stabilizes the tetrameric form of PKR (both wild type and R510Q mutant), thereby lowering the Michaelis-Menten constant for PEP [136]. FT-4202 is also in clinical trials (NCT03815695) showing that it can raise hemoglobin levels in 86% of patients and reduce the occurrence of vaso-occlusive crisis, a common complication in patients with sickle-cell disease [137, 138].
14-3-3 molecular glues
14-3-3 proteins are highly conserved eukaryotic adapter proteins involved in multiple cellular processes such as cell-cycle control, signal transduction, protein trafficking, and apoptosis [139]. They mediate their physiological effects by binding to other proteins, assisting with protein folding, and modulating their subcellular localization, enzymatic activity, and PPIs [140].
The 14-3-3 protein family consists of seven isoforms in mammals, encoded by seven separate genes, each denoted by a Greek letter (β, γ, ε, ζ, η, τ and σ). 14-3-3 proteins mostly associate in functional homodimers or heterodimers with each of the monomers displaying an amphipathic groove, or cup-like structure, which binds phosphorylated Ser/Thr interaction motifs of their partner proteins [141]. 14-3-3 proteins have over 500 interaction partners, many of which are disease relevant such as Raf kinases, p53, Cdc25, YAP, MLF1, PAD6, Tau, LRRK2, CTRF, among others [142,143,144,145,146,147,148,149,150,151]. Because of 14-3-3’s involvement in multiple diseases and because both inhibition and stabilization of 14-3-3 PPIs have been shown with small molecules, 14-3-3 proteins have become of interest because there is substantial potential for novel pharmacological PPI modulation [152].
Multiple molecular glues for the stabilization and modulation of 14-3-3 have been identified. An example is fusicoccin A, a diterpene glycoside produced by the fungus Phomopsis amygdali (Fig. 8A). In 1994, it was discovered that its molecular target was the binary complex between the regulatory domain of the plasma membrane H + -ATPase (PMA) and 14-3-3 adapter proteins. Fusicoccin A’s mechanism of action was described as “acting as a molecular glue” [153]. Since then, related natural products like cotylenin A and semisynthetic fusiococcane analogs such as fusicoccin THF, ISIR-005 (Fig. 8A) have been valuable tools to study the stabilization of 14-3-3 binary complexes with a molecular glue mechanism [152]. In the study of 14-3-3 complexes, molecular glues are best described as chemical inducers of dimerization [154].
In a high-throughput screening of a 37,000 small molecule library, two compounds were found to stabilize the interaction between 14-3-3 and PMA2 [155]. The compounds, epibestatin and pyrrolidone1 (Fig. 8B, C), were found to stabilize the 14-3-3/PMA2 complex in different ways to each other and to fusicoccin A. Crystal structures revealed that pyrrolidone1 shares most of its protein contact surface with 14-3-3 (288.2 Å from a total of 349.7 Å) and has limited contact with PMA2. In contrast, epibestatin is trapped between 14-3-3 and PMA2 and shares a roughly equal contact surface with 14-3-3 (164.4 Å) and PMA2 (135.5 Å) [155]. The structure of pyrrolidone1 was further optimized by converting the pyrrolinone scaffold into a pyrazole, adding a tetrazole moiety to the phenyl ring that contacts PMA2, and introducing a bromine to the phenyl ring that exclusively contacts the 14-3-3 protein. This optimization rendered the compound with a threefold increase in stabilization of the 14-3-3/PMA2 complex [156].
Recently, a molecular glue to the SLP76/14-3-3 was reported to increase degradation of SLP76 [157]. Under normal conditions, 14-3-3 interacts with Ser376 of SLP76, mediating the proteasomal degradation of SLP76. SLP76 is an adapter protein that regulates the downstream signaling of TCRs helping to modulate the immune response. SLP76 is phosphorylated on Ser376 by the kinase HPK1 (hematopoietic progenitor kinase 1), resulting in a negative feedback mechanism. Enhanced degradation of SLP76 could negatively regulate TCR signaling, which could be beneficial in the context of autoimmune or T cell mediated inflammatory conditions [158]. Through high-throughput screening of the “Diversity Set” small molecule library (20,000 compounds), HTRF dose–response and dose-ratio assays, there were 16 compounds that showed increased stabilization of the 14-3-3/SLP76 complex, and thus the degradation of SLP76. Moreover, 13 of the 16 compounds were orthogonally confirmed by SPR to stabilize the 14-3-3/SLP76 interaction [157].
Conclusion
PPIs are crucial for functional cellular processes. In the field of therapeutics, we aim at developing drugs that can modulate these PPIs when PPIs are dysregulated and lead to disease. Most conventional drug development has been focused on protein inhibitory functions. However, a substantial portion of the genome has remained undruggable by this approach. Molecular glues and heterobifunctional degrader technology have made the undruggable genome more amicable to modulation.
Molecular glues can strengthen and stabilize PPIs, repair PPIs, and facilitate de novo PPIs [16]. By doing so, they can modulate protein function by inhibition or activation of one or both protein partners or by recruiting an E3 ubiquitin ligase that targets the protein complex for degradation. Likewise, PROTACs and other heterobifunctional degraders such as LYTACs, AUTACs, ATTECs, HaloPROTACs, AbTACs, and hydrophobic mediated degraders can facilitate the interactions between macromolecules and degradation machinery such as an E3 ligase or the lysosome or the autophagosome.
Designing molecular glues in a rational fashion has been and continues to be a challenge. Most molecular glues and their mechanism of action has been found serendipitously. Several approaches for high-throughput screening of molecular glues have involved screening small molecule libraries in the hope of finding needles in a haystack. Moving forward, it is crucial to understand and study protein-protein interfaces and functional ternary structures to better (semi) rationally design molecular glues in a high-throughput manner [16].
Heterobifunctional degraders are in theory, easier to design due to their modular nature. However, their degradation functionality depends on productive ternary complex formation. Recently, two groups have independently published native mass spectrometry as a method to study and predict functional ternary complexes and intermediaries, which can be a great tool to screen to rationally design PROTACs and their functionality in a high-throughput fashion [71, 72].
