Abstract
Discoveries in the field of genomics have revealed that non-coding genomic regions are not merely "junk DNA", but rather comprise critical elements involved in gene expression. These gene regulatory elements (GREs) include enhancers, insulators, silencers, and gene promoters. Notably, new evidence shows how mutations within these regions substantially influence gene expression programs, especially in the context of cancer. Advances in high-throughput sequencing technologies have accelerated the identification of somatic and germline single nucleotide mutations in non-coding genomic regions. This review provides an overview of somatic and germline non-coding single nucleotide alterations affecting transcription factor binding sites in GREs, specifically involved in cancer biology. It also summarizes the technologies available for exploring GREs and the challenges associated with studying and characterizing non-coding single nucleotide mutations. Understanding the role of GRE alterations in cancer is essential for improving diagnostic and prognostic capabilities in the precision medicine era, leading to enhanced patient-centered clinical outcomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Human Genome Project, which generated the first map of the human reference genome, marked a pivotal milestone in genetics research. However, the significance of non-coding genomic regions, formerly considered “junk DNA,” remained largely unexplored. The convergence of large-scale sequencing technologies and computational biology pipelines in the field of functional genomics has revealed the importance of non-coding regions in orchestrating gene expression programs [1]. These regions, collectively known as gene regulatory elements (GREs) [2], have been classified based on their impact on gene expression into gene promoters, enhancer elements (EEs), insulator elements (IEs), and silencers. Additionally, genomic alterations on GREs, ranging from single nucleotide variants (SNVs) to larger structural variants (SV), can disrupt the expression of regional as well as distant genes in disease states, specifically in cancer [3]. Consequently, these previously overlooked genetic modifications can dramatically impact normal gene expression programs [4, 5] by affecting the binding of transcription factors (TFs) [6], altering genome organization [7], modulating chromatin accessibility [8], or changing regional DNA methylation levels [9] at GREs.
Two main types of mutations that play a pivotal role in various diseases are involved in GRE dysregulation: germline single nucleotide polymorphisms (SNPs) and somatic SNVs [10]. Notably, genome-wide association studies (GWAS) have linked different SNPs located within non-coding regions to various types of cancer [11]. In contrast, projects such as the Pan-Cancer Analysis Whole Genomes (PCAWG) have identified thousands of non-coding somatic SNVs in numerous cancer types [12, 13]. Regardless of the origin, these point mutations are enriched within the transcription factor binding sites (TFBS) of GRE sequences in cancer [14,15,16,17]. This area of functional genomics opens an opportunity to leverage the clinical utility of non-coding mutations in different disease states, specifically in the context of cancer, bringing a chance to improve diagnostic, prognostic, and predictive models to improve patient’s clinical outcomes. While the impact of large SV on precision oncology has been discussed elsewhere [5], this review provides an overview of the recent findings on the functional impact of non-coding somatic and germline single nucleotide alterations affecting GREs in cancer. Considering the growing body of evidence highlighting the clinical significance of SNVs within non-coding regions of the genome, there has been a surge of innovation in technologies aimed at their comprehensive characterization and the exploration of their intricate molecular mechanisms [18, 19], which is also discussed.
Types and definitions of gene regulatory elements
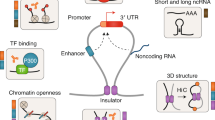
GREs are defined by a specific combination of histone marks and conglomerates of TFBS [20,21,22,23]. Based on the impact on the expression of regional as well as distant genes, GREs are classified into promoters, enhancer elements (EEs), insulator elements (IEs), and silencers (Fig. 1). Regarding the annotation of GRE, it is crucial to acknowledge the work conducted by the ENCODE (Encyclopedia of DNA Elements) consortium, which employed different experimental techniques – including ChIP-seq of TFs and histone marks, RNA-seq, among others – to characterize the regulatory elements in the human genome [24, 25]. This section provides information about each type of GRE to better understand the impact of single nucleotide variations on cancer biology.
Schematic overview of GREs and their chromatin interactions. The figure shows a DNA strand with promoters (P with red circles), enhancer elements or super-enhancers elements (EEs or SEEs in yellow), insulator elements (IEs in green) with CTCF binding, and silencer element (grey). The representation includes a Topologically Associating Domain (TAD), transcription factors (TFs), mediator protein complex, histones, and their marks in each type of GRE
Gene promoters comprise sequences upstream of the transcription start site (TSS), where the transcription machinery is assembled [26]. Many genes have been described to have alternative TSS [27]; as a result, different promoters can be associated with a single gene. However, the impact of gene promoters is usually associated with a nearby single gene. On the other hand, enhancer elements (EE) are defined by clusters of TFBS whose activation may affect the expression of both regional and distant genes by recruiting coactivators in cooperation [28]. Thus, these cis-regulatory elements have a highly variable location relative to the target genes [29]. EEs are activated or repressed in a spatial–temporal manner to define cellular fate during development [30]. As a consequence of its activation, the chromatin is looped allowing the proximity between EEs and promoters through the action of mediator proteins called cohesins [31]. Moreover, a single EE can regulate multiple genes, and one gene can be regulated by multiple EEs [32]. In addition, conglomerates of EEs have been defined as super-enhancer elements (SEEs). These GREs span large genomic regions and are enriched in binding motifs for master TFs and cofactors [33, 34]. Multiple TFs can occupy SEEs, modulating gene expression through SEE-promoter interactions, and forming core transcriptional regulatory circuits [35]. These elements are capable of driving cell-type-specific genes involved in key hemostatic functions and defining cell fates. Thus, the alteration of SEEs has been demonstrated to be crucial for tumor development and progression, as well as in therapeutic drug resistance or insensitivity [36, 37]. Another type of GREs known as silencer elements have the opposite effect compared to EEs. These regulatory elements repress gene expression by blocking the TF aggregation on either the gene promoter or upstream regulatory elements [23, 38]. Moreover, dual-function regulatory elements (REs) have been characterized in Drosophila [39], yet their presence in mammals remains unexplored. These genomic regions exhibit the capacity to function as both EEs and silencer elements. Notably, more than 5% of human silencers display regulatory element properties, underscoring the versatility of REs [40]. Finally, interactions between gene promoters and EEs can be influenced by another type of GRE that acts as boundary elements, known as insulator elements (IEs) [41]. These types of GREs are responsible for generating and maintaining the chromatin structural units called Topologically Associating Domains (TADs), which divide the genome into different compartments confining the interaction of GREs inside TADs [42]. Thus, alterations affecting IEs disrupt the TAD organization and have also been confirmed to contribute to tumorigenesis [43]. Activation of IEs mainly involves the binding of two critical proteins, CCCTC-binding factor (CTCF) and cohesin (RAD21) [44, 45]. Therefore, dysregulation of IEs alters gene expression programs by reshaping the landscape of promoter-EE interactions. Apart from single nucleotide mutations involving CTCF binding sites, many IEs can be impaired through abnormal DNA methylation [46, 47].
Cancer-associated non-coding single nucleotide mutations in GREs
Numerous SNPs and SNVs have been identified outside of coding genomic regions [48, 49]. Mechanistically, these alterations can influence the stability of GREs, leading to an alteration in the balance between the expression of tumor suppressor genes and oncogenes [50,51,52]. In this context, genomic alterations that lack measurable biological or phenotypic effects are often referred to as "passenger mutations" [53], whereas mutations conferring advantages to tumors are denoted as "driver mutations". The latter can be further categorized as either "major drivers" or "mini drivers", based on their magnitude of impact [54]. Another important factor in determining the impact of the SNV is the type of GRE affected. Tables 1, 2, 3 highlight the most important SNVs associated with cancer, including both SNPs and somatic mutations that affect promoters, EEs, and IEs, respectively.
Non-coding single nucleotide mutations within gene promoters
SNPs in promoter regions that disrupt the TFBS are studied across various tumor types, including lung cancer [55], hepatocellular carcinoma [56], neuroblastoma [57,58,59], and breast cancer [60,61,62]. A well-described example of germline single nucleotide mutations in tumorigenesis are the SNPs located on the promoter region of the oncogene Murine Double Minute 2 homolog (MDM2) [63]. MDM2, which is under the control of two distinct promoters, P1 and P2 [64], can negatively modulate the tumor suppressor p53, targeting it for proteasomal degradation [65]. For example, the G-allele of the rs2279744, known as SNP309 at the P2 promoter increases MDM2 expression by elongating the Sp1 TFBS. This alteration significantly reduces the tumor suppressor p53 levels [66], ultimately enhancing the risk of cancer development in humans, as depicted in Fig. 2A. In the context of melanoma pathogenesis, the SNP309 variation generates a stronger E2F1 binding site (Fig. 2B), which is responsible for cyclin D1 modulation and tumor proliferation [67]. Another germline mutation described within this promoter (rs117039649), located just 24 bp upstream of the SNP309, has the opposite impact, by reducing the Sp1 binding affinity and, therefore, the expression levels of MDM2 in ovarian and breast cancer [68]. Furthermore, a third SNP (rs2870820) found on the MDM2 promoter, known as SNP55, leads to an allele-specific expression by impairing NF-κB binding (Fig. 2C) [69]. Thus, the MDM2 gene highlights the complex interplay between genetic variations and gene regulation, demonstrating that the same promoter can be affected by different SNPs, causing a substantial differential effect in pathogenesis.
