Abstract
Recent advances in RNA sequencing technologies helped uncover what was once uncharted territory in the human genome—the complex and versatile world of long noncoding RNAs (lncRNAs). Previously thought of as merely transcriptional “noise”, lncRNAs have now emerged as essential regulators of gene expression networks controlling development, homeostasis and disease progression. The regulatory functions of lncRNAs are broad and diverse, and the underlying molecular mechanisms are highly variable, acting at the transcriptional, post-transcriptional, translational, and post-translational levels. In recent years, evidence has accumulated to support the important role of lncRNAs in the development and functioning of the lymphatic vasculature and associated pathological processes such as tumor-induced lymphangiogenesis and cancer metastasis. In this review, we summarize the current knowledge on the role of lncRNAs in regulating the key genes and pathways involved in lymphatic vascular development and disease. Furthermore, we discuss the potential of lncRNAs as novel therapeutic targets and outline possible strategies for the development of lncRNA-based therapeutics to treat diseases of the lymphatic system.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
About three quarters of the human genome can be transcribed [1], giving rise to millions of transcripts, most of which appear nonfunctional [2]. The remaining tens of thousands of functional transcripts correspond to either canonical protein-coding genes (PCGs) or their noncoding counterparts, which are often involved in the regulation of PCG expression. Some protein-coding RNAs are bifunctional and also possess noncoding functions and vice versa [3]. Depending on their size, noncoding functional transcripts (excluding ribosomal RNAs and transfer RNAs) can be classified into two broad and heterogeneous groups—small noncoding RNAs (sncRNAs) and long noncoding RNAs (lncRNAs). Defined as non-protein-coding transcripts longer than of 200 nucleotides (an arbitrary threshold, which was recently recommended to be raised to 500 nucleotides [4]), lncRNAs share many features with mRNAs. Like mRNAs, lncRNAs are predominantly transcribed by RNA polymerase II [5, 6], are often capped and polyadenylated, and undergo splicing [7]. Once dismissed as part of transcriptional “noise”, lncRNAs have made a remarkable comeback in recent years, providing a whole new layer of complexity to gene regulation. Despite lacking protein-coding potential, lncRNAs exert diverse gene regulatory functions through a variety of molecular mechanisms at the transcriptional, post-transcriptional, translational and post-translational levels [8,9,10]. Although these mechanisms typically involve lncRNA interactions with functional biomolecules like nucleic acids or proteins, the very process of lncRNA transcription may itself regulate gene expression [11]. The versatile regulatory functions of lncRNAs are being increasingly appreciated in almost every aspect of human physiology and disease [12] including developmental processes [13, 14], cell cycle [15], differentiation [16], metabolism [17], hematopoiesis [18], X-chromosome inactivation [19], stress response [20], aging [21], apoptosis [22], inflammation [23], viral infection [24] and cancer [25, 26].
The subcellular localization of an lncRNA largely determines its biological function. LncRNAs may reside in various subcellular compartments including the nucleus, cytosol, endoplasmic reticulum and mitochondria [27]. Furthermore, lncRNAs may be secreted from the cell in exosomes or other extracellular vesicles (EVs). A number of lncRNAs localize in different subcellular compartments and may have multiple functions depending on their subcellular localization. Nuclear lncRNAs typically regulate transcriptional programs through chromatin interactions and remodeling [28, 29], often serving as scaffolds for multiprotein transcriptional and epigenetic complexes. However, nuclear lncRNAs can also perform a variety of non-chromatin-related functions such as organization of nuclear paraspeckles [20], processing of preribosomal RNA in the nucleolus [30] and regulation of alternative splicing [31]. Cytoplasmic lncRNAs, on the other hand, are mainly involved in the post-transcriptional regulation of gene expression. For example, they may interact with protein-coding mRNAs to alter their translation and/or stability [32,33,34] or act as molecular “sponges”, also known as competing endogenous RNAs (ceRNAs), to sequester microRNAs (miRNAs) from their mRNA targets [35, 36]. MiRNAs are not the only molecules that can be “sponged” in this manner: proteins may also become sequestered through their interaction with lncRNAs [37]. In addition, protein-lncRNA interactions may participate in the scaffolding of protein–protein or protein-nucleic acid complexes [38]. Yet another function of lncRNAs in the cytoplasm is to mediate signal transduction pathways by influencing protein post-translational modifications [39] or serving as stabilizing scaffolds for signal transduction proteins [40].
In addition to subcellular localization, cell lineage specificity (or the lack of it) is another important influence on the function of lncRNAs. Certain lncRNAs are specifically expressed or enriched in particular cell types, defining their phenotypes. For example, the lineage-specific lncRNA NeST (nettoie Salmonella pas Theiler’s) is induced in type 1 T helper (Th1) cells, but not in type 2 T helper (Th2) cells, and regulates the expression of interferon-gamma via an epigenetic mechanism [41, 42]. Another example is spliced-transcript endothelial-enriched lncRNA (STEEL), which is enriched in endothelial cells and activates a pro-angiogenic transcriptional program [43]. On the other hand, there are lncRNAs that are widely expressed across almost all cell types such as the metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) [44]. Some lncRNAs are associated with certain diseases such as coronary artery disease [45], or respond to environmental stimuli such as hypoxia [46]. Overall, lncRNAs are extremely diverse, both functionally and spatially within the cell, and there is hardly any physiological or pathophysiological process that is not affected by them to some degree. In this review, we will highlight the lncRNAs that are involved in lymphatic vascular development and disease.