Against all odds, molecular glues have been found, optimized, and brought to the clinic. Of note is TP-1454, a molecular glue that stabilizes PMK2 and activates the kinase, the first of its kind to make it to clinical trials in oncology. Other molecular glues such as Mitapivat and FT-4202, both stabilizing PKR, are also success stories. Newly discovered CDK12 molecular glues that degrade cyclin K show promise as new cancer therapeutics [41,42,43,44]. Likewise, PROTACs have also been designed and brought to the clinic targeting androgen and estrogen receptor, BTK, BCL-XL, IRAK4, etc. and other targets show promising results. All in all, molecular glues and heterobifunctional degraders have the potential for enhanced PPIs or cell proteome editing and can have a significant impact on how we treat diseases in the future.
References
Amanda L, Garner KDJ. Protein-protein interactions and cancer: targeting the central dogma. Curr Top Medicinal Chem. 2011;11:258–80. https://doi.org/10.2174/156802611794072614.
Hopkins AL, Groom CR. The druggable genome. Nat Rev Drug Disco. 2002;1:727–30. https://doi.org/10.1038/nrd892.
Thiel P, Kaiser M, Ottmann C. Small-molecule stabilization of protein-protein interactions: an underestimated concept in drug discovery? Angew Chem Int Ed. 2012;51:2012–8. https://doi.org/10.1002/anie.201107616.
Ivanov AA, Khuri FR, Fu H. Targeting protein–protein interactions as an anticancer strategy. Trends Pharm Sci. 2013;34:393–400. https://doi.org/10.1016/j.tips.2013.04.007.
Wells JA, Mcclendon CL. Reaching for high-hanging fruit in drug discovery at protein–protein interfaces. Nature. 2007;450:1001–9. https://doi.org/10.1038/nature06526.
Dupont CA, Riegel K, Pompaiah M, Juhl H, Rajalingam K. Druggable genome and precision medicine in cancer: current challenges. FEBS J. 2021. https://doi.org/10.1111/febs.15788.
Lazo JS, Sharlow ER. Drugging undruggable molecular cancer targets. Annu Rev Pharm Toxicol. 2016;56:23–40. https://doi.org/10.1146/annurev-pharmtox-010715-103440.
Zeng S, Huang W, Zheng X, Liyan C, Zhang Z, Wang J, et al. Proteolysis targeting chimera (PROTAC) in drug discovery paradigm: Recent progress and future challenges. Eur J Medicinal Chem. 2021;210:112981. https://doi.org/10.1016/j.ejmech.2020.112981.
Dong G, Ding Y, He S, Sheng C. Molecular glues for targeted protein degradation: from serendipity to rational discovery. J Med Chem. 2021. https://doi.org/10.1021/acs.jmedchem.1c00895.
Mainolfi N, Rasmusson T. Targeted protein degradation. Platform technologies in drug discovery and validation. Annual reports in medicinal chemistry Volume 50, Chapter 9, 2017. pp. 301–334. https://doi.org/10.1016/bs.armc.2017.08.005
Komander D, Rape M. The ubiquitin code. Annu Rev Biochem. 2012;81:203–29. https://doi.org/10.1146/annurev-biochem-060310-170328.
Dermachi F, Brancolini C. Altering protein turnover in tumor cells: new opportunities for anti-cancer therapies. Drug Resist Updat. 2005;8:359–68. https://doi.org/10.1016/j.drup.2005.12.001.
Groll M, Clausen T. Molecular shredders: how proteasomes fulfill their role. Curr Opin Struct Biol. 2003;13:665–73.
Kisselev AF, Akopian TN, Woo KM, Goldberg AL. The sizes of peptides generated from protein by mammalian 26 and 20 S proteasomes. J Biol Chem. 1999;274:3363–71.
Raina K, Crews CM. Targeted protein knockdown using small molecule degraders. Curr Opin Chem Biol. 2017;39:46–53. https://doi.org/10.1016/j.cbpa.2017.05.016.
Kozicka Z, Thomä NH. Haven’t got a glue: protein surface variation for the design of molecular glue degraders. Cell Chem Biol. 2021;28:1032–47. https://doi.org/10.1016/j.chembiol.2021.04.009.
Simonetta KR, Taygerly J, Boyle K, Basham SE, Padovani C, Lou Y, et al. Prospective discovery of small molecule enhancers of an E3 ligase-substrate interaction. Nat Commun. 2019;10. https://doi.org/10.1038/s41467-019-09358-9.
Garcia-Seisdedos H, Empereur-Mot C, Elad N, Levy ED. Proteins evolve on the edge of supramolecular self-assembly. Nature. 2017;548:244–7. https://doi.org/10.1038/nature23320.
Che Y, Gilbert AM, Shanmugasundaram V, Noe MC. Inducing protein-protein interactions with molecular glues. Bioorg Medicinal Chem Lett. 2018;28:2585–92. https://doi.org/10.1016/j.bmcl.2018.04.046.
Leissing TM, Luh LM, Cromm PM. Structure driven compound optimization in targeted protein degradation. Drug Discov Today Technol. 2020. https://doi.org/10.1016/j.ddtec.2020.11.005.
Han B. A suite of mathematical solutions to describe ternary complex formation and their application to targeted protein degradation by heterobifunctional ligands. J Biol Chem. 2020;295:15280–91. https://doi.org/10.1074/jbc.ra120.014715.
Dale B, Cheng M, Park K-S, Kaniskan HÜ, Xiong Y, Jin J. Advancing targeted protein degradation for cancer therapy. Nat Rev Cancer. 2021. https://doi.org/10.1038/s41568-021-00365-x.
Scholes NS, Mayor-Ruiz C, Winter GE. Identification and selectivity profiling of small-molecule degraders via multi-omics approaches. Cell Chem Biol. 2021;28:1048–60. https://doi.org/10.1016/j.chembiol.2021.03.007.
Li Z, Wang C, Wang Z, Zhu C, Li J, Sha T, et al. Allele-selective lowering of mutant HTT protein by HTT–LC3 linker compounds. Nature. 2019;575:203–9. https://doi.org/10.1038/s41586-019-1722-1.