SNPs in MDM2-P2 promoter and its oncogenic consequences. A The presence of the germline alteration rs2279744 promotes Sp1 binding diminishing the p53 tumor suppressor pathway [66]. B The same mutation in the MDM2-P2 promoter generates a strong affinity with the E2F1 TF, modulating cyclin D1 and generating tumor proliferation [67]. C rs2870820 (SNP55C > T) is related to MDM2-P2 transcriptional activity. SNP55C has an affinity for NFκB p50 homodimers and it suppresses oncogene MDM2 transcription. However, the alteration does not retain this affinity with NFκB p50 favoring oncogene transcription [69]
Somatic SNVs have been identified as affecting gene promoters in different cancer types as well [70,71,72]. One of the most relevant findings was in the human telomerase reverse transcriptase (TERT) gene [73, 74]. In glioblastoma, Bell et al. discovered two somatic SNVs (chr5:1,295,411; G > A and chr5:1,295,433; G > A) in the TERT core promoter, which led to an enhanced GABP recruitment [75]. In melanoma, the TERT promoter contains two highly recurrent somatic SNVs (chr5:1,295,228; C > T, and chr5:1,295,250; C > T) allowing the binding of the ETS TF [76]. The consequence of the increased affinity of these TFs is the reactivation of TERT, a common mechanism in multiple cancers that allows bypassing the replicative senescence [76]. Another example is found in the promoter of SEMA3C, a gene related to tumor development in glioma stem cells [77]. The presence of a somatic SNV (chr7:80,552,013; T > C) has been found to modify the binding affinity of several TFs, such as RUNX1, ZNF354C, FOXA2, and EN1. Importantly, this mutation alters the binding site for FOXA1 in the SEMA3C promoter, leading to a reduced TF binding to the region [78]. Similarly, a somatic SNV in the FOXA1 promoter region (chr14:38,064,406; G > A) has been detected in primary breast cancers [79]. The mutant motif creates a stronger binding site for TF members of the E2F family, promoting high expression levels of FOXA1. This gene works as a transcriptional pioneer factor in breast cancer, enhancing chromatin accessibility for estrogen receptor interaction to its genomic targets [80], and has been linked to decreased response to fulvestrant, an estrogen receptor antagonist [81, 82]. In melanoma, the SDHD promoter contains different C > T transitions within the core ETS TF binding motifs, such as C524T and C523T, specifically affecting the binding of GABPA, GABPB1, and ETS1 [71, 83]. These alterations lead to a decreased expression of SDHD, which is associated with an unfavorable prognosis [83]. Furthermore, in primary liver cancer, Lowdon RF et al. identified a somatic mutation (chr4:81,187,908; A > T) in the FGF5 promoter region, which generates a new MYC binding site and enhances FGF5 expression [84]. SNVs at promoter regions affecting gene expression in cancer have been compiled in Table 1.
Single nucleotide mutations affecting enhancer and super-enhancer elements in cancer
Non-coding single nucleotide mutations within EEs and SEEs have been shown to disrupt critical TFBSs and influence transcriptional regulation through intricate interactions between these genetic variations and the epigenomic landscape. GWAS studies have demonstrated this phenomenon across a spectrum of cancer types, including but not limited to ovarian cancer [85, 86], colorectal cancer [87], and chronic lymphocytic leukemia [88], as summarized in Table 2.
Germline alterations have been shown to have an important role in EE abnormal activity in cancer. For instance, in lung cancer, two SNPs (rs9390123 and rs9399451) were detected within an EE located near the PHACTR2-AS1 gene, resulting in the creation of a new POU2F1 binding site that potentially modulates the DNA repair capacity of this cancer type (Fig. 3) [89]. Cardinale et al. characterized the role of rs2995264, an SNP located within an EE near the OBFC1 gene, in melanoma [90]. The presence of the G allele of this SNP reduces the binding affinity of the MEOX2 TF, thereby promoting carcinogenesis. In the context of low-grade glioma, the presence of rs55705857 in a brain-specific EE disrupts the OCT2/4 binding motif. This alteration leads to an abnormally higher expression of MYC by enhancing the interaction between the EE and MYC gene promoter [91]. Similarly, the SNP rs174575 exerts its influence on a long-range EE and modulates FADS2 gene expression through an increased binding affinity for E2F1. The upregulation of FADS2 leads to an increase in Prostaglandin E2 metabolism, a known oncogenic factor contributing to colorectal cancer development [92].
The rs9390123 and rs9399451 are located in the same EE and affect the binding of TF and PHACTR2-AS1 transcription. In lung cancer, the two germline mutations allow POU2F1 binding in an enhancer element (EE in yellow) and, consequently, the interaction between it and the promoter of the oncogene PHACTR2-AS1, favoring its expression [89]
In breast cancer, SNPs located in EEs have been demonstrated to influence tumorigenic gene expression programs. Notably, multiple breast cancer-associated SNPs exhibit enrichment in FOXA1 binding sites. As previously mentioned, FOXA1 acts as a pioneer factor by binding to highly compacted heterochromatin and exposing genomic areas to other transcription factors, hence influencing cancer-related pathways. In this context, the presence of the [T] rs4784227 allele in an EE leads to an elevated affinity of FOXA1 compared to the [C] reference allele. In vitro experimentation demonstrated that this SNP, which is located 18.4 kb upstream of the TOX3 gene, interacts with FOXA1/Groucho/TLE proteins, resulting in local chromatin condensation and transcriptional suppression. As a result, the [T] rs4784227 variant allele is found to have a repressive effect on TOX3 gene expression [93]. Moreover, the rs9383590 SNP impairs the interaction between GATA3 and an EE located upstream of the ESR1 gene TSS. In this context, GATA3 acts as a repressor and the SNP consequence was an increase in ESR1 gene expression [94]. Another noteworthy SNP (rs10941679) located within an EE alters the gene expression program of breast cancer cell lines by establishing interactions with the MRPS30 and FGF10 promoter regions. This leads to MRPS30 downregulation, a gene involved in the apoptosis process, and FGF10 upregulation, a well-known oncogene [95]. In lung adenocarcinoma, Li X et al. characterized another relevant SNP (rs2853677) within an EE near the TERT gene, which disrupts the Snail1 TFBS and enhances TERT gene expression [96].
Several somatic SNVs have also been identified in EEs. For example, a somatic SNV within an EE converges upon the TEAD4/PAX8-binding sites, leading to the perturbation of the expression levels of PAX8-target genes during the progression of ovarian cancer [97]. Interestingly, somatic and germline mutations can cooperate in favoring TFBS perturbations. For example, in a study on promyelocytic leukemia conducted by Song H et al., recurrent non-coding somatic and germline mutations were detected in an EE located inside the third intron of the WT1 gene. These mutations were found to reduce the binding of MYB, thereby disrupting the EE-promoter interaction. Consequently, it resulted in a decreased expression of WT1, a critical regulator of hematopoiesis [98].
Interestingly, new data indicates that approximately 64% of disease-associated SNPs are found within genomic regions with SEE activity [99]. One example is the rs6854845, which disrupts long-range chromosomal interaction between SEE and target genes CXCLs, EPGN, and EREG. This has been linked to a transcriptional switch that has a pivotal role in cell proliferation and inflammatory response in colon cancer [100]. Similarly, the rs11064124 G > A influences the binding of the vitamin D receptor (VDR), resulting in reduced expression of the tumor suppressor genes CD9 and PLEKHG6, ultimately promoting the development of colon cancer [101]. In diffuse large B-cell lymphoma, Kleinstern et al. identified two SNPs, rs6773363 and rs9831894, both located in the same SEE. While the presence of rs9831894 leads the SEE to interact with immune response genes, the rs6773363 variant promotes the interaction with oncogenes, consequently fostering tumor growth [102]. In a study of associations between SNPs and neuroblastoma, it has been observed that the rs2168101 G > T disrupts a binding site for the members of the GATA TF family within a SEE involved in LMO1 gene expression, ultimately contributing to neuroblastoma progression [103]. Finally, in chronic lymphocytic leukemia, the rs539846 variant disrupts a RELA binding site within an SEE. This disruption is associated with decreased expression of BMF, thereby enhancing the expression of the anti-apoptotic protein BCL2, a well-known oncogenic hallmark [104].
Alternatively, somatic SNVs can also contribute to generating new SEEs. In a subset of T-cell acute lymphoblastic leukemia cases, a singular somatic alteration has been observed to profoundly affect MYB binding affinity, resulting in the formation of a SEE located upstream of the TAL1 oncogene [105]. The evidence highlighting the involvement of non-coding mutations in governing SEE is just beginning to emerge. The characterization of non-coding mutations affecting these elements may unveil novel theranostic biomarkers to enhance the management of this disease.
Single nucleotide mutations on insulator elements in cancer
A comprehensive examination of SNPs and somatic SNVs affecting IEs is summarized in Table 3. As previously discussed, the activation of IEs relies on CTCF binding and the formation of homodimers with other CTCF-IE complexes. Somatic as well as germline single nucleotide mutations that interfere with the consensus CTCF motif can disrupt the binding of the CTCF protein and, therefore impact the activation of IEs [15]. This phenomenon has been observed in various types of cancers [106]. For example, the rs60507107 impacting a CTCF binding site (Fig. 4) has been identified as a susceptible SNP for lung cancer development [107]. An elevated risk of breast cancer development has been associated with the G/G variant of the rs11540855. Functional genomics studies in both tissue and cell lines have revealed that individuals with this variant have higher expression of the ANKLE1 gene due to the disruption of the CTCF binding to an IE that controls the expression of the ANKLE1 gene [108].
The SNP rs60507107 disturbs CTCF binding affinity in an IE. Lung cancer development is promoted by the presence of this SNP because it breaks an insulator element (IE in light green), increasing DAGLA expression, a gene that is related to carcinogenesis in different types of tumors [107]
Similarly, somatic SNVs have been recognized to influence IEs in cancer. In the context of melanoma, a somatic mutation (chr5:111,887,319 G > A) has been identified in a CTCF motif. This non-coding mutation disrupts the loop formation, resulting in the dysregulation of APC expression, a crucial tumor suppressor gene [109]. Another study conducted in melanoma identified an insulator (chr19:41,767,305–41771623) that displayed seven different somatic hotspots. Different somatic mutations on this IE increased the expression of TGFB1, contributing to aggressiveness. A mechanism detailing how UV-induced DNA damage leads to somatic SNVs in CTCF binding sites and, as a consequence, mutagenesis in human skin cells has been promoted [16].
In a separate study in gastrointestinal cancer, Guo YA et al. delved into the prediction and evaluation of three somatic non-coding mutations that have a discernible impact on CTCF binding sites, subsequently causing alterations in TFBSs [110]. Non-coding mutations in CTCF motifs near oncogenes such as KCNJ5, FLI1, and MYC have also been reported in gastrointestinal cancer [111]. Despite their potential implications for cancer development, non-coding single nucleotide mutations affecting IEs are currently relegated to the status of "passenger" mutations [112] and remain overlooked in cancer research.
Furthermore, when CTCF binding is disrupted, it can trigger the upregulation of genes that are typically protected within TADs, isolated from neighboring EEs [113]. Both SNPs and somatic SVs have been observed to interfere with these contact domains. This disruption can result in the activation of oncogenes through the formation of novel promoter-enhancer interactions [114]. In particular, some TADs exhibit SNP-driven alterations in a cancer-specific manner due to the organization of genes known to drive cancer progression [115]. For instance, recent findings by Osman et al. unveiled the presence of risk SNPs at the boundaries of certain TADs in prostate and breast cancer, specifically associated with GREs implicated in these pathologies [116].