The lymphatic vascular system is a unidirectional network of lymphatic capillaries and collecting lymphatic vessels lined by lymphatic endothelial cells (LECs), which is present in most of the body’s organs. The lymphatic vasculature drains interstitial fluid from tissues and returns it to blood circulation in the form of lymph. In addition to its central role in maintaining body fluid homeostasis [47], the lymphatic vasculature transports antigens and immune cells from peripheral tissues to lymph nodes (LNs), thereby contributing to immune surveillance [48,49,50]. Other tissue- and organ-specific functions of lymphatic vessels include dietary fat absorption in the intestine [51], antigen storage and presentation in lymphoid tissues [48], and outflow of cerebrospinal fluid in the cranial and spinal compartments of the central nervous system [52,53,54,55,56]. Structural and functional abnormalities of the lymphatic vasculature are associated with multiple diseases including lymphedema (accumulation of lymph in soft tissues) [57, 58], metabolic diseases such as obesity [59, 60] and diabetes [61, 62], chronic inflammation [63, 64], cardiovascular disease [65, 66], atherosclerosis [67,68,69], neurodegenerative diseases [70,71,72,73], glaucoma [74, 75] and Crohn’s disease [50]. Finally, the process of lymphatic vessel formation and expansion, termed lymphangiogenesis, plays an essential role in cancer progression and metastasis [49, 50, 76,77,78,79].
The present review does not attempt to cover the whole spectrum of molecular, cellular and morphological mechanisms underlying physiological and pathological lymphangiogenesis, as they have been extensively reviewed elsewhere [49, 50, 80,81,82]. Instead, we focus only on those molecular players and pathways for which regulatory mechanisms involving lncRNAs have been identified. Because the field is rapidly changing and new data constantly becomes available, the review does not aim at full coverage of all the existing literature. Nevertheless, we will discuss the most important findings showing how lncRNAs affect lymphatic vascular development, physiology and disease, and outline potential therapeutic approaches targeting lncRNAs for the amelioration of lymphatic vascular pathologies, including lymphedema and cancer metastasis.
LETR1: lymphatic vascular lineage-specific lncRNA
Lymphatic endothelial transcriptional regulator lncRNA 1 (LETR1), also known as LINC01197, was identified by Ducoli et al. [83] and represents the first, and so far only, example of LEC-specific lncRNA. The principal difference between LETR1 and the other lncRNAs discussed below is that LETR1 is specifically expressed in the lymphatic but not in the blood vessel endothelium, suggesting a unique, lineage-specific role in regulating LEC differentiation. Indeed, Ducoli et al. found that LETR1 acts as gatekeeper of the LEC transcriptome by modulating the expression of essential proliferation-related genes such as the tumor-suppressor transcription factor KLF4 [84] and genes involved in endothelial cell migration such as the secreted semaphorin SEMA3C [85, 86]. They also demonstrated that, consistent with the predominantly nuclear localization of LETR1 in LECs, LETR1 regulates gene expression through a chromatin-based epigenetic mechanism. This mechanism involves LETR1 being recruited to DNA regions near its target genes and interacting with the nucleosome remodeling factor RBBP7, a component of several histone deacetylase (HDAC) complexes including mSin3, NuRD and CoREST [87]. Another epigenetic complex containing RBBP7 is polycomb repressive complex 2 (PRC2), which is best known as an epigenetic “writer” of histone methylation associated with transcriptional repression [88]. Taken together, the findings of Ducoli et al. indicate that LETR1 regulates the expression of lymphatic lineage-specific genes by acting as a scaffold for epigenetic protein complexes [83]. Furthermore, LETR1 is the first bona fide lncRNA that could potentially serve as a lymphatic-specific biomarker.
While most of the lncRNAs discussed in this review are involved in tumor lymphangiogenesis and lymphatic metastasis, LETR1 is the first example of an lncRNA with a role in normal lymphatic vascular development. Therefore, the discovery of LETR1 marked a major paradigm shift away from the focus on the oncogenic roles of lymphatic-associated lncRNAs towards a broader consideration of the roles of lncRNAs in both physiological and pathological lymphangiogenesis.
LncRNAs as regulators of the lymphangiogenic growth factor VEGF-C
Paracrine signaling by vascular endothelial growth factor C (VEGF-C) plays a central role during lymphatic vascular development [89]. VEGF-C is a secreted ligand of vascular endothelial growth factor receptor-3 (VEGFR-3), the mitogenic tyrosine kinase receptor that drives LEC proliferation and migration [80, 90, 91]. To activate VEGFR-3, VEGF-C must first undergo stepwise proteolytic processing by several proteases to generate the mature form of the protein [92, 93]. Another receptor activated by the mature form of VEGF-C is the vascular endothelial growth factor receptor-2 (VEGFR-2) [90], which is also involved in lymphangiogenesis, albeit presumably in a more limited manner [80]. The VEGF-C-mediated activation of the VEGFR-3 signaling pathway is not only essential for lymphangiogenesis [94], but also plays a role in lymphatic vessel remodeling and homeostasis [95]. In the context of cancer, overexpression of VEGF-C induces the formation and remodeling of lymphatic vessels within and around primary tumors [96]. In addition, the aberrant activation of VEGFR-3 signaling by tumor-derived VEGF-C promotes metastatic spread of tumor cells via the lymphatics [96, 97]. Moreover, VEGF-C secreted by primary tumors stimulates lymphangiogenesis in the draining LNs even before metastasis occurs [98], inducing a permissive “lymphovascular niche” to ensure successful colonization and long-term survival of metastatic cells at the secondary site [99].
A growing number of lncRNAs have been shown to function as regulators of VEGF-C expression. For example, the lncRNA VEGFC-LNC was found to upregulate VEGF-C expression in human umbilical vein endothelial cells (HUVECs) [100], suggesting a role for this lncRNA in the activation of VEGF-C signaling under non-diseased physiological conditions. However, most lncRNAs implicated in VEGF-C regulation have been identified in cancers, where they either promote or inhibit tumor lymphangiogenesis and lymphatic metastasis. Some of these lncRNAs and their mechanisms of action are discussed below.