Chini A. Application of yeast-two hybrid assay to chemical genomic screens: a high-throughput system to identify novel molecules modulating plant hormone receptor complexes. Methods in molecular biology. Humana Press. 2014;1056:35-43. https://doi.org/10.1007/978-1-62703-592-7_4.
Zhai Q, Zhang X, Wu F, Feng H, Deng L, Xu L, et al. Transcriptional mechanism of jasmonate receptor COI1-mediated delay of flowering time in arabidopsis. Plant Cell. 2015;27:2814–28. https://doi.org/10.1105/tpc.15.00619.
Chini A, Fonseca S, Fernández G, Adie B, Chico JM, Lorenzo O, et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature. 2007;448:666–71. https://doi.org/10.1038/nature06006.
Katsir L, Schilmiller AL, Staswick PE, He SY, Howe GA. COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. Proc Natl Acad Sci. 2008;105:7100–5. https://doi.org/10.1073/pnas.0802332105.
Chini A, Monte I, Fernández-Barbero G, Boter M, Hicks G, Raikhel N, et al. A small molecule antagonizes jasmonic acid perception and auxin responses in vascular and non-vascular plants. bioRxiv. 2021. https://doi.org/10.1101/2021.02.02.429350.
Isobe Y, Okumura M, Mcgregor LM, Brittain SM, Jones MD, Liang X, et al. Manumycin polyketides act as molecular glues between UBR7 and P53. Nat Chem Biol. 2020;16:1189–98. https://doi.org/10.1038/s41589-020-0557-2.
Roberts AM, Ward CC, Nomura DK. Activity-based protein profiling for mapping and pharmacologically interrogating proteome-wide ligandable hotspots. Curr Opin Biotechnol. 2017;43:25–33. https://doi.org/10.1016/j.copbio.2016.08.003.
Weerapana E, Wang C, Simon GM, Richter F, Khare S, Dillon MBD, et al. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature. 2010;468:790–5. https://doi.org/10.1038/nature09472.
Isobe Y, Okumura M, White R, McGregor LM, McKenna JM, Tallarico JA, et al. Manumycin polyketides act as molecular glues between UBR7 and P53 to impair breast cancer pathogenicity. bioRxiv. 2019:814285. https://doi.org/10.1101/814285.
Gehrtz P, London N. Electrophilic natural products as drug discovery tools. Trends Pharm Sci. 2021;42:434–47. https://doi.org/10.1016/j.tips.2021.03.008.
Haritakun R, Srikitikulchai P, Khoyaiklang P, Isaka M, Isariotins A–D. Alkaloids from the insect pathogenic fungus isaria tenuipes BCC 7831. J Nat Products. 2007;70:1478–80. https://doi.org/10.1021/np070291q.
Wørmer GJ, Villadsen NL, Nørby P, Poulsen TB. Concise asymmetric syntheses of streptazone A and abikoviromycin**. Angew Chem. 2021;133:10615–9. https://doi.org/10.1002/ange.202101439.
Jones LH. Expanding chemogenomic space using chemoproteomics. Bioorg Medicinal Chem. 2019;27:3451–3. https://doi.org/10.1016/j.bmc.2019.06.022.
Castaldi MP, Hendricks JA, Zhang AX. Design, synthesis, and strategic use of small chemical probes toward identification of novel targets for drug development. Curr Opin Chem Biol. 2020;56:91–7. https://doi.org/10.1016/j.cbpa.2020.03.003.
Bussiere DE, Xie L, Srinivas H, Shu W, Burke A, Be C, et al. Structural basis of indisulam-mediated RBM39 recruitment to DCAF15 E3 ligase complex. Nat Chem Biol. 2020;16:15–23. https://doi.org/10.1038/s41589-019-0411-6.
Han T, Goralski M, Gaskill N, Capota E, Kim J, Ting TC, et al. Anticancer sulfonamides target splicing by inducing RBM39 degradation via recruitment to DCAF15. Science. 2017;356:eaal3755. https://doi.org/10.1126/science.aal3755.
Słabicki M, Kozicka Z, Petzold G, Li Y-D, Manojkumar M, Bunker RD, et al. The CDK inhibitor CR8 acts as a molecular glue degrader that depletes cyclin K. Nature. 2020;585:293–7. https://doi.org/10.1038/s41586-020-2374-x.
Mayor-Ruiz C, Bauer S, Brand M, Kozicka Z, Siklos M, Imrichova H, et al. Rational discovery of molecular glue degraders via scalable chemical profiling. Nat Chem Biol. 2020;16:1199–207. https://doi.org/10.1038/s41589-020-0594-x.
Lv L, Chen P, Cao L, Li Y, Zeng Z, Cui Y, et al. Discovery of a molecular glue promoting CDK12-DDB1 interaction to trigger cyclin K degradation. eLife. 2020;9. https://doi.org/10.7554/elife.59994.
Dieter SM, Siegl C, Codo PL, Huerta M, Ostermann-Parucha AL, Schulz E, et al. Degradation of CCNK/CDK12 is a druggable vulnerability of colorectal cancer. Cell Rep. 2021;36:109394. https://doi.org/10.1016/j.celrep.2021.109394.
Naito M, Ohoka N, Shibata N, Tsukumo Y. Targeted protein degradation by chimeric small molecules, PROTACs and SNIPERs. Front Chem. 2019;7. https://doi.org/10.3389/fchem.2019.00849.
Zhou X, Dong R, Zhang J-Y, Zheng X, Sun L-P. PROTAC: a promising technology for cancer treatment. Eur J Medicinal Chem. 2020;203:112539. https://doi.org/10.1016/j.ejmech.2020.112539.
Shanmugasundaram K, Shao P, Chen H, Campos B, Mchardy SF, Luo T, et al. A modular PROTAC design for target destruction using a degradation signal based on a single amino acid. J Biol Chem. 2019;294:15172–5. https://doi.org/10.1074/jbc.ac119.010790.
Sakamoto KM, Kim KB, Kumagai A, Mercurio F, Crews CM, Deshaies RJ. Protacs: Chimeric molecules that target proteins to the Skp1–Cullin–F box complex for ubiquitination and degradation. Proc Natl Acad Sci. 2001;98:8554. https://doi.org/10.1073/pnas.141230798.