In patients with lung squamous cell carcinoma, the presence of the T allele of the rs58163073 variant has been demonstrated to significantly enhance SOX2 binding affinity within the TAD boundary. This alteration in chromatin conformation near the VDAC3 gene results in an elevated expression that fosters cancer progression [117]. Colorectal cancer exhibits a distinct regulatory scenario, where the upregulation of the RPS24 gene is driven by the presence of three SNPs within a TAD boundary (rs3740253, rs7071351, and rs12263636). This enables the formation of a pathological promoter-EE interaction [118]. In pancreatic cancer patients, the G allele of the rs2001389 weakens the binding site for CTCF resulting in TAD disruption. This alteration diminishes the expression of the tumor suppressor gene MFSD13A, ultimately culminating in increased tumor proliferation [119].
Silencer elements affected by non-coding mutations
The impact of non-coding single nucleotide mutations on silencer elements remains poorly understood in cancer. While some SNPs have been identified, to date, no somatic SNVs have been reported within silencer elements. Interestingly, a study by Doni Jayavelu et al. showed that cancer-associated SNPs are significantly enriched in non-coding regions with function as silencer elements [23]. Huang et al. showed that the rs12631656 variant alters the binding affinity of SOX13 and ARID5B, two repressors in T cells, in a silencer element [40]. In patients with endometrial cancer, the rs2494737 overlaps a silencer element located within the AKT1 gene [120]. The variant risk allele A creates a new binding site for the YY1 TF, a positive regulator of AKT1. These discoveries emphasize the need to deepen investigations into how somatic mutations and SNPs affect silencer elements, holding the potential to unveil a more profound comprehension of their significance in the pathogenesis of cancer.
Technical approaches to identify and characterize non-coding single nucleotide mutations in GREs
Characterizing non-coding single nucleotide mutations within GREs in cancer requires technology that can precisely identify such mutations and delineate their impact on gene expression. Thus, the techniques can be classified according to the type of information that is generated, ranging from the identification and annotation to the functional validation. While there are diverse approaches available to achieve this objective, it is crucial to carefully weigh the merits and limitations of each approach, which have been summarized in Tables 4, 5, 6, and 7.
Identification of novel single nucleotide mutations affecting non-coding GREs Detecting somatic SNVs requires analyzing tumor-derived specimens and contrasting them with normal tissues, whereas the SNPs can be determined from virtually any tissue in the subject. Different technologies are available for this purpose. Table 4 provides an overview of these technologies.
Over the past decade, next-generation sequencing (NGS) has led to the discovery of new SNVs in non-coding genomic regions. Whole-genome sequencing (WGS) provides a comprehensive insight into an individual’s genetic makeup. Yet, it faces challenges due to its cost, increased computational expenditures, complex data analysis, and the added burden of multiple tests. Nonetheless, it provides versatility in detecting a wide range of somatic variants, from common to extremely rare, contingent upon sequencing depth [121, 122]. In the next-generation sequencing technologies abovementioned, identifying non-coding variants affecting the TF binding motifs within GREs is highly susceptible to false positives because of the short binding length. Also, analyzing and pinpointing these variants can be challenging due to numerous sequencing chemistry errors that commonly result in many false positive variants [123]. Thus, achieving a balance between eliminating false positive variants (specificity) and retaining true variants (sensitivity) is essential [124]. The progress in computational biology has also radically improved the discovery of novel variants associated with cancer traits. For example, the use of phyloP scores generated from the genomic constraint based on base-pair level conservation across 240 mammals, spanning 100 million years of evolution, can be used for fine-mapping of disease-related non-coding mutations, including cancer [125].
Another technological approach to perform WGS and identify novel SNVs in cancer is Nanopore sequencing, which determines a DNA sequence through the electrical potential perturbations occurring as the DNA strand passes through a pore. It offers distinct advantages, including the generation of long-reads, real-time insights, and direct DNA sequencing without the need for a prior amplification. However, researchers must consider its limitations, such as elevated error rates, lower throughput, high economic cost, and base-calling challenges [126]. Alternatively, SNPs can also be determined using microarray technologies, which utilize reliable genotyping technology, offering a cost-effective approach to identifying risk loci. These arrays rely on established genetic variant reference panels and are inadequate to detect novel or rare disease-contributing SNPs [121]. Finally, once a novel non-coding SNV is identified, the validation on a larger sample cohort can be performed using cost-effective targeted approaches like Sanger sequencing [127] or Digital PCR [128].
Assessing the impact of non-coding mutations in GREs. Two main approaches are used to evaluate the effects of germline as well as somatic SNVs in GREs, each with its advantages and limitations. Indirect methods, like whole-genome epigenetic assays, provide a broad overview of a region’s regulatory status but may not pinpoint the impact of a specific genetic alteration [24, 25]. On the other hand, direct methods assess how individual alleles affect gene expression, either in an episomal or native context. However, these direct methods are currently low-throughput and require substantial resources for comprehensive evaluation of non-coding regions, such as repetition of experiments with short DNA fragments. But perhaps one of the main limitations of the latter is the impossibility of assessing the contribution of distal intrachromosomal or interchromosomal regions.
A high-throughput indirect method termed single-nucleotide polymorphisms evaluation by systematic evolution of ligands by exponential enrichment (SNP-SELEX) made estimations of TF relative affinity to predict the effects of non-coding variants [129]. In cancer, the integration of allelic imbalance of chromatin accessibility, TF motif discovery, and Regulome-Wide Association Study help identify potential causal risk variants and elucidate their underlying mechanisms [130].
Regarding direct techniques, the Multiplex Parallel Reporter Assays (MPRA) is based on the introduction of a plasmid construct into a cell containing a reporter gene (luciferase or green fluorescent protein), a promoter, and the mutant GRE candidate. These assays measure changes in luciferase activity or GFP expression to identify whether the mutation induces an activation or inactivation of the gene expression [131]. These approaches have been used in various studies of non-coding mutations in GREs, both in in vitro [131,132,133] and in vivo models [134, 135]. A specific application of MPRA is the self-transcribing active regulatory region sequencing (STARR-seq) approach, which quantifies the activity of multiple non-coding mutations simultaneously [136]. The STARR-seq method has been useful in systematically assessing the impact of non-coding mutations on GRE function [137, 138].
A primary drawback of these approaches is their inability to effectively evaluate the functional impact of the mutation within the native genomic context. To address these concerns, genome editing techniques provide a more physiologically relevant method for assessing the impact of non-coding mutations on tumor development. One promising approach involves the use of CRISPR and base editing screens with a phenotypic readout achieved by employing a single-guide RNA (sgRNA) dropout [139,140,141,142]. However, in cases where the target region is larger, leading to cellular heterogeneity, clonal selection may be necessary. Alternatively, protein binding assays, such as the Electrophoretic Mobility Shift Assay (EMSA), can be employed to elucidate the molecular functions of non-coding mutations. In an in vitro setting, DNA probes are exposed to antibodies targeting the candidate transcription factors to assess the binding affinity of different alleles surrounding a candidate mutation [143]. For unbiased techniques, DNA-affinity pulldown followed by mass spectrometry offers a valuable option [144].
In parallel, ATAC-seq (Assay for Transposase-Accessible Chromatin using sequencing) is a complementary and cost-effective approach that can determine the status of GREs based on chromatin accessibility. A hyperactive Tn5 transposase inserts sequencing adapters into open chromatin regions, which are subsequently subjected to NGS [145]. However, accessibility alone cannot unveil the nature of the GRE and the functional impact of single nucleotide mutations. This prompts the combination of ATAC-seq data with the mapping of histone marks defining promoters, EEs, and IEs, respectively. Table 5 summarizes each strategy, along with its benefits and drawbacks.
Techniques to identify DNA–Protein interactions. Since non-coding single nucleotide mutations can alter TFBS sequences influencing the ability of a TF to bind DNA, various approaches are employed to study changes in protein-DNA interactions. These techniques involve ChIP-seq (Chromatin Immunoprecipitation), CUT&RUN (Cleavage under targets and release using nuclease), and CUT&TAG (Cleavage Under Targets and Tagmentation). Table 6 provides a quick overview of these technical strategies.
ChIP-seq is an antibody-based technique that involves crosslinking between DNA–protein complexes, chromatin shearing, and antibody pulldowns for the studied factor [146]. The precipitated DNA fragments are then purified and sequenced or quantified by real-time PCR. However, ChIP-seq can be challenging, particularly when studying target proteins that are part of multiprotein complexes or do not directly interact with DNA, and due to variability introduced during sonication.
To address these challenges, novel techniques have emerged. CUT&RUN employs a recombinant Protein A-MNase fusion construct that binds to the factor of interest’s primary antibody and cleaves DNA around TFBS, generating small fragments for sequencing or real-time PCR [147]. Another innovative method, CUT&Tag, utilizes pA-Tn5 carrying sequencing adapters to generate DNA amplicons for tagmentation-based sequencing [148].
Techniques to unveil the impact of non-coding mutations on chromatin conformation. Massive alterations in TADs and chromatin conformation, due to somatic SNVs or pathological SNPs, can be assessed using a variety of techniques. Chromatin conformation methodologies provide essential validation for the impact of non-coding mutations in GREs that affect chromatin looping; nevertheless, their limited throughput makes them less suitable for variant screening [149]. Despite this limitation, it is important to highlight their utility in establishing connections between GREs and specific target genes. Additionally, they can enhance the scope of GWAS by elucidating functional links between non-coding point mutations and GRE activity [121]. Among these techniques, we find 4C (Circular Chromosome Conformation Capture), 5C (Chromosome Conformation Capture Carbon Copy), Hi-C, Promoter Capture Hi-C (PCHi-C), HiChIP, and ChIA-PET. All these methods used for unveiling the impact of non-coding mutations on chromatin conformation are displayed in Table 7.
