Abstract
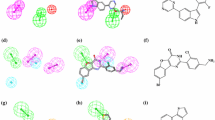
PIM3 (Proviral Integration site for Maloney murine leukemia virus kinase 3) is a proto-oncogene with serine/threonine kinase activity that prevents apoptosis, promotes cell survival, and stimulates protein translation. In addition, PIM3 functions in inflammation and immunity pathways. PIM3 inhibitors are being developed to treat cancer and inflammation-related disorders. Here we screen a 98,000 compound virtual library of natural products to identify those that are predicted to fit in the ATP site of PIM3. Since the structure of PIM3 has not been determined experimentally, we performed molecular structure prediction using the SWISS-MODEL tool to generate a PIM3 model structure for in silico screening. Compounds predicted to fit the ATP binding site of PIM3 were validated using biochemical assays, revealing activity against PIM3 for all eight candidates, with potencies mostly in the micromolar range. We tested several analogs of two validated candidates, the diosgenin glycoside dioscin and the biflavonoid hinokiflavone. Among five dioscin analogs, three exhibit similar potency against PIM3, and with some selectivity for PIM3 versus PIM1 and 2. Meanwhile, three of seven biflavonoid analogs exhibit sub-micromolar IC50 potency against PIM3, but with less selectivity for PIM3 versus PIM1 and 2.









Similar content being viewed by others
References
Koblish H, et al. Preclinical characterization of INCB053914, a novel pan-PIM kinase inhibitor, alone and in combination with anticancer agents, in models of hematologic malignancies. PLoS ONE. 2018;13:e0199108.
Brault L, et al. PIM serine/threonine kinases in the pathogenesis and therapy of hematologic malignancies and solid cancers. Haematologica. 2010;95:1004–15.
Li YY, Mukaida N. Pathophysiological roles of Pim-3 kinase in pancreatic cancer development and progression. World J Gastroenterol. 2014;20:9392–404.
Li YY, et al. Pim-3, a proto-oncogene with serine/threonine kinase activity, is aberrantly expressed in human pancreatic cancer and phosphorylates bad to block bad-mediated apoptosis in human pancreatic cancer cell lines. Cancer Res. 2006;66:6741–7.
Blanco-Aparicio C, Carnero A. Pim kinases in cancer: diagnostic, prognostic and treatment opportunities. Biochem Pharm. 2013;85:629–43.
Yang H, et al. Pim protein kinase-3 is regulated by TNF-α and promotes endothelial cell sprouting. Molecules cells. 2011;32:235–41.
Jackson LJ, et al. The role of PIM kinases in human and mouse CD4+ T cell activation and inflammatory bowel disease. Cell Immunol. 2012;272:200–13.
Zhang X, et al. PIM kinase as an executional target in cancer. J Cancer Prev. 2018;23:109–16.
Foulks JM, et al. A small-molecule inhibitor of PIM kinases as a potential treatment for urothelial carcinomas. Neoplasia. 2014;16:403–12.
Nakano H, et al. Design and synthesis of potent and selective PIM kinase inhibitors by targeting unique structure of ATP-Binding pocket. ACS Med Chem Lett. 2017;8:504–9.
Kirschner AN. et al. PIM kinase inhibitor AZD1208 for treatment of MYC-driven prostate cancer. J Natl Cancer Inst. 2015;107:dju407.
Chang M, et al. PIM kinase inhibitors downregulate STAT3(Tyr705) phosphorylation. Mol Cancer Ther. 2010;9:2478–87.
Jie W, et al. Inhibition of Pim-1 attenuates the proliferation and migration in nasopharyngeal carcinoma cells. Asian Pac J Trop Med. 2012;5:645–50.
Liu K, et al. Hispidulin suppresses cell growth and metastasis by targeting PIM1 through JAK2/STAT3 signaling in colorectal cancer. Cancer Sci. 2018;109:1369–81.
Fazio NF, et al. A natural product biflavonoid scaffold with anti-tryptase activity. Naunyn Schmiedebergs Arch Pharm. 2021;394:107–15.
Waterhouse A, et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 2018;46:W296–W303.
Morris GM, et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem. 2009;30:2785–91.
Fedorov O, et al. A systematic interaction map of validated kinase inhibitors with Ser/Thr kinases. Proc Natl Acad Sci USA. 2007;104:20523–8.
Ntie-Kang F, et al. AfroDb: a select highly potent and diverse natural product library from African medicinal plants. PLoS ONE. 2013;8:e78085.
Ru J, et al. TCMSP: a database of systems pharmacology for drug discovery from herbal medicines. J Cheminform. 2014;6:13.
Ntie-Kang F, et al. NANPDB: a resource for natural products from Northern African Sources. J Nat Prod. 2017;80:2067–76.