BLACAT2
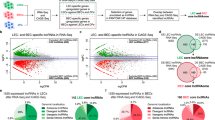
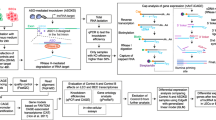
The bladder cancer-associated transcript 2 (BLACAT2), also known as LINC00958, was initially identified by Seitz et al. [101] as a candidate oncogene in bladder cancer and is regarded as one of the first examples of an lncRNA promoting VEGF-C-induced tumor lymphangiogenesis and lymphatic metastasis. He et al. [102] found that BLACAT2 is markedly overexpressed in metastatic bladder cancer, and its overexpression is positively correlated with LN metastasis and poor prognosis. They also showed that overexpression of BLACAT2 promotes lymphangiogenesis and lymphatic metastasis in animal models. Furthermore, they suggested that BLACAT2 exerts its oncogenic effect through an epigenetic mechanism that upregulates the expression of VEGF-C (Fig. 1a). In this mechanism, BLACAT2 forms an RNA–DNA triplex with the VEGF-C promoter and recruits WD repeat-containing protein 5 (WDR5), a core component of the histone H3 lysine 4 (H3K4) methyltransferase complexes [103], to epigenetically promote VEGF-C transcription through WDR5-mediated H3K4 methylation. Thus BLACAT2 promotes tumor lymphangiogenesis and lymphatic metastasis by epigenetically upregulating VEGF-C expression and pathologically activating the VEGFR-3 signaling pathway [102].
Representative lncRNA-mediated transcriptional and epigenetic mechanisms regulating the expression of the lymphangiogenic growth factor VEGF-C. a In the nucleus of bladder cancer cells, lncRNA BLACAT2 interacts with the core subunit WDR5 of the histone H3 lysine 4 (H3K4) methyltransferase complex and guides it to the VEGF-C promoter by forming a RNA–DNA triplex with the promoter sequence. The promoter H3K4 trimethylation (H3K4me3) by the catalytic subunit MLL (mixed lineage leukemia) of the methyltransferase complex drives epigenetic activation of VEGF-C transcription, which leads to the activation of the VEGF-C signaling pathway, thereby inducing tumor lymphangiogenesis and lymphatic metastasis. Based on He et al. [102]. b In triple-negative breast cancer cells, the nuclear pool of lncRNA HUMT activates FOXK1 transcription by recruiting the Y-box transcription factor YBX1 to the FOXK1 promoter. YBX1 is a known inhibitor of the histone methyltransferase activity of the polycomb repressive complex 2 (PRC2). Therefore, the YBX1-mediated inhibition of H3K27 trimethylation (H3K27me3) at the FOXK1 promoter may further contribute to the activation of FOXK1 transcription. FOXK1 subsequently activates the expression of the hypoxia-inducible transcription factor HIF-1a, which in turn activates the expression of VEGF-C. Based on Zheng et al. [134]
LNMAT1
Lymph node metastasis-associated transcript 1 (LNMAT1), alternatively named DUXAP9 or LINC01296, was identified as a candidate oncogene in bladder cancer in the same study by Seitz et al. [101] that discovered the oncogenic properties of BLACAT2. Chen et al. [104] showed that LNMAT1 is a nuclear-enriched lncRNA that is overexpressed in LN-positive bladder cancer and is associated with lymphatic metastasis and poor prognosis. Furthermore they demonstrated that, consistent with its oncogenic role in human bladder cancer, LNMAT1 overexpression induces lymphangiogenesis and lymphatic metastasis in animal models. According to Chen et al., the function of LNMAT1 in metastatic bladder cancer revolves around epigenetic activation of the small chemokine CCL2, which manifests itself in the deposition of H3K4 trimethyl activating marks at the CCL2 promoter region. Mechanistically, LNMAT1 forms a DNA-RNA triplex with the CCL2 promoter and interacts with the heterogeneous nuclear ribonucleoprotein L (HNRNPL) known for its role in transcriptional regulation [23]. By this means, LNMAT1 recruits HNRNPL to the CCL2 promoter and activates its transcription in the nucleus of bladder cancer cells. CCL2 activation and subsequent secretion by bladder cancer cells in turn upregulates VEGF-C expression in tumor-associated macrophages, promoting lymphangiogenesis and lymphatic metastasis [104]. It should be noted, however, that despite the established role of CCL2 in developmental and tumor-associated lymphangiogenesis [105], the exact molecular mechanism linking CCL2 activation with the enhanced VEGF-C expression remains to be elucidated.
BCYRN1
Brain cytoplasmic RNA 1 (BCYRN1), also known as LINC00004, is another example of an oncogenic lncRNA that promotes lymphatic metastasis through the activation of VEGF-C expression. Zheng et al. [106] found that BCYRN1 was significantly enriched in urinary exosomes from patients with bladder cancer compared to healthy controls. Furthermore, elevated levels of exosomal BCYRN1 were associated with increased lymphatic metastasis, decreased survival and poor prognosis in these patients. The authors proposed a dual mechanism of action for BCYRN1 in bladder cancer. On one hand, BCYRN1 activates the Wnt/β‐catenin signaling pathway to promote the secretion of VEGF‐C. To achieve this, BCYRN1 forms a DNA‐RNA triplex with the promoter of the WNT5A gene and, in association with the RNA‐binding protein HNRNPA1, induces its expression via H3K4 trimethylation [106]. Since WNT5A is a Wnt ligand [107], its overexpression activates the Wnt/β‐catenin signaling pathway, increasing the expression and secretion of VEGF‐C, which is a known Wnt target gene [108, 109]. On the other hand, Zheng et al. showed that BCYRN1 plays an important role in exosome‐mediated communication between bladder cancer cells and LECs. They found that BCYRN1 is transported via exosomes from bladder cancer cells to LECs, where it stabilizes VEGFR-3 mRNA by interacting with its 3′‐UTR. This results in the increased expression of VEGFR-3 on the surface of LECs, making the cells more sensitive to VEGF-C. Thus, BCYRN1 simultaneously increases both VEGFR-3 expression in LECs and VEGF-C secretion from bladder cancer cells, creating a feedforward loop that promotes tumor lymphangiogenesis and LN metastasis in bladder cancer through amplified VEGF‐C/VEGFR-3 signaling [106].