In 4C, a circularization step is performed to screen physical interactions between chromosomes associated with the genomic region of interest. Subsequently, target genes are amplified to identify genome-wide interactions [150]. On the other hand, 5C involves the relegation of DNA fragments from crosslinked cells to promote ligation between cross-linked interacting DNA fragments, followed by ligation-mediated amplification and sequencing of the target fragment [151]. In Hi-C, after DNA digestion is completed, the ends of the fragments are labeled with biotinylated nucleotides for ligation and reversal of crosslinks, followed by sequencing using paired-end sequencing [152]. Additionally, PCHi-C allows the genome-wide detection of distal promoter-interacting regions using Hi-C libraries enriched in promoter sequences. This is achieved by selecting biotinylated RNA baits that are complementary to promoter-containing restriction fragments. The objective is to capture promoter sequences and their interacting GREs, thereby increasing the number of reads covering promoter regions and improving the sensitivity of the technique for these regions [153]. HiChIP has recently been introduced by Mumbach MR, et al., incorporates in situ Hi-C and transposase-mediated on-bead library construction with a robust, reproducible, and two-day protocol [154]. In HiChIP, long-range DNA interactions are initially formed within the nucleus before lysis, reducing the potential for false-positive interactions [155] and significantly enhancing the efficiency of DNA contact capture.
Finally, ChIA-PET takes a different approach to explore chromatin conformation by crosslinking DNA–protein complexes with formaldehyde in the nucleus, followed by sonication-induced breaks. After reversing the crosslinking, protein complexes are digested, and DNA fragments are extracted for sequencing [156]. The sequencing reads are then aligned and scrutinized to unveil long-distance interactions between TFs.
Challenges and future perspectives in the research of non-coding mutations with functional impact on GRE activity
Understanding the intricate landscape of non-coding mutations within GREs is pivotal for deciphering their roles in cancer initiation and progression, and their potential diagnostic and therapeutic implications. The main challenge lies in precisely pinpointing the genomic coordinates of the mutation and discerning the impact on the affected GRE [157]. However, an unsolved issue is determining when these non-coding mutations occur during tumor development and progression. Recent advances in single-cell technology have started to identify sub-stoichiometric alterations and their possible contributions to cancer providing a tool to potentially predict the timeline of occurrence [158].
The limited understanding of the non-coding genomic space has led to disparities in variant annotations across various databases, resulting in divergent predictions. Therefore, standardizing non-coding mutation annotation is an imperative step forward in this field [159]. Moreover, due to the lack of experimental data, many annotations in these databases rely on in-silico predictions. While international collaborative efforts have yielded proficient tools for variant calling, such as GATK (https://gatk.broadinstitute.org/hc/en-us), the extensive annotation of the non-coding genome remains an ongoing challenge. Additionally, the activity of GREs may vary with the tissue and site of origin, as well as the intrinsic heterogeneity present within cells, especially in the context of cancer, which brings additional challenges to predicting functional impact and outcomes. The coexistence of multiple genes within the same genomic region further complicates the endeavor of defining driver non-coding mutations on GREs.
The next major challenge involves translating these non-coding mutations into their causative roles in altering oncogenic networks. Researchers employ various experimental and in silico methods to characterize potential pathogenic non-coding mutations. Due to the insufficient experimental data, current in silico approaches utilize multiple machine learning and mathematical modeling to make predictions with available published data [160, 161]. For example, TURF [162] and GRAM [163] are computational tools that integrate various layers of information to prioritize non-coding regulatory variants across the human genome. Important databases such as the Ensembl project (https://www.ensembl.org) include the Ensembl Variant Effect Predictor, a robust toolset for analyzing, annotating, and prioritizing genomic variants in both coding and non-coding regions [164]. Fu et al. created FunSeq2, a computational framework designed to annotate and prioritize noncoding mutations by integrating extensive genomics and cancer datasets within a customizable context [165]. Additionally, the Chromatin-Chromatin Spatial Interaction (CCSI) database displays chromatin interactions along with associated genes, EEs, and SNPs, offering comprehensive interaction maps and providing an analysis pipeline for annotating interactions [166]. GWAS4D (https://mulinlab.org/gwas4d) is a free web server that systematically analyzes genetic variants that could influence GREs by integrating annotations from cell type-specific chromatin states, epigenetic modifications, sequence motifs, and cross-species conservation [167]. Furthermore, Li et al. developed OncoBase, a valuable resource for the functional annotation of non-coding regulatory regions and for systematically benchmarking the regulatory effects of embedded non-coding somatic mutations in human carcinogenesis [168]. Lee PH et al. provide a comprehensive review of existing data resources and advanced analytical methods for aiding the in-silico prioritization of non-coding mutations [169]. Nonetheless, it’s crucial to acknowledge that each bioinformatic approach has its limitations, and variability exists between them [170].
While numerous assays have been conducted in cancer cell lines, they often overlook cellular diversity and physiological context. To address these limitations, researchers are turning to animal models and the raising application of genome editing techniques. For example, these approaches have been employed to create mice with mutations in the TERT promoter region, providing insights into non-coding TERT mutations detected in melanomas [171]. For EEs, although in vivo studies specific to cancer are still lacking, studies related to type 2 diabetes [172] and orofacial clefting [173] exist in the zebrafish model and polydactyly [174] and neuropsychiatric disorders [175] in mouse models. In the case of SEEs, Cui, S et al. deleted the EphA2-SEE in a xenograft model, which is present in various tumor types, effectively suppressing tumor proliferation [176]. Finally, in the context of IEs, mouse models incorporating mutations at CTCF binding sites were employed for developmental studies [177, 178], although their relevance to cancer research is limited. Collectively, these studies suggest that rectifying non-coding mutations within GREs offers a promising avenue for cancer therapeutics, even though in vivo research faces throughput limitations.
Equally important is the urgent need to translate non-coding GRE mutations into clinical significance, which could reshape cancer genomic medicine. Continuous advancements in CRISPR/Cas and base editing technologies are pivotal in this endeavor. For instance, in patients with β-thalassemia, an erythroid-specific EE within BCL11A contains numerous non-coding mutations that suppress γ-globin expression and fetal hemoglobin in erythroid cells [179]. Utilizing CRISPR/Cas9, researchers disrupted GATA1 binding sequences within the BCL11A EE, ultimately restoring γ-globin synthesis and fetal hemoglobin production in patients with β-hemoglobinopathies [180, 181]. Currently, there are no similar clinical trials applied to cancer using the CRISPR system. However, it is important to consider a series of potential limitations such as the presence of pre-existing immunity against CRISPR components restricting the safety and feasibility of in vivo delivery [182].
Conclusions
Recent advancements in sequencing techniques have significantly enriched our understanding of the impact of germline and somatic non-coding mutations in cancer. These alterations can occur in various non-coding gene regulatory regions of the genome, including promoters, EEs, IEs, and silencer elements. Notably, single nucleotide mutations within these regions can disrupt TFBSs, thereby altering TF recognition on gene regulatory elements. Consequently, this disruption can lead to a perturbation in the gene expression networks, ultimately resulting in an imbalanced expression of tumor suppressor genes and oncogenes.
Different techniques are available for the detection and functional inference of non-coding single nucleotide mutations in cancer. Both in vitro and in vivo models can be employed to assess the targetability of candidate variants, which, in turn, may inform the development of novel drugs and gene therapy strategies, or the development of prognostic or predictive biomarkers.
Despite the challenges posed by technical limitations, population heterogeneity, and inconsistencies in SNV annotations, recent research findings indicate a substantial impact of non-coding alterations on cancer development and progression. It is, therefore, essential that ongoing research efforts continue to elucidate the intricate links between non-coding mutations in gene regulatory regions and pathology. Moreover, the translation of this knowledge from laboratory research to clinical application is utterly important, specifically for aggressive forms of cancers that still do not have effective treatments. Thus, research in this field may ultimately fill the gap between benchside discoveries and bedside patient care.
Summary
-
Genomic alterations in non-coding regions, including somatic single nucleotide variations (SNVs) and single nucleotide polymorphisms (SNPs), can impact gene regulatory elements (GREs) and play a role in human disorders, including cancer.
-
Sequencing technology advances have revealed that over 90% of mutations in cancer are in non-coding genome regions.
-
Oncogenic somatic SNVs and SNPs within GREs can disrupt transcription factor binding sites (TFBS), leading to alterations in epigenetic mechanisms, including changes in chromatin accessibility and DNA methylation.
-
Cutting-edge and emerging high-throughput sequencing technologies allow the identification and cataloging of non-coding genomic alterations, thereby enabling a comprehensive exploration of the intricate landscape of genetic mutations within GREs.
-
Additional challenges lie in the massive data interpretation and understanding of the functional impact of these single nucleotide mutations on gene regulatory elements.
-
Understanding these alterations is crucial for identifying new theranostic biomarkers to add a new layer of information to improve the clinical management of patients with cancer.
Data availability
Not applicable.