Schneekloth AR, Pucheault M, Tae HS, Crews CM. Targeted intracellular protein degradation induced by a small molecule: en route to chemical proteomics. Bioorg Medicinal Chem Lett. 2008;18:5904–8. https://doi.org/10.1016/j.bmcl.2008.07.114.
Bondeson DP, Mares A, Smith IED, Ko E, Campos S, Miah AH, et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat Chem Biol. 2015;11:611–7. https://doi.org/10.1038/nchembio.1858.
Itoh Y, Ishikawa M, Kitaguchi R, Sato S, Naito M, Hashimoto Y. Development of target protein-selective degradation inducer for protein knockdown. Bioorg Medicinal Chem. 2011;19:3229–41. https://doi.org/10.1016/j.bmc.2011.03.057.
Winter GE, Buckley DL, Paulk J, Roberts JM, Souza A, Dhe-Paganon S, et al. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science. 2015;348:1376–81. https://doi.org/10.1126/science.aab1433.
Zengerle M, Chan K-H, Ciulli A. Selective small molecule induced degradation of the BET bromodomain protein BRD4. ACS Chem Biol. 2015;10:1770–7. https://doi.org/10.1021/acschembio.5b00216.
Lai AC, Toure M, Hellerschmied D, Salami J, Jaime-Figueroa S, Ko E, et al. Modular PROTAC design for the degradation of oncogenic BCR-ABL. Angew Chem Int Ed. 2016;55:807–10. https://doi.org/10.1002/anie.201507634.
Xie H, Liang J-J, Wang Y-L, Hu T-X, Wang J-Y, Yang R-H, et al. The design, synthesis and anti-tumor mechanism study of new androgen receptor degrader. Eur J Medicinal Chem. 2020;204:112512. https://doi.org/10.1016/j.ejmech.2020.112512.
Neklesa TK, Winkler JD, Crews CM. Targeted protein degradation by PROTACs. Pharm Ther. 2017;174:138–44. https://doi.org/10.1016/j.pharmthera.2017.02.027.
Gadd MS, Testa A, Lucas X, Chan K-H, Chen W, Lamont DJ, et al. Structural basis of PROTAC cooperative recognition for selective protein degradation. Nat Chem Biol. 2017;13:514–21. https://doi.org/10.1038/nchembio.2329.
Pettersson M, Crews CM. PROteolysis TArgeting Chimeras (PROTACs)—past, present and future. Drug Disco Today Technol. 2019;31:15–27. https://doi.org/10.1016/j.ddtec.2019.01.002.
Bondeson DP, Smith BE, Burslem GM, Buhimschi AD, Hines J, Jaime-Figueroa S, et al. Lessons in PROTAC design from selective degradation with a promiscuous warhead. Cell Chem Biol. 2018;25:78–87. https://doi.org/10.1016/j.chembiol.2017.09.010. e5
Shibata N, Shimokawa K, Nagai K, Ohoka N, Hattori T, Miyamoto N, et al. Pharmacological difference between degrader and inhibitor against oncogenic BCR-ABL kinase. Sci Rep. 2018;8:13549. https://doi.org/10.1038/s41598-018-31913-5.
Steinebach C, Ng YLD, Sosič I, Lee C-S, Chen S, Lindner S, et al. Systematic exploration of different E3 ubiquitin ligases: an approach towards potent and selective CDK6 degraders. Chem Sci. 2020;11:3474–86. https://doi.org/10.1039/d0sc00167h.
Ren H, Santner A, Pozo JCD, Murray JAH, Estelle M. Degradation of the cyclin-dependent kinase inhibitor KRP1 is regulated by two different ubiquitin E3 ligases. Plant J. 2008;53:705–16. https://doi.org/10.1111/j.1365-313X.2007.03370.x.
Ottis P, Toure M, Cromm PM, Ko E, Gustafson JL, Crews CM. Assessing different E3 ligases for small molecule induced protein ubiquitination and degradation. ACS Chem Biol. 2017;12:2570–8. https://doi.org/10.1021/acschembio.7b00485.
Okamoto T, Imaizumi K, Kaneko M. The role of tissue-specific ubiquitin ligases, RNF183, RNF186, RNF182 and RNF152, in disease and biological function. Int J Mol Sci. 2020;21:3921. https://doi.org/10.3390/ijms21113921.
Kaneko M, Iwase I, Yamasaki Y, Takai T, Wu Y, Kanemoto S, et al. Genome-wide identification and gene expression profiling of ubiquitin ligases for endoplasmic reticulum protein degradation. Sci Rep. 2016;6:30955. https://doi.org/10.1038/srep30955.
Hou X, Zhang W, Xiao Z, Gan H, Lin X, Liao S, et al. Mining and characterization of ubiquitin E3 ligases expressed in the mouse testis. BMC Genomics. 2012;13:495. https://doi.org/10.1186/1471-2164-13-495.
Xie P, Guo S, Fan Y, Zhang H, Gu D, Li H. Atrogin-1/MAFbx enhances simulated ischemia/reperfusion-induced apoptosis in cardiomyocytes through degradation of MAPK phosphatase-1 and sustained JNK activation. J Biol Chem. 2009;284:5488–96. https://doi.org/10.1074/jbc.m806487200.
Mayor-Ruiz C, Jaeger MG, Bauer S, Brand M, Sin C, Hanzl A, et al. Plasticity of the Cullin-RING ligase repertoire shapes sensitivity to ligand-induced protein degradation. Mol Cell. 2019;75:849–58. https://doi.org/10.1016/j.molcel.2019.07.013.
Nowak RP, DeAngelo SL, Buckley D, He Z, Donovan KA, An J, et al. Plasticity in binding confers selectivity in ligand-induced protein degradation. Nat Chem Biol. 2018;14:706–14. https://doi.org/10.1038/s41589-018-0055-y.
Burslem GM, Smith BE, Lai AC, Jaime-Figueroa S, McQuaid DC, Bondeson DP, et al. The advantages of targeted protein degradation over inhibition: an RTK case study. Cell Chem Biol. 2018;25:67–77. https://doi.org/10.1016/j.chembiol.2017.09.009.