VESTAR
VEGF-C mRNA stability-associated lncRNA (VESTAR), otherwise known as LINC00638, was identified by Wang et al. [110] in a chromosomal region frequently amplified in esophageal squamous cell carcinoma. They found that VESTAR is overexpressed in esophageal and several other digestive system cancers, including cancers of the liver, stomach, and colon. Furthermore, they showed that VESTAR overexpression in esophageal squamous cell carcinoma tissues is accompanied by a change in its subcellular localization from the nucleus to the cytoplasm, suggesting a nuclear export mechanism. Indeed, VESTAR interacts with nuclear RNA export factor 1 (NXF1) and its most potent export adaptor SRSF3 [111] to facilitate its export from the nucleus. Notably, nuclear export of VESTAR correlates with regional LN metastasis, and this correlation was attributed to the VESTAR-mediated pathological activation of VEGF-C signaling [110]. The activation is due to an increase in VEGF-C mRNA stability caused by its direct interaction with VESTAR in the cytoplasm of esophageal cancer cells. In addition, VESTAR interacts with the RNA-binding protein HuR, enhancing its previously reported stabilizing effect on VEGF-C mRNA [112]. Consequently, overexpressed cytoplasmic VESTAR functions in association with HuR as a VEGF-C mRNA stabilizing factor, thereby promoting tumor-associated lymphangiogenesis and LN metastasis via aberrant activation of the VEGF-C signaling pathway [110].
ASLNC07322
SMAD family member 4 (SMAD4) is a critical component of the TGF-β pathway that acts as a tumor suppressor in several cancers, including pancreatic, bile duct and colon cancer [113]. Li et al. [114] demonstrated that SMAD4 represses VEGF-C expression in colon cancer by activating the transcription of the gene encoding miR-128-3p, a microRNA targeting the 3′ UTR region of VEGF-C mRNA. Furthermore, they found that the repression of VEGF-C by miR-128-3p could be relieved by the oncogenic lncRNA ASLNC07322, which acts as the miR-128-3p sponge. Thus, ASLNC07322 overexpression abrogates the tumor-suppressive effect of SMAD4, leading to the uncontrolled expression of VEGF-C, which in turn promotes tumor lymphangiogenesis and lymphatic metastasis [114].
AFAP1-AS1
A similar mechanism involving the activation of VEGF-C expression via microRNA sponging is employed by the lncRNA AFAP1-AS1, which stands for actin filament-associated protein 1 antisense RNA 1. AFAP1-AS1 is an oncogenic lncRNA [115] that, according to a recent study by Xia et al. [116], acts as a sponge for miR-27b-3p in cervical cancer cells, sequestering it away from its target VEGF-C. The study suggested that the derepression of VEGF-C by AFAP1-AS1 could be responsible for promoting lymphatic metastasis and enhancing cervical cancer stemness.
DANCR
Differentiation antagonistic non-protein coding RNA (DANCR), also called ANCR or AGU2, plays an important role in the progression, invasion and metastasis of several cancers, including cervical, pancreatic and bladder cancers [117,118,119]. In the case of bladder cancer, this lncRNA has been shown to promote lymphatic metastasis through sponging of miR-335 and derepression of its target gene, VEGF-C [120]. In this way, DANCR overexpression in bladder cancer leads to a pathological activation of pro-lymphangiogenic VEGF-C signaling, thereby promoting tumor lymphangiogenesis and LN metastasis.
MFSD4A-AS1
A recent study by Liu et al. [121] provides yet another example of how lncRNAs can regulate VEGF-C expression through miRNA sponging. The study found that the lymphatic node metastatsis-related lncRNA MFSD4A-AS1 is upregulated in papillary thyroid cancer (PTC) tissues with LN metastasis. Furthermore, the study showed that MFSD4A-AS1 promotes lymphangiogenesis and lymphatic metastasis in PTC by acting as a sponge for miR-30c-2-3p and miR-145-3p to induce VEGF-C expression via the ceRNA mechanism [121].
HNF1A-AS1
Hepatocyte nuclear factor 1 homeobox A antisense RNA 1 (HNF1A-AS1), also known as HAS1, is a tumor-associated lncRNA with an established role in the development and progression of many cancers, including those of the head and neck, breast, lung, bone, liver, colon, esophagus, bladder, and cervix [122]. Liu et al. [123] reported that HNF1A-AS1 overexpression correlates with LN metastasis in gastric cancer patients and promotes metastasis of gastric cancer in a xenograft mouse model. The authors attributed this effect to the ability of HNF1A-AS1 to induce lymphangiogenesis through the activation of the PI3K/AKT signaling pathway. PI3K/AKT signaling is one of the most frequently dysregulated pathways in cancer [124], and its aberrant activation promotes tumor lymphagiogenesis and lymphatic metastasis via increased expression and secretion of VEGF-C [125]. According to Liu et al., HNF1A-AS1 acts as a ceRNA for miR-30b-3p to upregulate the expression of the PIK3CD gene, which encodes the delta isoform of the catalytic subunit of phosphoinositide 3-kinase (PI3K-delta), a key component of the PI3K/AKT signaling pathway and a known oncogene [126]. Thus, the HNF1A-AS1-mediated upregulation of PI3K-delta activates PI3K/AKT signaling in gastric cancer cells, inducing VEGF-C secretion and ultimately promoting tumor lymphangiogenesis and lymphatic metastasis [123].
circNFIB1
The circular lncRNA NFIB1 (hsa_circ_0086375) differs from the other lncRNAs discussed here in that it has anti-lymphangiogenic properties and is downregulated in pancreatic ductal adenocarcinoma (PDAC) patients with LN metastasis [127]. Circular RNAs (circRNAs) are characterized by a covalently closed circular structure produced through a non-canonical form of splicing called “back-splicing” [128]. Unlike linear lncRNAs, circular RNAs lack a 5′ cap and a 3′ polyadenylated tail, making them more resistant to RNase-mediated degradation [129]. Although the majority of circRNAs are thought to be the products of splicing errors [130], some do have functional roles in development and disease. One such is circNFIB1, which was shown by Kong et al. [127] to inhibit lymphangiogenesis in vitro and suppress LN metastasis of PDAC in a mouse model. Circular lncRNAs typically act as post-transcriptional regulators through miRNA sponging [131,132,133], and circNFIB1 is no exception. According to Kong et al., circNFIB1 functions as a sponge for the oncogenic miR-486-5p in the cytoplasm of PDAC cells to derepress its target, the regulatory subunit of phosphoinositide 3-kinase (PIK3R1). The derepression of PIK3R1, in turn, leads to the downregulation of VEGF-C expression via inhibition of the PI3K/Akt pathway. Overall, the findings of Kong et al. indicate that circNFIB1 is an anti-lymphangiogenic lncRNA that suppresses lymphangiogenesis and LN metastasis in PDAC via the miR-486-5p/PI3KR1/VEGF-C axis [127].