References
Ohno S (1972) So much “junk” DNA in our genome. Brookhaven Symp Biol 23:366–370
Doane AS, Elemento O (2017) Regulatory elements in molecular networks. Wiley Interdiscip Rev Syst Biol Med. https://doi.org/10.1002/wsbm.1374
Kleinjan DJ, van Heyningen V (1998) Position effect in human genetic disease. Hum Mol Genet 7:1611–1618. https://doi.org/10.1093/hmg/7.10.1611
Herz H-M (2016) Enhancer deregulation in cancer and other diseases. BioEssays 38:1003–1015. https://doi.org/10.1002/bies.201600106
van Belzen IAEM, Schönhuth A, Kemmeren P, Hehir-Kwa JY (2021) Structural variant detection in cancer genomes: computational challenges and perspectives for precision oncology. NPJ Precis Oncol 5:15. https://doi.org/10.1038/s41698-021-00155-6
Degtyareva AO, Antontseva EV, Merkulova TI (2021) Regulatory SNPs: altered transcription factor binding sites implicated in complex traits and diseases. Int J Mol Sci. https://doi.org/10.3390/ijms22126454
Hawley JR, Zhou S, Arlidge C et al (2021) Reorganization of the 3D genome pinpoints noncoding drivers of primary prostate tumors. Cancer Res 81:5833–5848. https://doi.org/10.1158/0008-5472.CAN-21-2056
Lee S, Osmanbeyoglu HU (2022) Chromatin accessibility landscape and active transcription factors in primary human invasive lobular and ductal breast carcinomas. Breast Cancer Res 24:54. https://doi.org/10.1186/s13058-022-01550-y
Bird AP (1986) CpG-rich islands and the function of DNA methylation. Nature 321:209–213. https://doi.org/10.1038/321209a0
Karki R, Pandya D, Elston RC, Ferlini C (2015) Defining “mutation” and “polymorphism” in the era of personal genomics. BMC Med Genom 8:37. https://doi.org/10.1186/s12920-015-0115-z
Yang J, Adli M (2019) Mapping and making sense of noncoding mutations in the genome. Cancer Res 79:4309–4314. https://doi.org/10.1158/0008-5472.CAN-19-0905
Rheinbay E, Nielsen MM, Abascal F et al (2020) Analyses of non-coding somatic drivers in 2658 cancer whole genomes. Nature 578:102–111. https://doi.org/10.1038/s41586-020-1965-x
(2020) Pan-cancer analysis of whole genomes. Nature 578:82–93. https://doi.org/10.1038/s41586-020-1969-6
Morova T, McNeill DR, Lallous N et al (2020) Androgen receptor-binding sites are highly mutated in prostate cancer. Nat Commun 11:832. https://doi.org/10.1038/s41467-020-14644-y
Katainen R, Dave K, Pitkänen E et al (2015) CTCF/cohesin-binding sites are frequently mutated in cancer. Nat Genet 47:818–821. https://doi.org/10.1038/ng.3335
Sivapragasam S, Stark B, Albrecht A V, et al (2021) CTCF binding modulates UV damage formation to promote mutation hot spots in melanoma. EMBO J 40:e107795. https://doi.org/10.15252/embj.2021107795
Kaiser VB, Taylor MS, Semple CA (2016) Mutational biases drive elevated rates of substitution at regulatory sites across cancer types. PLoS Genet 12:e1006207. https://doi.org/10.1371/journal.pgen.1006207
Pihlajamaa P, Kauko O, Sahu B et al (2023) A competitive precision CRISPR method to identify the fitness effects of transcription factor binding sites. Nat Biotechnol 41:197–203. https://doi.org/10.1038/s41587-022-01444-6
Sahu B, Hartonen T, Pihlajamaa P et al (2022) Sequence determinants of human gene regulatory elements. Nat Genet 54:283–294. https://doi.org/10.1038/s41588-021-01009-4
Gates LA, Foulds CE, O’Malley BW (2017) Histone marks in the “Driver’s Seat”: functional roles in steering the transcription cycle. Trends Biochem Sci 42:977–989. https://doi.org/10.1016/j.tibs.2017.10.004
Valencia AM, Kadoch C (2019) Chromatin regulatory mechanisms and therapeutic opportunities in cancer. Nat Cell Biol 21:152–161. https://doi.org/10.1038/s41556-018-0258-1
Das ND, Chang J-C, Hon C-C et al (2023) Defining super-enhancers by highly ranked histone H4 multi-acetylation levels identifies transcription factors associated with glioblastoma stem-like properties. BMC Genom 24:574. https://doi.org/10.1186/s12864-023-09659-w
Doni Jayavelu N, Jajodia A, Mishra A, Hawkins RD (2020) Candidate silencer elements for the human and mouse genomes. Nat Commun 11:1061. https://doi.org/10.1038/s41467-020-14853-5
ENCODE Project Consortium (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489:57–74. https://doi.org/10.1038/nature11247
ENCODE Project Consortium (2014) An atlas of active enhancers across human cell types and tissues. Nature 507:455–461. https://doi.org/10.1038/nature12787
Haberle V, Stark A (2018) Eukaryotic core promoters and the functional basis of transcription initiation. Nat Rev Mol Cell Biol 19:621–637. https://doi.org/10.1038/s41580-018-0028-8
Landry J-R, Mager DL, Wilhelm BT (2003) Complex controls: the role of alternative promoters in mammalian genomes. Trends Genet 19:640–648. https://doi.org/10.1016/j.tig.2003.09.014
Benveniste D, Sonntag H-J, Sanguinetti G, Sproul D (2014) Transcription factor binding predicts histone modifications in human cell lines. Proc Natl Acad Sci U S A 111:13367–13372. https://doi.org/10.1073/pnas.1412081111
Blackwood EM, Kadonaga JT (1998) Going the distance: a current view of enhancer action. Science 281:60–63. https://doi.org/10.1126/science.281.5373.60
Bonifer C, Cockerill PN (2017) Chromatin priming of genes in development: Concepts, mechanisms and consequences. Exp Hematol 49:1–8. https://doi.org/10.1016/j.exphem.2017.01.003
Bulger M, Groudine M (2011) Functional and mechanistic diversity of distal transcription enhancers. Cell 144:327–339. https://doi.org/10.1016/j.cell.2011.01.024
Wu J BM (2018) Chapter 2 - Epigenetics and Epigenomics. In: Hematology (Seventh Edition): Elsevier. pp 17–24
Whyte WA, Orlando DA, Hnisz D et al (2013) Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153:307–319. https://doi.org/10.1016/j.cell.2013.03.035
Wang X, Cairns MJ, Yan J (2019) Super-enhancers in transcriptional regulation and genome organization. Nucleic Acids Res 47:11481–11496. https://doi.org/10.1093/nar/gkz1038
Saint-André V, Federation AJ, Lin CY et al (2016) Models of human core transcriptional regulatory circuitries. Genome Res 26:385–396. https://doi.org/10.1101/gr.197590.115
Jiang Y, Jiang Y-Y, Lin D-C (2021) Super-enhancer-mediated core regulatory circuitry in human cancer. Comput Struct Biotechnol J 19:2790–2795. https://doi.org/10.1016/j.csbj.2021.05.006
Li G-H, Qu Q, Qi T-T et al (2021) Super-enhancers: a new frontier for epigenetic modifiers in cancer chemoresistance. J Exp Clin Cancer Res 40:174. https://doi.org/10.1186/s13046-021-01974-y
Ogbourne S, Antalis TM (1998) Transcriptional control and the role of silencers in transcriptional regulation in eukaryotes. Biochem J 331;( Pt 1):1–14. https://doi.org/10.1042/bj3310001
Erceg J, Pakozdi T, Marco-Ferreres R et al (2017) Dual functionality of cis-regulatory elements as developmental enhancers and Polycomb response elements. Genes Dev 31:590–602. https://doi.org/10.1101/gad.292870.116
Huang D, Ovcharenko I (2022) Enhancer-silencer transitions in the human genome. Genome Res 32:437–448. https://doi.org/10.1101/gr.275992.121
Swygert SG, Kim S, Wu X et al (2019) Condensin-dependent chromatin compaction represses transcription globally during quiescence. Mol Cell 73:533-546.e4. https://doi.org/10.1016/j.molcel.2018.11.020
Bintu B, Mateo LJ, Su J-H et al (2018) Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science. https://doi.org/10.1126/science.aau1783
Tena JJ, Santos-Pereira JM (2021) Topologically associating domains and regulatory landscapes in development, evolution and disease. Front cell Dev Biol 9:702787. https://doi.org/10.3389/fcell.2021.702787
Sesé B, Ensenyat-Mendez M, Iñiguez S et al (2021) Chromatin insulation dynamics in glioblastoma: challenges and future perspectives of precision oncology. Clin Epigenet 13:150. https://doi.org/10.1186/s13148-021-01139-w
Ong C-T, Corces VG (2014) CTCF: an architectural protein bridging genome topology and function. Nat Rev Genet 15:234–246. https://doi.org/10.1038/nrg3663
Xu J, Huo D, Chen Y et al (2010) CpG island methylation affects accessibility of the proximal BRCA1 promoter to transcription factors. Breast Cancer Res Treat 120:593–601. https://doi.org/10.1007/s10549-009-0422-1
Renaud S, Loukinov D, Alberti L et al (2011) BORIS/CTCFL-mediated transcriptional regulation of the hTERT telomerase gene in testicular and ovarian tumor cells. Nucleic Acids Res 39:862–873. https://doi.org/10.1093/nar/gkq827
Edwards SL, Beesley J, French JD, Dunning AM (2013) Beyond GWASs: illuminating the dark road from association to function. Am J Hum Genet 93:779–797. https://doi.org/10.1016/j.ajhg.2013.10.012
Khurana E, Fu Y, Chakravarty D et al (2016) Role of non-coding sequence variants in cancer. Nat Rev Genet 17:93–108. https://doi.org/10.1038/nrg.2015.17
Mazrooei P, Kron KJ, Zhu Y et al (2019) Cistrome partitioning reveals convergence of somatic mutations and risk variants on master transcription regulators in primary prostate tumors. Cancer Cell 36:674-689.e6. https://doi.org/10.1016/j.ccell.2019.10.005
Zhang X, Meyerson M (2020) Illuminating the noncoding genome in cancer. Nat cancer 1:864–872. https://doi.org/10.1038/s43018-020-00114-3
Elliott K, Larsson E (2021) Non-coding driver mutations in human cancer. Nat Rev Cancer 21:500–509. https://doi.org/10.1038/s41568-021-00371-z
Stratton MR, Campbell PJ, Futreal PA (2009) The cancer genome. Nature 458:719–724. https://doi.org/10.1038/nature07943