Sternicki LM, Nonomiya J, Liu M, Mulvihill MM, Quinn RJ. Native mass spectrometry for the study of PROTAC GNE-987-containing ternary complexes. ChemMedChem. 2021;16:2206–10. https://doi.org/10.1002/cmdc.202100113.
Beveridge R, Kessler D, Rumpel K, Ettmayer P, Meinhart A, Clausen T. Native mass spectrometry can effectively predict PROTAC efficacy. ACS Cent Sci. 2020;6:1223–30. https://doi.org/10.1021/acscentsci.0c00049.
Simard JR, Lee L, Vieux E, Improgo R, Tieu T, Phillips AJ, et al. High-throughput quantitative assay technologies for accelerating the discovery and optimization of targeted protein degradation therapeutics. SLAS Disco Advancing Sci Drug Disco. 2021;26:503–17. https://doi.org/10.1177/2472555220985049.
Weng G, Shen C, Cao D, Gao J, Dong X, He Q, et al. PROTAC-DB: an online database of PROTACs. Nucleic Acids Res 2020;49:D1381–7. https://doi.org/10.1093/nar/gkaa807.
Whitworth C, Ciulli A. New class of molecule targets proteins outside cells for degradation. Nature. 2020;584:193–4. https://doi.org/10.1038/d41586-020-02211-w.
Deane C. It’s a trap! Nat Chem Biol. 2020;16:1153. https://doi.org/10.1038/s41589-020-00680-8.
Eldeeb MA, Zorca CE, Goiran T. Extracellular protein degradation via the lysosome. Commun Chem. 2020;3. https://doi.org/10.1038/s42004-020-00397-8.
Banik SM, Pedram K, Wisnovsky S, Ahn G, Riley NM, Bertozzi CR. Lysosome-targeting chimaeras for degradation of extracellular proteins. Nature. 2020;584:291–7. https://doi.org/10.1038/s41586-020-2545-9.
Alabi SB, Crews CM. Major advances in targeted protein degradation: PROTACs, LYTACs, and MADTACs. J Biol Chem. 2021;296:100647. https://doi.org/10.1016/j.jbc.2021.100647.
Takahashi D, Moriyama J, Nakamura T, Miki E, Takahashi E, Sato A, et al. AUTACs: cargo-specific degraders using selective autophagy. Mol Cell. 2019;76:797–810. https://doi.org/10.1016/j.molcel.2019.09.009. e10
De Vita E, Lucy D, Tate EW. Beyond targeted protein degradation: LD·ATTECs clear cellular lipid droplets. Cell Res. 2021;31:945–6. https://doi.org/10.1038/s41422-021-00546-1.
Fu Y, Lu B. Targeting lipid droplets for autophagic degradation by ATTEC. Autophagy. 2021:1–3. https://doi.org/10.1080/15548627.2021.1967616.
Buckley DL, Raina K, Darricarrere N, Hines J, Gustafson JL, Smith IE, et al. HaloPROTACS: use of small molecule PROTACs to induce degradation of HaloTag fusion proteins. ACS Chem Biol. 2015;10:1831–7. https://doi.org/10.1021/acschembio.5b00442.
Tomoshige S, Hashimoto Y, Ishikawa M. Efficient protein knockdown of HaloTag-fused proteins using hybrid molecules consisting of IAP antagonist and HaloTag ligand. Bioorg Medicinal Chem. 2016;24:3144–8. https://doi.org/10.1016/j.bmc.2016.05.035.
Tovell H, Testa A, Maniaci C, Zhou H, Prescott AR, Macartney T, et al. Rapid and reversible knockdown of endogenously tagged endosomal proteins via an optimized HaloPROTAC degrader. ACS Chem Biol. 2019;14:882–92. https://doi.org/10.1021/acschembio.8b01016.
Cotton AD, Nguyen DP, Gramespacher JA, Seiple IB, Wells JA. Development of antibody-based PROTACs for the degradation of the cell-surface immune checkpoint protein PD-L1. J Am Chem Soc. 2021;143:593–8. https://doi.org/10.1021/jacs.0c10008.
Labrijn AF, Janmaat ML, Reichert JM, Parren PWHI. Bispecific antibodies: a mechanistic review of the pipeline. Nat Rev Drug Disco. 2019;18:585–608. https://doi.org/10.1038/s41573-019-0028-1.
Lin J, Jin J, Shen Y, Zhang L, Gong G, Bian H, et al. Emerging protein degradation strategies: expanding the scope to extracellular and membrane proteins. Theranostics. 2021;11:8337–49. https://doi.org/10.7150/thno.62686.
Deeks ED. Fulvestrant: a review in advanced breast cancer not previously treated with endocrine therapy. Drugs. 2018;78:131–7. https://doi.org/10.1007/s40265-017-0855-5.
Osborne CK, Wakeling A, Nicholson RI. Fulvestrant: an oestrogen receptor antagonist with a novel mechanism of action. Br J Cancer. 2004;90:S2–6. https://doi.org/10.1038/sj.bjc.6601629.
Carlson RW. The history and mechanism of action of fulvestrant. Clin Breast Cancer. 2005;6:S5–8. https://doi.org/10.3816/CBC.2005.s.008.
Iorfida M, Mazza M, Munzone E. Fulvestrant in combination with CDK4/6 inhibitors for HER2- metastatic breast cancers: current perspectives. Breast Cancer. 2020;12:45–56. https://doi.org/10.2147/BCTT.S196240.
Jones RH, Casbard A, Carucci M, Cox C, Butler R, Alchami F, et al. Fulvestrant plus capivasertib versus placebo after relapse or progression on an aromatase inhibitor in metastatic, oestrogen receptor-positive breast cancer (FAKTION): a multicentre, randomised, controlled, phase 2 trial. Lancet Oncol. 2020;21:345–57. https://doi.org/10.1016/S1470-2045(19)30817-4.
Smyth LM, Tamura K, Oliveira M, Ciruelos EM, Mayer IA, Sablin M-P, et al. Capivasertib, an AKT kinase inhibitor, as monotherapy or in combination with fulvestrant in patients with AKT1E17K-mutant, ER-positive metastatic breast cancer. Clin Cancer Res. 2020;26:3947–57. https://doi.org/10.1158/1078-0432.Ccr-19-3953.