HUMT
LINC00857, better known as HUMT, which stands for lncRNA highly upregulated in metastatic triple-negative breast cancer (TNBC), is yet another lncRNA that regulates, albeit indirectly, the expression of VEGF-C in cancer. As its name implies, HUMT is highly expressed in TNBC [134], which is the most malignant subtype of breast cancer with the highest lymphatic metastatic potential [135]. Using a combination of bioinformatic and biochemical approaches, Zheng et al. [134] demonstrated that in the nucleus of TNBC cells, HUMT recruits the Y-box transcription factor YBX1, a PRC2 interactor and inhibitor of H3K27me3 [136], to the promoter region of the forkhead box K1 transcription factor (FOXK1) and activates its transcription [134] (Fig. 1b). FOXK1 is a known inducer of the hypoxia-inducible transcription factor 1 alpha (HIF-1α) [137], which in turn activates the expression of VEGF-C [138,139,140]. Thus HUMT promotes tumor-induced lymphangiogenesis and lymphatic metastasis in TNBC by pathologically activating the VEGF-C signaling pathway via the FOXK1/HIF-1α axis [134].
LncRNA-mediated regulation of pro-lymphangiogenic VEGF-A signaling
VEGF-A is a hypoxia-driven secreted growth factor that signals through vascular endothelial growth factor receptors 1 and 2 (VEGFR-1 and VEGFR-2) to induce proliferation, migration, sprouting, permeability and survival of endothelial cells [141]. VEGF-A belongs to the same family of proteins as VEGF-C and plays a critical role in developmental and pathological angiogenesis. In addition to its role in angiogenesis, VEGF-A signaling through VEGFR-2 is also implicated in lymphangiogenesis [80], along with VEGF-C signaling through VEGFR-3/VEGFR-2. Therefore, dysregulation of VEGF-A expression by lncRNAs may contribute to the pathogenesis of lymphatic-associated diseases including metastatic cancer. Indeed, Shi et al. [142] showed that the aberrant induction of VEGF-A by the lncRNA HANR (also known as RPL13AP20) promotes tumor lymphangiogenesis in hepatocellular carcinoma. They also found that the underlying molecular mechanism involves the HANR-mediated sponging of miR‐296 in hepatocellular carcinoma cells, reducing the release of miR‐296 from the cells in the form of exosomes. Consequently, the LECs are able to internalize less exosomal miR‐296, causing derepression of the miR‐296 target EAG1, a potassium channel protein known to induce HIF-1α and promote the expression of VEGF-A [143]. Thus the overexpression of HANR in hepatic cancer cells leads to the activation of the VEGF-A/VEGFR-2 signaling pathway in LECs, thereby promoting tumor-associated lymphangiogenesis [142]. Another example of how an lncRNA can induce tumor lymphangiogenesis by upregulating VEGF-A expression comes from the above-mentioned study on the role of MFSD4A-AS1 in papillary thyroid cancer. In addition to demonstrating that MFSD4A-AS1 controls VEGF-C expression through miRNA sponging, the study identified a similar mechanism regulating the expression of VEGF-A. According to this mechanism, MFSD4A-AS1 acts as a sponge for miR-139-5p to upregulate the expression of its target gene, VEGF-A. This leads to pathological activation of the VEGF-A/VEGFR-2 signaling pathway, which in turn stimulates lymphangiogenesis and lymphatic metastasis in papillary thyroid cancer [121].
NEAT1: an IRES-dependent translational regulator of mRNAs encoding lymphangiogenic growth factors
As one of the best-studied oncogenic lncRNAs, nuclear paraspeckle assembly transcript 1 (NEAT1) is known to promote metastasis of various cancers, including those of the breast, lung, thyroid gland, colon, ovary, prostate, and liver [144]. Despite the rapidly accumulating knowledge about the diverse mechanisms through which NEAT1 exerts its oncogenic activity [144, 145], our understanding of how NEAT1 dysregulates the key lymphangiogenic pathways to promote lymphatic metastasis is still limited. In a recent study, Godet et al. [146] identified the essential nuclear paraspeckle component NEAT1 as a novel translational regulator which enhances the translation of fibroblast growth factor 1 (FGF-1) as well as VEGF-C and VEGF-A via internal ribosome entry sites (IRESs) in the corresponding mRNAs. Since FGF-1, VEGF-C and VEGF-A are all known inducers of lymphangiogenesis [89, 141, 147], the results of Godet et al. shed light on a possible mechanism by which NEAT1 may activate FGF, VEGF-C or VEGF-A signaling in cancer, thereby promoting tumor lymphangiogenesis and lymphatic metastasis. Furthermore, the notion that NEAT1 is a pro-lymphangiogenic lncRNA is also supported by an independent study demonstrating that NEAT1 upregulates VEGF-C expression in bladder cancer by sponging its negative regulator miR-101 [148].
Regulation of lymphangiogenic growth factor IGF-1 by LncCCLM
Cancer lymphatic metastasis-associated lncRNA (LncCCLM), also known as RP11-7K24.3, is an example of an lncRNA that acts as a suppressor of lymphatic metastasis. Chen et al. [149] found that LncCCLM is downregulated in cervical cancer tissues, and its low expression is associated with an increased risk of distant lymphatic metastasis. They also showed that LncCCLM decreases cervical cancer cell migration and invasion in vitro and inhibits lymphatic metastasis of cervical cancer in a mouse model. The authors proposed a mechanism for the action of LncCCLM in cervical cancer, according to which cytoplasmic LncCCLM interacts with Staufen double-stranded RNA binding protein 1 (STAU1) to promote the decay of the insulin-like growth factor 1 (IGF-1) mRNA. This leads to a decrease in the amount of IGF-1 protein, which is a known inducer of tumor lymphangiogenesis and lymphatic metastasis [150]. Based on these findings, Chen et al. concluded that LncCCLM functions as a suppressor of lymphatic metastasis in cervical cancer by inhibiting the pro-lymphangiogenic IGF-1/IGF-1R signaling pathway via the STAU1-mediated degradation of IGF-1 mRNA [149].