Castro-Giner F, Ratcliffe P, Tomlinson I (2015) The mini-driver model of polygenic cancer evolution. Nat Rev Cancer 15:680–685. https://doi.org/10.1038/nrc3999
Wang Y, Ma R, Liu B et al (2020) SNP rs17079281 decreases lung cancer risk through creating an YY1-binding site to suppress DCBLD1 expression. Oncogene 39:4092–4102. https://doi.org/10.1038/s41388-020-1278-4
Schonfeld M, Zhao J, Komatz A et al (2020) The polymorphism rs975484 in the protein arginine methyltransferase 1 gene modulates expression of immune checkpoint genes in hepatocellular carcinoma. J Biol Chem 295:7126–7137. https://doi.org/10.1074/jbc.RA120.013401
Gamble LD, Purgato S, Henderson MJ et al (2021) A G316A polymorphism in the ornithine decarboxylase gene promoter modulates MYCN-driven childhood neuroblastoma. Cancers (Basel). https://doi.org/10.3390/cancers13081807
Avitabile M, Lasorsa VA, Cantalupo S et al (2020) Association of PARP1 polymorphisms with response to chemotherapy in patients with high-risk neuroblastoma. J Cell Mol Med 24:4072–4081. https://doi.org/10.1111/jcmm.15058
Jin Y, Wang H, Han W et al (2016) Single nucleotide polymorphism rs11669203 in TGFBR3L is associated with the risk of neuroblastoma in a Chinese population. Tumour Biol J Int Soc Oncodevelopmental Biol Med 37:3739–3747. https://doi.org/10.1007/s13277-015-4192-6
Zhou Y-T, Zheng L-Y, Wang Y-J et al (2020) Effect of functional variant rs11466313 on breast cancer susceptibility and TGFB1 promoter activity. Breast Cancer Res Treat 184:237–248. https://doi.org/10.1007/s10549-020-05841-w
Chen Q, Deng X, Hu X et al (2019) Breast cancer risk-associated SNPs in the mtor promoter form De Novo KLF5- and ZEB1-binding sites that influence the cellular response to paclitaxel. Mol Cancer Res 17:2244–2256. https://doi.org/10.1158/1541-7786.MCR-18-1072
Chen L, Liang Y, Qiu J et al (2013) Significance of rs1271572 in the estrogen receptor beta gene promoter and its correlation with breast cancer in a southwestern Chinese population. J Biomed Sci 20:32. https://doi.org/10.1186/1423-0127-20-32
Gansmo LB, Bjørnslett M, Halle MK et al (2017) MDM2 promoter polymorphism del1518 (rs3730485) and its impact on endometrial and ovarian cancer risk. BMC Cancer 17:97. https://doi.org/10.1186/s12885-017-3094-y
Barak Y, Gottlieb E, Juven-Gershon T, Oren M (1994) Regulation of mdm2 expression by p53: alternative promoters produce transcripts with nonidentical translation potential. Genes Dev 8:1739–1749. https://doi.org/10.1101/gad.8.15.1739
Haupt Y, Maya R, Kazaz A, Oren M (1997) Mdm2 promotes the rapid degradation of p53. Nature 387:296–299. https://doi.org/10.1038/387296a0
Bond GL, Hu W, Bond EE et al (2004) A single nucleotide polymorphism in the MDM2 promoter attenuates the p53 tumor suppressor pathway and accelerates tumor formation in humans. Cell 119:591–602. https://doi.org/10.1016/j.cell.2004.11.022
Yang Z-H, Zhou C-L, Zhu H et al (2014) A functional SNP in the MDM2 promoter mediates E2F1 affinity to modulate cyclin D1 expression in tumor cell proliferation. Asian Pac J Cancer Prev 15:3817–3823. https://doi.org/10.7314/apjcp.2014.15.8.3817
Knappskog S, Bjørnslett M, Myklebust LM et al (2011) The MDM2 promoter SNP285C/309G haplotype diminishes Sp1 transcription factor binding and reduces risk for breast and ovarian cancer in Caucasians. Cancer Cell 19:273–282. https://doi.org/10.1016/j.ccr.2010.12.019
Okamoto K, Tsunematsu R, Tahira T et al (2015) SNP55, a new functional polymorphism of MDM2-P2 promoter, contributes to allele-specific expression of MDM2 in endometrial cancers. BMC Med Genet 16:67. https://doi.org/10.1186/s12881-015-0216-8
Smith KS, Yadav VK, Pedersen BS et al (2015) Signatures of accelerated somatic evolution in gene promoters in multiple cancer types. Nucleic Acids Res 43:5307–5317. https://doi.org/10.1093/nar/gkv419
Dietlein F, Wang AB, Fagre C et al (2022) Genome-wide analysis of somatic noncoding mutation patterns in cancer. Science. https://doi.org/10.1126/science.abg5601
Colebatch AJ, Di Stefano L, Wong SQ, et al (2016) Clustered somatic mutations are frequent in transcription factor binding motifs within proximal promoter regions in melanoma and other cutaneous malignancies. Oncotarget 7:66569–66585. https://doi.org/10.18632/oncotarget.11892
Gupta S, Vanderbilt CM, Lin Y-T et al (2021) A pan-cancer study of somatic TERT promoter mutations and amplification in 30,773 tumors profiled by clinical genomic sequencing. J Mol Diagn 23:253–263. https://doi.org/10.1016/j.jmoldx.2020.11.003
Fredriksson NJ, Ny L, Nilsson JA, Larsson E (2014) Systematic analysis of noncoding somatic mutations and gene expression alterations across 14 tumor types. Nat Genet 46:1258–1263. https://doi.org/10.1038/ng.3141
Bell RJA, Rube HT, Kreig A et al (2015) Cancer. The transcription factor GABP selectively binds and activates the mutant TERT promoter in cancer. Science 348:1036–1039. https://doi.org/10.1126/science.aab0015
Huang FW, Hodis E, Xu MJ et al (2013) Highly recurrent TERT promoter mutations in human melanoma. Science 339:957–959. https://doi.org/10.1126/science.1229259
Man J, Shoemake J, Zhou W et al (2014) Sema3C promotes the survival and tumorigenicity of glioma stem cells through Rac1 activation. Cell Rep 9:1812–1826. https://doi.org/10.1016/j.celrep.2014.10.055
Sakthikumar S, Roy A, Haseeb L et al (2020) Whole-genome sequencing of glioblastoma reveals enrichment of non-coding constraint mutations in known and novel genes. Genome Biol 21:127. https://doi.org/10.1186/s13059-020-02035-x
Rheinbay E, Parasuraman P, Grimsby J et al (2017) Recurrent and functional regulatory mutations in breast cancer. Nature 547:55–60. https://doi.org/10.1038/nature22992
Seachrist DD, Anstine LJ, Keri RA (2021) FOXA1: a pioneer of nuclear receptor action in breast cancer. Cancers (Basel). https://doi.org/10.3390/cancers13205205
Fu X, Jeselsohn R, Pereira R et al (2016) FOXA1 overexpression mediates endocrine resistance by altering the ER transcriptome and IL-8 expression in ER-positive breast cancer. Proc Natl Acad Sci USA 113:E6600–E6609. https://doi.org/10.1073/pnas.1612835113
Jeselsohn R, Barry WT, Migliaccio I et al (2016) TransCONFIRM: identification of a genetic signature of response to fulvestrant in advanced hormone receptor-positive breast cancer. Clin Cancer Res Off J Am Assoc Cancer Res 22:5755–5764. https://doi.org/10.1158/1078-0432.CCR-16-0148
Zhang T, Xu M, Makowski MM et al (2017) SDHD promoter mutations ablate GABP transcription factor binding in melanoma. Cancer Res 77:1649–1661. https://doi.org/10.1158/0008-5472.CAN-16-0919
Lowdon RF, Wang T (2017) Epigenomic annotation of noncoding mutations identifies mutated pathways in primary liver cancer. PLoS ONE 12:e0174032. https://doi.org/10.1371/journal.pone.0174032
Lawrenson K, Song F, Hazelett DJ et al (2019) Genome-wide association studies identify susceptibility loci for epithelial ovarian cancer in east Asian women. Gynecol Oncol 153:343–355. https://doi.org/10.1016/j.ygyno.2019.02.023
Jones MR, Peng P-C, Coetzee SG et al (2020) Ovarian cancer risk variants are enriched in histotype-specific enhancers and disrupt transcription factor binding sites. Am J Hum Genet 107:622–635. https://doi.org/10.1016/j.ajhg.2020.08.021
Yu C-Y, Han J-X, Zhang J et al (2020) A 16q22.1 variant confers susceptibility to colorectal cancer as a distal regulator of ZFP90. Oncogene 39:1347–1360. https://doi.org/10.1038/s41388-019-1055-4
Yan H, Tian S, Kleinstern G et al (2020) Chronic lymphocytic leukemia (CLL) risk is mediated by multiple enhancer variants within CLL risk loci. Hum Mol Genet 29:2761–2774. https://doi.org/10.1093/hmg/ddaa165
Shi Q, Shi Q-N, Xu J-W et al (2022) rs9390123 and rs9399451 influence the DNA repair capacity of lung cancer by regulating PEX3 and PHACTR2-AS1 expression instead of PHACTR2. Oncol Rep. https://doi.org/10.3892/or.2022.8270
Cardinale A, Cantalupo S, Lasorsa VA et al (2022) Functional annotation and investigation of the 10q24.33 melanoma risk locus identifies a common variant that influences transcriptional regulation of OBFC1. Hum Mol Genet 31:863–874. https://doi.org/10.1093/hmg/ddab293
Yanchus C, Drucker KL, Kollmeyer TM et al (2022) A noncoding single-nucleotide polymorphism at 8q24 drives IDH1-mutant glioma formation. Science 378:68–78. https://doi.org/10.1126/science.abj2890
Tian J, Lou J, Cai Y et al (2020) Risk SNP-mediated enhancer-promoter interaction drives colorectal cancer through both FADS2 and AP002754.2. Cancer Res 80:1804–1818. https://doi.org/10.1158/0008-5472.CAN-19-2389
Cowper-Sal lari R, Zhang X, Wright JB et al (2012) Breast cancer risk-associated SNPs modulate the affinity of chromatin for FOXA1 and alter gene expression. Nat Genet 44:1191–1198. https://doi.org/10.1038/ng.2416
Bailey SD, Desai K, Kron KJ et al (2016) Noncoding somatic and inherited single-nucleotide variants converge to promote ESR1 expression in breast cancer. Nat Genet 48:1260–1266. https://doi.org/10.1038/ng.3650
Ghoussaini M, French JD, Michailidou K et al (2016) Evidence that the 5p12 variant rs10941679 confers susceptibility to estrogen-receptor-positive breast cancer through FGF10 and MRPS30 regulation. Am J Hum Genet 99:903–911. https://doi.org/10.1016/j.ajhg.2016.07.017
Li X, Xu X, Fang J, et al (2016) Rs2853677 modulates Snail1 binding to the TERT enhancer and affects lung adenocarcinoma susceptibility. Oncotarget 7:37825–37838. https://doi.org/10.18632/oncotarget.9339