Chemicals, I Indofine Natural Products. 2016; Available from: https://zinc.docking.org/catalogs/indofinenp/.
Introbioscreen. IBScreen. 2018; Available from: http://zinc.docking.org/catalogs/ibs/.
Zhao X. et al. Dioscin induces apoptosis in human cervical carcinoma HeLa and SiHa cells through ROS-mediated DNA damage and the mitochondrial signaling pathway. Molecules. 2016;21:730.
Pei JS, et al. Amentoflavone induces cell-cycle arrest and apoptosis in MCF-7 human breast cancer cells via mitochondria-dependent pathway. Vivo. 2012;26:963–70.
Zhang G, et al. Dioscin suppresses hepatocellular carcinoma tumor growth by inducing apoptosis and regulation of TP53, BAX, BCL2 and cleaved CASP3. Phytomedicine. 2016;23:1329–36.
Guruvayoorappan C, Kuttan G. Amentoflavone stimulates apoptosis in B16F-10 melanoma cells by regulating bcl-2, p53 as well as caspase-3 genes and regulates the nitric oxide as well as proinflammatory cytokine production in B16F-10 melanoma cells, tumor associated macrophages and peritoneal macrophages. J Exp Ther Oncol. 2008;7:207–18.
Wang C, et al. Dioscin strengthens the efficiency of adriamycin in MCF-7 and MCF-7/ADR cells through autophagy induction: more than just down-regulation of MDR1. Sci Rep. 2016;6:28403.
Lee YJ, et al. Suppression of ERK/NF-κB activation is associated with amentoflavone-inhibited osteosarcoma progression. Anticancer Res. 2019;39:3669–75.
Zhou L, et al. Cdh1-mediated Skp2 degradation by dioscin reprogrammes aerobic glycolysis and inhibits colorectal cancer cells growth. EBioMedicine. 2020;51:102570.
Liu H, Yue Q, He S. Amentoflavone suppresses tumor growth in ovarian cancer by modulating Skp2. Life Sci. 2017;189:96–105.
Liu B, Yu S. Amentoflavone suppresses hepatocellular carcinoma by repressing hexokinase 2 expression through inhibiting JAK2/STAT3 signaling. Biomed Pharmacother. 2018;107:243–53.
Mao Z, et al. Potent effects of dioscin against hepatocellular carcinoma through regulating TP53-induced glycolysis and apoptosis regulator (TIGAR)-mediated apoptosis, autophagy, and DNA damage. Br J Pharm. 2019;176:919–37.
Chen Y, et al. Amentoflavone suppresses cell proliferation and induces cell death through triggering autophagy-dependent ferroptosis in human glioma. Life Sci. 2020;247:117425.
Chen CH, et al. Anticancer efficacy and mechanism of amentoflavone for sensitizing oral squamous cell carcinoma to cisplatin. Anticancer Res. 2020;40:6723–32.
Lim WC, et al. Dioscin suppresses TGF-β1-induced epithelial-mesenchymal transition and suppresses A549 lung cancer migration and invasion. Bioorg Med Chem Lett. 2017;27:3342–8.
Kim GL, et al. Amentoflavone, active compound of Selaginella tamariscina, inhibits in vitro and in vivo TGF-β-induced metastasis of human cancer cells. Arch Biochem Biophys. 2020;687:108384.
Lee JS, et al. Fatty acid synthase inhibition by amentoflavone induces apoptosis and antiproliferation in human breast cancer cells. Biol Pharm Bull. 2009;32:1427–32.
Hu XL, et al. Identification of amentoflavone as a potent highly selective PARP-1 inhibitor and its potentiation on carboplatin in human non-small cell lung cancer. Phytomedicine. 2018;50:88–98.
Pan X, et al. Amentoflavone and its derivatives as novel natural inhibitors of human Cathepsin B. Bioorg Med Chem. 2005;13:5819–25.
Seaver B, Smith JR. Inhibition of COX isoforms by nutraceuticals. J Herb Pharmacother. 2004;4:11–8.
Lv X, et al. Amentoflavone is a potent broad-spectrum inhibitor of human UDP-glucuronosyltransferases. Chem Biol Interact. 2018;284:48–55.
Park SY. et al. Strong and selective inhibitory effects of the biflavonoid selamariscina A against CYP2C8 and CYP2C9 enzyme activities in human liver microsomes. Pharmaceutics. 2020;12:343.
Acknowledgements
Funding for the research was provided by the Department of Physiology and Developmental Biology at the College of Life Sciences at Brigham Young University, Provo.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Russell, M.H., Fazio, N.F., Webster, J. et al. Selectivity and potency of natural product PIM kinase inhibitors identified by in silico docking. Med Chem Res 30, 1117–1124 (2021). https://doi.org/10.1007/s00044-021-02713-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-021-02713-w