LncRNA-mediated control of PROX1, the master regulator of lymphatic differentiation and development
The prospero homeodomain transcription factor (PROX1) is the master regulator of LEC identity, initiating and maintaining the specific transcriptional program that governs LEC differentiation from a subpopulation of venous endothelial cells (VECs) [49, 82]. PROX1 serves as a specific marker of developing and adult lymphatic vasculature [151] and deregulation of its expression is linked to several lymphatic-associated diseases comprising the metabolic syndrome [62] such as hyperlipidemia, obesity and diabetes [152, 153]. In addition, PROX1 plays an important but ambivalent role in cancer [154] as either oncogene [155,156,157,158] or tumor suppressor [159,160,161,162], depending on the cancer type and context. Therefore, a better understanding of the molecular mechanisms regulating PROX1 expression not only expands our knowledge of normal lymphatic development and function, but also provides novel insights into the pathophysiology of lymphatic-associated diseases and cancer.
ANRIL and GAS5
The antisense noncoding RNA in the INK4 locus (ANRIL), also called CDKN2B-AS1, was first identified in patients with familial melanoma [163]. Cunnington et al. [164] subsequently showed that ANRIL is associated with coronary artery disease and diabetes, where its expression is downregulated. On the other hand, Sun et al. [165] found that ANRIL promotes lymphangiogenesis and lymphatic metastasis in colorectal cancer by upregulating the expression of VEGF-C and VEGFR-3. These findings prompted He et al. [167] to investigate whether ANRIL might promote lymphangiogenesis to accelerate the process of wound healing, which is impaired in diabetes [166]. Indeed, the authors showed that ANRIL upregulates the expression of PROX1 in LECs on a post-transcriptional level, thereby promoting lymphangiogenesis and accelerating wound healing [167]. Mechanistically, this is achieved by ANRIL-mediated sponging of miR‐181a (Fig. 2a), which was characterized previously as a negative regulator of PROX1 expression [168].
Examples of cytoplasmic mechanisms by which lncRNAs regulate key molecular players involved in lymphangiogenesis. a MicroRNA sponging. (i) miR-181a binds to the 3′-UTR of the PROX1 mRNA inducing translational repression and mRNA decay. (ii) The lncRNA ANRIL functions as a decoy to sponge miR-181a away from PROX1 mRNA, derepressing its translation. The increased translation of PROX1 leads to the activation of genes that promote lymphangiogenesis, thereby accelerating diabetic wound healing. Based on He et al. [167]. b Regulation of mRNA stability. The natural antisense lncRNA FOXC2-AS1 forms an RNA-RNA duplex with the FOXC2 mRNA, stabilizing it and protecting it from RNase-mediated cleavage. The resulting aberrant increase in FOXC2 translation promotes EMT and tumor metastasis. In a non-cancer context, the same mechanism might regulate lymphatic valve formation and collecting lymphatic vessel specialization. Based on Zhang et al. [208] and Missaglia et al. [216]. eIF4F eukaryotic initiation factor 4F, m7G methyl-7-guanosine (cap), miRISC microRNA-induced silencing complex, ORF open reading frame, PABP poly(A)-binding protein
The lncRNA growth arrest-specific transcript 5 (GAS5), also known as SNHG2, was originally identified as a non-protein-coding tumor suppressor gene that is highly expressed during growth arrest [169]. GAS5 is downregulated in many cancers including breast, prostate, lung and colorectal cancer, and its reduced expression correlates with poor prognosis [170]. In a non-cancer setting, GAS5 participates in diabetic wound healing promoted by topical mevastatin [171]. As just described in the case of ANRIL, GAS5 promotes lymphangiogenesis during wound healing via a PROX1-dependent mechanism. To establish this mechanism, He et al. [172] first carried out bioinformatics analysis and found that both GAS5 and PROX1 contain binding sites for the same microRNA, miR-217. Consequently, they showed that GAS5 acts in a similar fashion to ANRIL, sponging miR-217 and thereby derepressing the expression of PROX1.
Given the similarities in the pro-lymphangiogenic mechanisms of ANRIL and GAS5, it is not surprising that both lncRNAs behave quite similarly [167, 172]. Both ANRIL and GAS5 are downregulated in the skin of diabetic mice or human patients. Furthermore, treating LECs with high glucose downregulates the expression of both lncRNAs, leading to the inhibition of lymphangiogenesis. On the other hand, overexpression of ANRIL or GAS5 accelerates wound healing, underscoring the prominent role of lymphangiogenesis in this process. Interestingly, the ANRIL- or GAS5-mediated sponging of their respective miRNAs upregulates the expression of not only PROX1, but also its direct target VEGFR-3 [173]. Taken together, the above findings indicate that ANRIL and GAS5 effectively improve wound healing by promoting lymphangiogenesis via the PROX1/miR‐181a and PROX1/miR-217 axes, thus highlighting the importance of these two lncRNAs in the regulation of the PROX1-mediated LEC-specific transcriptional program. Therefore, both ANRIL and GAS5 deserve further evaluation as potential therapeutic targets for the treatment of delayed wound healing in diabetic patients [167, 172]. Finally, it should be noted that the opposing roles of ANRIL and GAS5 as tumor promoter versus tumor suppressor likely reflect the ambivalent role of PROX1 in cancer.