Corona RI, Seo J-H, Lin X et al (2020) Non-coding somatic mutations converge on the PAX8 pathway in ovarian cancer. Nat Commun 11:2020. https://doi.org/10.1038/s41467-020-15951-0
Song H, Liu Y, Tan Y et al (2022) Recurrent noncoding somatic and germline WT1 variants converge to disrupt MYB binding in acute promyelocytic leukemia. Blood 140:1132–1144. https://doi.org/10.1182/blood.2021014945
Hnisz D, Abraham BJ, Lee TI et al (2013) Super-enhancers in the control of cell identity and disease. Cell 155:934–947. https://doi.org/10.1016/j.cell.2013.09.053
Cong Z, Li Q, Yang Y et al (2019) The SNP of rs6854845 suppresses transcription via the DNA looping structure alteration of super-enhancer in colon cells. Biochem Biophys Res Commun 514:734–741. https://doi.org/10.1016/j.bbrc.2019.04.190
Ke J, Tian J, Mei S et al (2020) Genetic predisposition to colon and rectal adenocarcinoma is mediated by a super-enhancer polymorphism coactivating CD9 and PLEKHG6. Cancer Epidemiol Biomarkers Prev a Publ Am Assoc Cancer Res Cosponsored by Am Soc Prev Oncol 29:850–859. https://doi.org/10.1158/1055-9965.EPI-19-1116
Kleinstern G, Yan H, Hildebrandt MAT et al (2020) Inherited variants at 3q13.33 and 3p24.1 are associated with risk of diffuse large B-cell lymphoma and implicate immune pathways. Hum Mol Genet 29:70–79. https://doi.org/10.1093/hmg/ddz228
Oldridge DA, Wood AC, Weichert-Leahey N et al (2015) Genetic predisposition to neuroblastoma mediated by a LMO1 super-enhancer polymorphism. Nature 528:418–421. https://doi.org/10.1038/nature15540
Kandaswamy R, Sava GP, Speedy HE et al (2016) Genetic predisposition to chronic lymphocytic leukemia is mediated by a BMF super-enhancer polymorphism. Cell Rep 16:2061–2067. https://doi.org/10.1016/j.celrep.2016.07.053
Mansour MR, Abraham BJ, Anders L et al (2014) Oncogene regulation. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science 346:1373–1377. https://doi.org/10.1126/science.1259037
Pinoli P, Stamoulakatou E, Nguyen A-P et al (2020) Pan-cancer analysis of somatic mutations and epigenetic alterations in insulated neighbourhood boundaries. PLoS One 15:e0227180. https://doi.org/10.1371/journal.pone.0227180
Dai J, Zhu M, Wang C et al (2015) Systematical analyses of variants in CTCF-binding sites identified a novel lung cancer susceptibility locus among Chinese population. Sci Rep 5:7833. https://doi.org/10.1038/srep07833
Liu Y, Walavalkar NM, Dozmorov MG et al (2017) Identification of breast cancer associated variants that modulate transcription factor binding. PLoS Genet 13:e1006761. https://doi.org/10.1371/journal.pgen.1006761
Poulos RC, Thoms JAI, Guan YF et al (2016) Functional mutations form at CTCF-cohesin binding sites in melanoma due to uneven nucleotide excision repair across the motif. Cell Rep 17:2865–2872. https://doi.org/10.1016/j.celrep.2016.11.055
Guo YA, Chang MM, Huang W et al (2018) Mutation hotspots at CTCF binding sites coupled to chromosomal instability in gastrointestinal cancers. Nat Commun 9:1520. https://doi.org/10.1038/s41467-018-03828-2
Umer HM, Cavalli M, Dabrowski MJ et al (2016) A significant regulatory mutation burden at a high-affinity position of the CTCF motif in gastrointestinal cancers. Hum Mutat 37:904–913. https://doi.org/10.1002/humu.23014
Vogelstein B, Papadopoulos N, Velculescu VE et al (2013) Cancer genome landscapes. Science 339:1546–1558. https://doi.org/10.1126/science.1235122
Guo Y, Perez AA, Hazelett DJ et al (2018) CRISPR-mediated deletion of prostate cancer risk-associated CTCF loop anchors identifies repressive chromatin loops. Genome Biol 19:160. https://doi.org/10.1186/s13059-018-1531-0
Sidiropoulos N, Mardin BR, Rodríguez-González FG et al (2022) Somatic structural variant formation is guided by and influences genome architecture. Genome Res 32:643–655. https://doi.org/10.1101/gr.275790.121
Jablonski KP, Carron L, Mozziconacci J et al (2022) Contribution of 3D genome topological domains to genetic risk of cancers: a genome-wide computational study. Hum Genom 16:2. https://doi.org/10.1186/s40246-022-00375-2
Osman N, Shawky A-E-M, Brylinski M (2022) Exploring the effects of genetic variation on gene regulation in cancer in the context of 3D genome structure. BMC Genom Data 23:13. https://doi.org/10.1186/s12863-021-01021-x
Chyr J, Guo D, Zhou X (2018) LSCC SNP variant regulates SOX2 modulation of VDAC3. Oncotarget 9:22340–22352. https://doi.org/10.18632/oncotarget.24918
Zou D, Zhang H, Ke J et al (2020) Three functional variants were identified to affect RPS24 expression and significantly associated with risk of colorectal cancer. Arch Toxicol 94:295–303. https://doi.org/10.1007/s00204-019-02600-9
Mei S, Ke J, Tian J et al (2019) A functional variant in the boundary of a topological association domain is associated with pancreatic cancer risk. Mol Carcinog 58:1855–1862. https://doi.org/10.1002/mc.23077
Painter JN, Kaufmann S, O’Mara TA et al (2016) A common variant at the 14q32 endometrial cancer risk locus activates AKT1 through YY1 binding. Am J Hum Genet 98:1159–1169. https://doi.org/10.1016/j.ajhg.2016.04.012
Tam V, Patel N, Turcotte M et al (2019) Benefits and limitations of genome-wide association studies. Nat Rev Genet 20:467–484. https://doi.org/10.1038/s41576-019-0127-1
Shigemizu D, Fujimoto A, Akiyama S et al (2013) A practical method to detect SNVs and indels from whole genome and exome sequencing data. Sci Rep 3:2161. https://doi.org/10.1038/srep02161
Ledergerber C, Dessimoz C (2011) Base-calling for next-generation sequencing platforms. Brief Bioinform 12:489–497. https://doi.org/10.1093/bib/bbq077
Luedtke A, Powers S, Petersen A et al (2011) Evaluating methods for the analysis of rare variants in sequence data. BMC Proc 5(Suppl 9):S119. https://doi.org/10.1186/1753-6561-5-S9-S119
Sullivan PF, Meadows JRS, Gazal S, et al (2023) Leveraging base pair mammalian constraint to understand genetic variation and human disease. Science (80–) 380:6643. https://doi.org/10.1126/science.abn2937
Ying Y-L, Hu Z-L, Zhang S et al (2022) Nanopore-based technologies beyond DNA sequencing. Nat Nanotechnol 17:1136–1146. https://doi.org/10.1038/s41565-022-01193-2
Hasanau T, Pisarev E, Kisil O, et al (2022) Detection of TERT promoter mutations as a prognostic biomarker in gliomas: methodology, prospects, and advances. Biomedicines https://doi.org/10.3390/biomedicines10030728
Quan P-L, Sauzade M, Brouzes E (2018) dPCR: a technology review. Sensors (Basel). https://doi.org/10.3390/s18041271
Yan J, Qiu Y, Ribeiro Dos Santos AM et al (2021) Systematic analysis of binding of transcription factors to noncoding variants. Nature 591:147–151. https://doi.org/10.1038/s41586-021-03211-0
Grishin D, Gusev A (2022) Allelic imbalance of chromatin accessibility in cancer identifies candidate causal risk variants and their mechanisms. Nat Genet 54:837–849. https://doi.org/10.1038/s41588-022-01075-2
Tewhey R, Kotliar D, Park DS et al (2016) Direct identification of hundreds of expression-modulating variants using a multiplexed reporter assay. Cell 165:1519–1529. https://doi.org/10.1016/j.cell.2016.04.027
Bin KS, Ernst J (2018) Investigating enhancer evolution with massively parallel reporter assays. Genome Biol 19:114. https://doi.org/10.1186/s13059-018-1502-5
Ulirsch JC, Nandakumar SK, Wang L et al (2016) Systematic functional dissection of common genetic variation affecting red blood cell traits. Cell 165:1530–1545. https://doi.org/10.1016/j.cell.2016.04.048
Patwardhan RP, Hiatt JB, Witten DM et al (2012) Massively parallel functional dissection of mammalian enhancers in vivo. Nat Biotechnol 30:265–270. https://doi.org/10.1038/nbt.2136
Zheng Y, VanDusen NJ (2023) Massively parallel reporter assays for high-throughput in vivo analysis of cis-regulatory elements. J Cardiovasc Dev Dis. https://doi.org/10.3390/jcdd10040144
Morova T, Ding Y, Huang C-CF et al (2023) Optimized high-throughput screening of non-coding variants identified from genome-wide association studies. Nucleic Acids Res 51:e18. https://doi.org/10.1093/nar/gkac1198
Wang X, He L, Goggin SM et al (2018) High-resolution genome-wide functional dissection of transcriptional regulatory regions and nucleotides in human. Nat Commun 9:5380. https://doi.org/10.1038/s41467-018-07746-1
Arnold CD, Gerlach D, Stelzer C et al (2013) Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339:1074–1077. https://doi.org/10.1126/science.1232542
Korkmaz G, Lopes R, Ugalde AP et al (2016) Functional genetic screens for enhancer elements in the human genome using CRISPR-Cas9. Nat Biotechnol 34:192–198. https://doi.org/10.1038/nbt.3450
Sanjana NE, Wright J, Zheng K et al (2016) High-resolution interrogation of functional elements in the noncoding genome. Science 353:1545–1549. https://doi.org/10.1126/science.aaf7613
Martin-Rufino JD, Castano N, Pang M et al (2023) Massively parallel base editing to map variant effects in human hematopoiesis. Cell 186:2456-2474.e24. https://doi.org/10.1016/j.cell.2023.03.035
Eleveld TF, Bakali C, Eijk PP et al (2021) Engineering large-scale chromosomal deletions by CRISPR-Cas9. Nucleic Acids Res 49:12007–12016. https://doi.org/10.1093/nar/gkab557
Peña-Martínez EG, Rivera-Madera A, Pomales-Matos DA et al (2023) Disease-associated non-coding variants alter NKX2-5 DNA-binding affinity. Biochim Biophys acta Gene Regul Mech 1866:194906. https://doi.org/10.1016/j.bbagrm.2023.194906