Dent S, Cortés J, Im YH, Diéras V, Harbeck N, Krop IE, et al. Phase III randomized study of taselisib or placebo with fulvestrant in estrogen receptor-positive, PIK3CA-mutant, HER2-negative, advanced breast cancer: the SANDPIPER trial☆. Ann Oncol. 2021;32:197–207. https://doi.org/10.1016/j.annonc.2020.10.596.
André F, Ciruelos EM, Juric D, Loibl S, Campone M, Mayer IA, et al. Alpelisib plus fulvestrant for PIK3CA-mutated, hormone receptor-positive, human epidermal growth factor receptor-2–negative advanced breast cancer: final overall survival results from SOLAR-1. Ann Oncol. 2021;32:208–17. https://doi.org/10.1016/j.annonc.2020.11.011.
Schreiber SL. The rise of molecular glues. Cell. 2021;184:3–9. https://doi.org/10.1016/j.cell.2020.12.020.
Schreiber SL, Crabtree GR. The mechanism of action of cyclosporin A and FK506. Immunol Today. 1992;13:136–42. https://doi.org/10.1016/0167-5699(92)90111-J.
Dumont FJ, Su Q. Mechanism of action of the immunosuppressant rapamycin. Life Sci. 1995;58:373–95. https://doi.org/10.1016/0024-3205(95)02233-3.
Sabatini DM, Erdjument-Bromage H, Lui M, Tempst P, Snyder SH. RAFT1: a mammalian protein that binds to FKBP12 in a rapamycin-dependent fashion and is homologous to yeast TORs. Cell. 1994;78:35–43. https://doi.org/10.1016/0092-8674(94)90570-3.
Dudkin L, Dilling MB, Cheshire PJ, Harwood FC, Hollingshead M, Arbuck SG, et al. Biochemical correlates of mTOR inhibition by the rapamycin ester CCI-779 and tumor growth inhibition. Clin Cancer Res. 2001;7:1758–64.
Zou Z, Tao T, Li H, Zhu X. mTOR signaling pathway and mTOR inhibitors in cancer: progress and challenges. Cell Biosci. 2020;10. https://doi.org/10.1186/s13578-020-00396-1.
Tan X, Calderon-Villalobos LIA, Sharon M, Zheng C, Robinson CV, Estelle M, et al. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature. 2007;446:640–5. https://doi.org/10.1038/nature05731.
Vargesson N. Thalidomide‐induced teratogenesis: history and mechanisms. Birth Defects Res Part C: Embryo Today Rev. 2015;105:140–56. https://doi.org/10.1002/bdrc.21096.
Sheskin J. Thalidomide in the treatment of lepra reactions. Clin Pharm Therapeutics. 1965;6:303–6. https://doi.org/10.1002/cpt196563303.
Ito T, Handa H. Molecular mechanisms of thalidomide and its derivatives. Proc Jpn Acad Ser B. 2020;96:189–203. https://doi.org/10.2183/pjab.96.016.
Millrine D, Kishimoto T. A brighter side to thalidomide: its potential use in immunological disorders. Trends Mol Med. 2017;23:348–61. https://doi.org/10.1016/j.molmed.2017.02.006.
Ito T, Ando H, Suzuki T, Ogura T, Hotta K, Imamura Y, et al. Identification of a primary target of thalidomide teratogenicity. Science. 2010;327:1345–50. https://doi.org/10.1126/science.1177319.
Lu G, Middleton RE, Sun H, Naniong M, Ott CJ, Mitsiades CS, et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science. 2014;343:305–9. https://doi.org/10.1126/science.1244917.
Stewart AK. Medicine. How thalidomide works against cancer. Science. 2014;343:256–7. https://doi.org/10.1126/science.1249543.
Du X, Volkov OA, Czerwinski RM, Tan H, Huerta C, Morton ER, et al. Structural basis and kinetic pathway of RBM39 recruitment to DCAF15 by a sulfonamide molecular glue E7820. Structure. 2019;27:1625–33. https://doi.org/10.1016/j.str.2019.10.005. e3
Zahra K, Dey T, Ashish, Mishra SP, Pandey U. Pyruvate kinase M2 and cancer: the role of PKM2 in promoting tumorigenesis. Front Oncol. 2020;10. https://doi.org/10.3389/fonc.2020.00159.
Israelsen WJ, Dayton TL, Davidson SM, Fiske BP, Hosios AM, Bellinger G. et al. PKM2 isoform-specific deletion reveals a differential requirement for pyruvate kinase in tumor cells. Cell. 2013;155:397–409. https://doi.org/10.1016/j.cell.2013.09.025.
Warner SL, Carpenter KJ, Bearss DJ. Activators of PKM2 in cancer metabolism. Future Medicinal Chem. 2014;6:1167–78. https://doi.org/10.4155/fmc.14.70.
Noguchi T, Inoue H, Tanaka T. The M1- and M2-type isozymes of rat pyruvate kinase are produced from the same gene by alternative RNA splicing. J Biol Chem. 1986;261:13807–12. https://doi.org/10.1016/s0021-9258(18)67091-7.
Zhang Z, Deng X, Liu Y, Liu Y, Sun L, Chen F. PKM2, function and expression and regulation. Cell Biosci. 2019;9. https://doi.org/10.1186/s13578-019-0317-8.
Mazurek S. Pyruvate kinase type M2: a key regulator of the metabolic budget system in tumor cells. Int J Biochem Cell Biol. 2011;43:969–80. https://doi.org/10.1016/j.biocel.2010.02.005.
Bluemlein K, Grüning N-M, Feichtinger RG, Lehrach H, Kofler B, Ralser M. No evidence for a shift in pyruvate kinase PKM1 to PKM2 expression during tumorigenesis. Oncotarget. 2011;2:393–400. https://doi.org/10.18632/oncotarget.278.
Lv L, Xu Y-P, Zhao D, Li F-L, Wang W, Sasaki N, et al. Mitogenic and oncogenic stimulation of K433 acetylation promotes PKM2 protein kinase activity and nuclear localization. Mol Cell. 2013;52:340–52. https://doi.org/10.1016/j.molcel.2013.09.004.