MIAT
Another example of PROX1 regulation by miRNA sponging is provided by the lncRNA myocardial infarction‑associated transcript (MIAT), also known as RNCR2 or Gomafu. MIAT plays an important role in development and various diseases [174] and is involved in the differentiation of mesenchymal stem cells (MSCs) into endothelial cells (ECs) [175]. In a recent study, MIAT was shown to promote the differentiation of adipose‑derived mesenchymal stem cells (ADMSCs) into LECs by regulating the expression of PROX1 [176]. To achieve this, MIAT acts as a molecular sponge of miR-495, for which a binding site has been identified in the 3′‑untranslated region of the PROX1 mRNA. Thus, MIAT upregulates PROX1 expression through competitive binding to miR-495, thereby promoting the transcriptional reprogramming of ADMSCs into LECs. Since the induced differentiation of ADMSCs into LECs is emerging as a novel avenue for the treatment of lymphedema [177,178,179], MIAT has been proposed as a potential therapeutic target in this disease [176].
LNMAT2
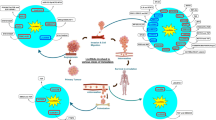
Yet another mechanism by which an lncRNA is able to regulate PROX1 expression has been identified in the context of bladder cancer. The lncRNA lymph node metastasis-associated transcript 2 (LNMAT2), otherwise known as LINC00858, was found by Chen et al. [180] to be overexpressed in bladder cancer cells, and its overexpression positively correlated with LN metastasis. They also showed that, in order to fulfill its lymphatic metastatic potential, the overexpressed LNMAT2 interacts with the RNA‐binding protein HNRNPA2B1 (heterogeneous nuclear ribonucleoprotein A2/B1) and the resulting complex is released by cancer cells via exosomes (Fig. 3). The exosomes are subsequently internalized by LECs, promoting tumor-associated lymphangiogenesis and lymphatic metastasis through an epigenetic mechanism upregulating the transcription of PROX1. The mechanism involves the formation of a DNA-RNA triplex between LNMAT2 and the PROX1 promoter, thereby recruiting the LNMAT2-tethered HNRNPA2B1 to the promoter region and increasing its H3K4 trimethylation (H3K4me3) [180]. The resulting aberrant epigenetic activation of PROX1 induces transcriptional reprogramming, which leads to the uncontrolled expression of lymphatic genes. Interestingly, the LNMAT2-mediated mechanism of PROX1 activation is VEGF-C-independent, explaining why approximately 20% of bladder cancers with LN metastasis have low VEGF-C expression [181, 182].
Representative mechanism of intercellular communication between cancer cells and lymphatic endothelial cells (LECs) mediated by exosomal lncRNA. The lncRNA LNMAT2, which is overexpressed in bladder cancer cells, contains the exo-motif GGAG recognized by the RNA binding protein HNRNPA2B1. The interaction with HNRNPA2B1 facilitates LNMAT2 sorting into exosomes, which are formed during endosome maturation through inward membrane budding of multivesicular bodies (MVBs). The exosomes are secreted out of cancer cells and subsequently internalized by LECs. Upon entering LECs, the exosomes dissociate and their cargo translocates into the nucleus, where LNMAT2 forms a DNA-RNA triplex with the PROX1 promoter. The LNMAT2-tethered HNRNPA2B1 activates PROX1 transcription by increasing the levels of H3K4 trimethylation (H3K4me3) in the promoter region. The epigenetically induced overexpression of PROX1 results in aberrant transcriptional reprogramming, which promotes tumor lymphangiogenesis and LN metastasis. Based on Chen et al. [180]
PROX1-AS1
PROX1-AS1, a natural antisense transcript overlapping the PROX1 gene, is an oncogenic lncRNA that is upregulated in renal and lung cancer and positively correlates with tumor metastasis [183, 184]. However, despite these findings, the role of PROX1-AS1 in the regulation of PROX1 during lymphatic vascular development and disease is currently unknown and requires further investigation.
Regulation of lymphangiogenic transcription factor SOX18 by ELNAT1
In addition to PROX1, several other transcription factors play a critical role in the specification of endothelial cells into lymphatic lineage. One of them is SRY-Box transcription factor 18 (SOX18), which activates the expression of PROX1 [185] in cooperation with chick ovalbumin upstream promoter transcription factor 2 (COUP-TFII) [186]. Recently, an lncRNA named extracellular lymph node-associated transcript 1 (ELNAT1), also known as SNHG16, has been shown by Chen et al. [187] to epigenetically upregulate the expression of SOX18, thereby inducing tumor lymphangiogenesis and lymphatic metastasis in bladder cancer. The molecular mechanism underlying this pathological process resembles that of LNMAT2. In the nucleus of bladder cancer cells, ELNAT1 forms a DNA-RNA triplex with the promoter of small ubiquitin-like modifier (SUMO)-conjugating enzyme 9 (UBC9) and, in association with the RNA‐binding protein HNRNPA1, activates the transcription of the UBC9 gene. The overexpressed UBC9 promotes the SUMOylation of HNRNPA1 to facilitate sorting and packaging of ELNAT1 into extracellular vesicles. The packaged vesicles are secreted by cancer cells and then internalized by LECs, where ELNAT1 is translocated into the nucleus, once again forming a DNA-RNA triplex in association with HNRNPA1, but this time with the SOX18 promoter. The epigenetic activation of the SOX18 gene in LECs is evident from the increased levels of HNRNPA1-induced H3K4me3 at the promoter region. Based on these findings, Chen et al. concluded that the overexpression and subsequent secretion of ELNAT1 from bladder cancer cells promotes tumor lymphangiogenesis and lymphatic metastasis through the upregulation of SOX18, which in turn leads to the aberrant activation of the PROX1-driven lymphatic transcriptional program [187].
MAPK8IP1P2 as a regulator of the Hippo pathway
The Hippo-YAP/TAZ signaling pathway plays an essential role in organ growth and tissue homeostasis [188, 189] and its dysregulation is commonly associated with cancer development and progression [190]. Increasing evidence suggests that the Hippo pathway is also critical during lymphatic vascular development. For instance, Hippo signaling negatively regulates PROX1 expression during LEC specification and sprouting [191]. In line with this, it has been proposed that the Hippo signaling effectors YAP/TAZ play a role in the PROX1/VEGFR-3 feedback loop during LEC specification and migration and lymphatic valve maturation [192].