Xia Q, Deliard S, Yuan C-X et al (2015) Characterization of the transcriptional machinery bound across the widely presumed type 2 diabetes causal variant, rs7903146, within TCF7L2. Eur J Hum Genet 23:103–109. https://doi.org/10.1038/ejhg.2014.48
Chen X, Shen Y, Draper W et al (2016) ATAC-see reveals the accessible genome by transposase-mediated imaging and sequencing. Nat Methods 13:1013–1020. https://doi.org/10.1038/nmeth.4031
Schmidl C, Rendeiro AF, Sheffield NC, Bock C (2015) ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat Methods 12:963–965. https://doi.org/10.1038/nmeth.3542
Hainer SJ, Fazzio TG (2019) High-resolution chromatin profiling using CUT&RUN. Curr Protoc Mol Biol 126:e85. https://doi.org/10.1002/cpmb.85
Kaya-Okur HS, Janssens DH, Henikoff JG et al (2020) Efficient low-cost chromatin profiling with CUT&Tag. Nat Protoc 15:3264–3283. https://doi.org/10.1038/s41596-020-0373-x
Pudjihartono M, Perry JK, Print C et al (2022) Interpretation of the role of germline and somatic non-coding mutations in cancer: expression and chromatin conformation informed analysis. Clin Epigenetics 14:120. https://doi.org/10.1186/s13148-022-01342-3
Zhao Z, Tavoosidana G, Sjölinder M et al (2006) Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat Genet 38:1341–1347. https://doi.org/10.1038/ng1891
Ferraiuolo MA, Sanyal A, Naumova N et al (2012) From cells to chromatin: capturing snapshots of genome organization with 5C technology. Methods 58:255–267. https://doi.org/10.1016/j.ymeth.2012.10.011
Kong S, Zhang Y (2019) Deciphering Hi-C: from 3D genome to function. Cell Biol Toxicol 35:15–32. https://doi.org/10.1007/s10565-018-09456-2
Schoenfelder S, Javierre B-M, Furlan-Magaril M, et al (2018) Promoter Capture Hi-C: High-resolution, Genome-wide Profiling of Promoter Interactions. J Vis Exp. https://doi.org/10.3791/57320
Mumbach MR, Rubin AJ, Flynn RA et al (2016) HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat Methods 13:919–922. https://doi.org/10.1038/nmeth.3999
Nagano T, Várnai C, Schoenfelder S et al (2015) Comparison of Hi-C results using in-solution versus in-nucleus ligation. Genome Biol 16:175. https://doi.org/10.1186/s13059-015-0753-7
Li G, Cai L, Chang H et al (2014) Chromatin interaction analysis with paired-end tag (ChIA-PET) sequencing technology and application. BMC Genom 15(Suppl 1):S11. https://doi.org/10.1186/1471-2164-15-S12-S11
Martincorena I, Campbell PJ (2015) Somatic mutation in cancer and normal cells. Science 349:1483–1489. https://doi.org/10.1126/science.aab4082
Lei Y, Tang R, Xu J et al (2021) Applications of single-cell sequencing in cancer research: progress and perspectives. J Hematol Oncol 14:91. https://doi.org/10.1186/s13045-021-01105-2
Yen JL, Garcia S, Montana A et al (2017) A variant by any name: quantifying annotation discordance across tools and clinical databases. Genome Med 9:7. https://doi.org/10.1186/s13073-016-0396-7
Wells A, Heckerman D, Torkamani A et al (2019) Ranking of non-coding pathogenic variants and putative essential regions of the human genome. Nat Commun 10:5241. https://doi.org/10.1038/s41467-019-13212-3
Lee D, Gorkin DU, Baker M et al (2015) A method to predict the impact of regulatory variants from DNA sequence. Nat Genet 47:955–961. https://doi.org/10.1038/ng.3331
Dong S, Boyle AP (2022) Prioritization of regulatory variants with tissue-specific function in the non-coding regions of human genome. Nucleic Acids Res 50:e6. https://doi.org/10.1093/nar/gkab924
Lou S, Cotter KA, Li T et al (2019) GRAM: a GeneRAlized model to predict the molecular effect of a non-coding variant in a cell-type specific manner. PLoS Genet 15:e1007860. https://doi.org/10.1371/journal.pgen.1007860
McLaren W, Gil L, Hunt SE, HS, Ritchie GRS, Thormann A, Flicek P, Cunningham F (2016) The ensembl variant effect predictor. Genome Biol 17(1):122. https://doi.org/10.1186/s13059-016-0974-4
Fu Y, Liu Z, Lou S, Bedford J, Mu XJ, Yip KY, Khurana E, Gerstein M (2014) FunSeq2: a framework for prioritizing noncoding regulatory variants in cancer. Genome Biol 15:480. https://doi.org/10.1186/s13059-014-0480-5
Xie X, Ma W, Songyang Z, Luo Z, Huang J, Dai Z, Xiong Y (2016) CCSI: a database providing chromatin–chromatin spatial interaction information. Database 2016:bav124. https://doi.org/10.1093/database/bav124
Huang D, Yi X, Zhang S, Zheng Z, Wang P, Xuan C, Sham PC, Wang J, Li MJ (2018) GWAS4D: multidimensional analysis of context-specific regulatory variant for human complex diseases and traits. Nucleic Acid Res 46(1):114–120. https://doi.org/10.1093/nar/gky407
Li X, Shi L, Wang Y, Zhong J, Zhao X, Teng H, Shi X, Yang H, Ruan S, Li M, Sun ZS, Zhan Q, Mao F (2019) OncoBase: a platform for decoding regulatory somatic mutations in human cancers. Nucleic Acid Res 47(1):1044–1055. https://doi.org/10.1093/nar/gky1139
Lee PH, Lee C, Li X et al (2018) Principles and methods of in-silico prioritization of non-coding regulatory variants. Hum Genet 137:15–30. https://doi.org/10.1007/s00439-017-1861-0
Wang Z, Zhao G, Li B et al (2022) Performance comparison of computational methods for the prediction of the function and pathogenicity of non-coding variants. Genom Proteom Bioinform. https://doi.org/10.1016/j.gpb.2022.02.002
Wang Y, Chen Y, Li C et al (2022) TERT promoter revertant mutation inhibits melanoma growth through intrinsic apoptosis. Biology (Basel). https://doi.org/10.3390/biology11010141
Eufrásio A, Perrod C, Ferreira FJ et al (2020) In vivo reporter assays uncover changes in enhancer activity caused by type 2 diabetes-associated single nucleotide polymorphisms. Diabetes 69:2794–2805. https://doi.org/10.2337/db19-1049
Liu H, Duncan K, Helverson A et al (2020) Analysis of zebrafish periderm enhancers facilitates identification of a regulatory variant near human KRT8/18. Elife. https://doi.org/10.7554/eLife.51325
Kvon EZ, Zhu Y, Kelman G et al (2020) Comprehensive in vivo interrogation reveals phenotypic impact of human enhancer variants. Cell 180:1262-1271.e15. https://doi.org/10.1016/j.cell.2020.02.031
Lambert JT, Su-Feher L, Cichewicz K et al (2021) Parallel functional testing identifies enhancers active in early postnatal mouse brain. Elife. https://doi.org/10.7554/eLife.69479
Cui S, Wu Q, Liu M et al (2021) EphA2 super-enhancer promotes tumor progression by recruiting FOSL2 and TCF7L2 to activate the target gene EphA2. Cell Death Dis 12:264. https://doi.org/10.1038/s41419-021-03538-6
Amândio AR, Beccari L, Lopez-Delisle L et al (2021) Sequential in cis mutagenesis in vivo reveals various functions for CTCF sites at the mouse HoxD cluster. Genes Dev 35:1490–1509. https://doi.org/10.1101/gad.348934.121
Anania C, Acemel RD, Jedamzick J et al (2022) In vivo dissection of a clustered-CTCF domain boundary reveals developmental principles of regulatory insulation. Nat Genet 54:1026–1036. https://doi.org/10.1038/s41588-022-01117-9
Bauer DE, Kamran SC, Lessard S et al (2013) An erythroid enhancer of BCL11A subject to genetic variation determines fetal hemoglobin level. Science 342:253–257. https://doi.org/10.1126/science.1242088
Frangoul H, Altshuler D, Cappellini MD et al (2021) CRISPR-Cas9 gene editing for Sickle cell disease and β-Thalassemia. N Engl J Med 384:252–260. https://doi.org/10.1056/NEJMoa2031054
Fu B, Liao J, Chen S et al (2022) CRISPR-Cas9-mediated gene editing of the BCL11A enhancer for pediatric β(0)/β(0) transfusion-dependent β-thalassemia. Nat Med 28:1573–1580. https://doi.org/10.1038/s41591-022-01906-z
Doudna JA (2020) The promise and challenge of therapeutic genome editing. Nature 578:229–236. https://doi.org/10.1038/s41586-020-1978-5
Whale AS, Jones GM, Pavšič J et al (2018) Assessment of digital pcr as a primary reference measurement procedure to support advances in precision medicine. Clin Chem 64:1296–1307. https://doi.org/10.1373/clinchem.2017.285478
Funding
This study was supported by the Instituto de la Salud Carlos III Miguel Servet Project (#CP17/00188 to D.M.M.), AES2019 (#I19/01514 to D.M.M.) and Sara Borrell project (#CD22/00026), the Institut d’Investigació Sanitària Illes Balears (IdISBa), the Scientific Foundation of the Spanish Association Against Cancer (M.E.M. and A.F.B.-L) Foundation, the Balearic Islands Government FPI program (#FPI/037/2021 to S.Í.-M), and the “CONTIGO Contra el Cancer de Mujer” foundation (#MERIT project).
Author information
Authors and Affiliations
Contributions
All authors listed a relevant, direct, and intellectual contribution to the work and approved the final version of the manuscript for publication. S.Í.-M. wrote the original draft and designed the figures. K.F-N, A.R, J.I.J.O., J.C., and M.L.D. provided significant inputs and feedback for the whole manuscript. M.E.-M., A.F.B.-L, and P.L.-A. actively participated in draft editing and review. D.M.M. provided the main guidelines and supervised the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that this research was conducted in the absence of any commercial or financial relationship that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Iñiguez-Muñoz, S., Llinàs-Arias, P., Ensenyat-Mendez, M. et al. Hidden secrets of the cancer genome: unlocking the impact of non-coding mutations in gene regulatory elements. Cell. Mol. Life Sci. 81, 274 (2024). https://doi.org/10.1007/s00018-024-05314-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-024-05314-z