Gao X, Wang H, Yang JJ, Liu X, Liu ZR. Pyruvate kinase M2 regulates gene transcription by acting as a protein kinase. Mol Cell. 2012;45:598–609. https://doi.org/10.1016/j.molcel.2012.01.001.
Tanaka F, Yoshimoto S, Okamura K, Ikebe T, Hashimoto S. Nuclear PKM2 promotes the progression of oral squamous cell carcinoma by inducing EMT and post-translationally repressing TGIF2. Oncotarget. 2018;9:33745–61. https://doi.org/10.18632/oncotarget.25850.
Parnell KM, Foulks JM, Nix RN, Clifford A, Bullough J, Luo B, et al. Pharmacologic activation of PKM2 slows lung tumor xenograft growth. Mol Cancer Therapeutics. 2013;12:1453–60. https://doi.org/10.1158/1535-7163.mct-13-0026.
Pathi S, Peterson P, Mangelson R, Tyagi E, Foulks JM, Whatcott CJ, et al. Abstract B080: PKM2 activation modulates metabolism and enhances immune response in solid tumor models. Mol Cancer Therapeutics. 2019;18:B080–B. https://doi.org/10.1158/1535-7163.Targ-19-b080.
Jiang JK, Walsh MJ, Brimacombe KR, Anastasiou D, Yu Y, Israelsen WJ, et al. ML265: a potent PKM2 activator induces tetramerization and reduces tumor formation and size in a mouse xenograft model. Probe Reports from the NIH Molecular Libraries Program. Bethesda (MD): National Center for Biotechnology Information (US); 2010.
Tee SS, Park JM, Hurd RE, Brimacombe KR, Boxer MB, Massoud TF, et al. PKM2 activation sensitizes cancer cells to growth inhibition by 2-deoxy-D-glucose. Oncotarget. 2017;8:90959–68. https://doi.org/10.18632/oncotarget.19630.
Brimacombe KR, Anastasiou D, Hong BS, Tempel W, Dimov S, Veith H, et al. ML285 affects reactive oxygen species’ inhibition of pyruvate kinase M2. Probe Reports from the NIH Molecular Libraries Program. Bethesda (MD); 2010.
Arora S, Joshi G, Chaturvedi A, Heuser M, Patil S, Kumar R. A perspective on medicinal chemistry approaches for targeting pyruvate kinase M2. J Med Chem. 2021. https://doi.org/10.1021/acs.jmedchem.1c00981.
Xu Y, Liu X-H, Saunders M, Pearce S, Foulks JM, Parnell KM, et al. Discovery of 3-(trifluoromethyl)-1H-pyrazole-5-carboxamide activators of the M2 isoform of pyruvate kinase (PKM2). Bioorg Medicinal Chem Lett. 2014;24:515–9. https://doi.org/10.1016/j.bmcl.2013.12.028.
Sommakia S, Pathi S, Matsumura Y, Allred C, Tyagi E, Lalonde M, et al. Abstract 606: Pkm2 activation modulates the tumor-immune microenvironment and enhances response to checkpoint inhibitors in preclinical solid tumor models. Cancer Res. 2021;81:606 https://doi.org/10.1158/1538-7445.Am2021-606.
Zanella A, Fermo E, Bianchi P, Chiarelli LR, Valentini G. Pyruvate kinase deficiency: the genotype-phenotype association. Blood Rev. 2007;21:217–31. https://doi.org/10.1016/j.blre.2007.01.001.
Grace RF, Bianchi P, Van Beers EJ, Eber SW, Glader B, Yaish HM, et al. Clinical spectrum of pyruvate kinase deficiency: data from the Pyruvate Kinase Deficiency Natural History Study. Blood. 2018;131:2183–92. https://doi.org/10.1182/blood-2017-10-810796.
Matte A, Federti E, Kung C, Kosinski PA, Narayanaswamy R, Russo R, et al. The pyruvate kinase activator mitapivat reduces hemolysis and improves anemia in a β-thalassemia mouse model. J Clin Investig. 2021;131. https://doi.org/10.1172/jci144206.
Kung C, Hixon J, Kosinski PA, Cianchetta G, Histen G, Chen Y, et al. AG-348 enhances pyruvate kinase activity in red blood cells from patients with pyruvate kinase deficiency. Blood. 2017;130:1347–56. https://doi.org/10.1182/blood-2016-11-753525.
Yang H, Merica E, Chen Y, Cohen M, Goldwater R, Kosinski PA, et al. Phase 1 single‐ and multiple‐ascending‐dose randomized studies of the safety, pharmacokinetics, and pharmacodynamics of AG‐348, a first‐in‐class allosteric activator of pyruvate kinase R, in healthy volunteers. Clin Pharm Drug Dev. 2019;8:246–59. https://doi.org/10.1002/cpdd.604.
Grace RF, Rose C, Layton DM, Galactéros F, Barcellini W, Morton DH, et al. Safety and efficacy of mitapivat in pyruvate kinase deficiency. N. Engl J Med. 2019;381:933–44. https://doi.org/10.1056/nejmoa1902678.
Ericsson A, Green N, Gustafson G, Han B, Lancia JR, Mitchell L, et al., Inventors; FORMA Therapeutics Inc, Assignee. Pyrrolopyrrole compositions as pyruvate kinase (PKR) activators. SG patent SG 11201908670S A. 2018/03/20. 2019.
Brown RC, Cruz K, Kalfa TA, Kuypers FA, Saraf SL, Estepp JH, et al. FT-4202, an allosteric activator of pyruvate kinase-R, demonstrates proof of mechanism and proof of concept after a single dose and after multiple daily doses in a phase 1 study of patients with sickle cell disease. Blood. 2020;136:19–20. https://doi.org/10.1182/blood-2020-134269.
Wood KW, Geib J, Wu E, Berlin J, Webster I, Ataga KI, et al. An adaptive, randomized, placebo-controlled, double-blind, multi-center study of oral FT-4202, a pyruvate kinase activator in patients with sickle cell disease (PRAISE). Blood 2020;136:19–20. https://doi.org/10.1182/blood-2020-134266.
Ottmann C. Small-molecule modulators of 14-3-3 protein–protein interactions. Bioorg Medicinal Chem. 2013;21:4058–62. https://doi.org/10.1016/j.bmc.2012.11.028.
Aitken A. 14-3-3 proteins: a historic overview. Semin Cancer Biol. 2006;16:162–72. https://doi.org/10.1016/j.semcancer.2006.03.005.
Cornell B, Toyo-oka K. 14-3-3 proteins in brain development: neurogenesis, neuronal migration and neuromorphogenesis. Front Mol Neurosci. 2017;10. https://doi.org/10.3389/fnmol.2017.00318.
Molzan M, Schumacher B, Ottmann C, Baljuls A, Polzien L, Weyand M, et al. Impaired binding of 14-3-3 to C-RAF in noonan syndrome suggests new approaches in diseases with increased Ras signaling. Mol Cell Biol. 2010;30:4698–711. https://doi.org/10.1128/MCB.01636-09.
Conklin DS, Galaktionov K, Beach D. 14-3-3 proteins associate with cdc25 phosphatases. Proc Natl Acad Sci USA. 1995;92:7892–6. https://doi.org/10.1073/pnas.92.17.7892.
Vassilev A. TEAD/TEF transcription factors utilize the activation domain of YAP65, a Src/Yes-associated protein localized in the cytoplasm. Genes Dev. 2001;15:1229–41. https://doi.org/10.1101/gad.888601.
Schumacher B, Mondry J, Thiel P, Weyand M, Ottmann C. Structure of the p53 C-terminus bound to 14-3-3: implications for stabilization of the p53 tetramer. FEBS Lett. 2010;584:1443–8. https://doi.org/10.1016/j.febslet.2010.02.065.
Sun Y, Fu A, Xu W, Chao J-R, Moshiach S, Morris SW. Myeloid leukemia factor 1 interfered with Bcl-XL to promote apoptosis and its function was regulated by 14-3-3. J Physiol Biochem. 2015;71:807–21. https://doi.org/10.1007/s13105-015-0445-5.
Higo J, Kawabata T, Kusaka A, Kasahara K, Kamiya N, Fukuda I, et al. Molecular interaction mechanism of a 14-3-3 protein with a phosphorylated peptide elucidated by enhanced conformational sampling. 2020.
Rose R, Rose M, Ottmann C. Identification and structural characterization of two 14-3-3 binding sites in the human peptidylarginine deiminase type VI. J Struct Biol. 2012;180:65–72. https://doi.org/10.1016/j.jsb.2012.05.010.
Sadik G, Tanaka T, Kato K, Yamamori H, Nessa BN, Morihara T, et al. Phosphorylation of tau at Ser214 mediates its interaction with 14-3-3 protein: implications for the mechanism of tau aggregation. J Neurochem. 2009;108:33–43. https://doi.org/10.1111/j.1471-4159.2008.05716.x.
Stevers LM, Lam CV, Leysen SFR, Meijer FA, van Scheppingen DS, de Vries RMJM, et al. Characterization and small-molecule stabilization of the multisite tandem binding between 14-3-3 and the R domain of CFTR. Proc Natl Acad Sci USA. 2016;113:E1152–61. https://doi.org/10.1073/pnas.1516631113.
Muda K, Bertinetti D, Gesellchen F, Hermann JS, von Zweydorf F, Geerlof A, et al. Parkinson-related LRRK2 mutation R1441C/G/H impairs PKA phosphorylation of LRRK2 and disrupts its interaction with 14-3-3. Proc Natl Acad Sci USA. 2014;111:E34–43. https://doi.org/10.1073/pnas.1312701111.
Stevers LM, Sijbesma E, Botta M, Mackintosh C, Obsil T, Landrieu I, et al. Modulators of 14-3-3 protein–protein interactions. J Medicinal Chem. 2018;61:3755–78. https://doi.org/10.1021/acs.jmedchem.7b00574.
Oecking C, Eckerskorn C, Weiler EW. The fusicoccin receptor of plants is a member of the 14-3-3 superfamily of eukaryotic regulatory proteins. FEBS Lett. 1994;352:163–6. https://doi.org/10.1016/0014-5793(94)00949-x.
Bosica F. Rational design of small molecule 14-3-3 protein-protein interaction stabilizers. Phd Thesis 1 (Research TU/e / Graduation TU/e): Technische Universiteit Eindhoven; 2020.
Rose R, Erdmann S, Bovens S, Wolf A, Rose M, Hennig S, et al. Identification and structure of small-molecule stabilizers of 14-3-3 protein-protein interactions. Angew Chem Int Ed. 2010;49:4129–32. https://doi.org/10.1002/anie.200907203.
Richter A, Rose R, Hedberg C, Waldmann H, Ottmann C. An optimised small-molecule stabiliser of the 14-3-3–PMA2 protein–protein interaction. Chem Eur J. 2012;18:6520–7. https://doi.org/10.1002/chem.201103761.
Soini L, Redhead M, Westwood M, Leysen S, Davis J, Ottmann C. Identification of molecular glues of the SLP76/14-3-3 protein–protein interaction. RSC Medicinal Chem. 2021;12:1555–64. https://doi.org/10.1039/d1md00172h.
Koretzky GA, Abtahian F, Silverman MA. SLP76 and SLP65: complex regulation of signalling in lymphocytes and beyond. Nat Rev Immunol. 2006;6:67–78. https://doi.org/10.1038/nri1750.
Acknowledgements
We would like to honor and celebrate Dr Laurence Hurley for a magnificent career studded with many highly significant scientific contributions. Especially, we would like to thank Dr Hurley for shaping the careers of so many undergraduate and graduate students that passed through his labs. In the fight against cancer, the efforts of those he inspired, trained, and mentored will extend far into the future.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
EGW, JMF, ASJ, and SLW are employed by Sumitomo Dainippon Pharma Oncology.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Weagel, E.G., Foulks, J.M., Siddiqui, A. et al. Molecular glues: enhanced protein-protein interactions and cell proteome editing. Med Chem Res 31, 1068–1087 (2022). https://doi.org/10.1007/s00044-022-02882-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-022-02882-2