Liu et al. [193] reported that the lncRNA MAPK8IP1P2 was downregulated in thyroid carcinoma with lymphatic metastasis. They also found that MAPK8IP1P2 acts as a sponge for miR-146b-3p to relieve the repression of three tumor suppressor genes involved in Hippo signaling. The first gene, NF2, is often mutated or inactivated in cancer [194, 195] and serves as an activator of the Hippo pathway [196, 197]. The two other genes, RASSF1 and RASSF5, belong to the C-terminal Ras-association domain family (RASSF), whose members suppress tumorigenesis by binding to and regulating the upstream Hippo kinase MST1/2 [198,199,200]. Liu et al. showed that the MAPK8IP1P2-mediated upregulation of NF2, RASSF1 and RASSF5 leads to the activation of the antitumorigenic Hippo signaling pathway in thyroid cancer cells. Thus MAPK8IP1P2 acts as an oncosuppressor lncRNA with anti-lymphangiogenic properties, inhibiting lymphatic metastasis in thyroid cancer via the activation of Hippo signaling [193].
LncRNA-mediated control of the master regulator of lymphatic valve morphogenesis FOXC2
FOXC2 is a lymph flow-induced forkhead transcription factor which acts as a master regulator of lymphatic valve morphogenesis [201, 202]. In addition to its central role in the formation of lymphatic valves, FOXC2 also controls specialization of collecting lymphatic vessels during embryonic and postnatal development [203]. Inactivating mutations in FOXC2 are the underlying cause of lymphedema-distichiasis syndrome (LD; OMIM 153400) characterized by late-onset hereditary lymphedema and the presence of a double row of eyelashes (distichiasis) [204, 205]. Furthermore, FOXC2 acts as a potent oncogene, promoting cancer cell proliferation and epithelial–mesenchymal transition (EMT), which is considered a key step in tumor metastasis [206].
A natural antisense transcript (NAT) overlapping the FOXC2 gene, designated FOXC2-AS1, was first identified by microarray analysis in doxorubicin-resistant osteosarcoma cell lines [207]. Zhang et al. [208] found that in these cells FOXC2-AS1 was predominantly localized in the cytoplasm in close proximity to FOXC2 mRNA. This finding, together with the fact that FOXC2-AS1 is fully complementary to 145 nucleotides in the first exon of FOXC2, prompted the authors to suggest that the two RNA molecules could interact, forming an RNA-RNA duplex. Indeed, Zhang et al. observed the formation of a double-stranded complex of FOXC2-AS1 and FOXC2 mRNA in the overlapping complementary sequence region. The complex formation has a stabilizing effect on FOXC2 mRNA, protecting it from RNase cleavage, and thus leading to an increase in the FOXC2 protein synthesis (Fig. 2b). Given the well-established role of FOXC2 dysregulation in cancer, the results of Zhang et al. implicated the FOXC2 inducer FOXC2-AS1 as a novel oncogene, whose aberrant overexpression contributes to cancer development, progression and metastasis, and correlates positively with poor prognosis [208]. The oncogenic potential of FOXC2-AS1 has been since confirmed in several other cancers including breast, lung, skin, colorectal, gastric and prostate tumors [209,210,211,212,213,214]. In colorectal and gastric cancers, FOXC2-AS1 utilizes a similar mechanism involving FOXC2 mRNA stabilization [212, 213]. However, in prostate cancer cells, cytoplasmic FOXC2-AS1 acts as a molecular sponge to sequester miR-1253 from its target EZH2 [214], which encodes an oncogenic histone methyltransferase and catalytic subunit of PRC2 [215]. In addition to the above cytoplasmic mechanisms involving mRNA stabilization and miRNA sponging, another oncogenic mechanism of FOXC2-AS1 action has been identified in the nucleus of non-small cell lung cancer (NSCLC) and melanoma cells. In these cells, the nuclear pool of FOXC2-AS1 mediates epigenetic repression of the tumor suppressor gene p15 via recruitment of PRC2 to its promoter [210, 211].
Recently experimental evidence has emerged suggesting that at least some of the above FOXC2-AS1-mediated mechanisms are operational in non-cancer cells. Missaglia et al. [216] detected endogenous expression of FOXC2-AS1 in blood cells of healthy individuals and LD patients. Furthermore, they found that, consistent with the results obtained in cancer cell lines, endogenous FOXC2-AS1 positively regulates the expression of wild-type FOXC2 and its frameshift LD mutant in normal, non-cancer cells. So far, such regulation has only been demonstrated in peripheral blood cells; the next step would be to confirm the expression of FOXC2-AS1 in LECs and study its regulatory role in lymphatic vascular development and homeostasis.
In summary, recent research has implicated lncRNAs in the regulation of several key molecular players and pathways involved in lymphangiogenesis (Table 1). This is a fast-moving area of research and some results still require validation in vivo before they can be considered fully reliable. Future studies in this emerging field should move beyond the predominantly in vitro work carried out so far and more thoroughly examine the functions of lymphatic-associated lncRNAs in animal models.
LncRNAs as diagnostic tools and therapeutic targets in lymphatic vascular diseases and cancer
LncRNAs are emerging as promising targets for therapeutic intervention and as potential diagnostic biomarkers. The fact that some lncRNAs can be lineage-, tissue- or disease-specific [4] underscores their diagnostic potential. For example, the lncRNA prostate cancer antigen 3 (PCA3) is specifically overexpressed in prostate cancer compared to normal prostate tissue and has been approved as a diagnostic biomarker for the early detection of prostate cancer [217]. As the pioneering lncRNA biomarker, PCA3 demonstrates a diagnostic accuracy superior to the most commonly used prostate-specific protein biomarker, prostate-specific antigen (PSA) [218]. In light of the above, the identification of LETR1, the first lymphatic lineage-specific lncRNA, paves the way for the development of novel diagnostic approaches to diseases involving the lymphatics such as lymphedema. Furthermore, lncRNAs associated with lymphatic pathologies, such as those promoting tumor lymphangiogenesis and lymphatic metastasis, may also have diagnostic and prognostic value.
LncRNAs represent promising therapeutic targets for the following reasons: (1) lncRNAs are generally expressed at lower levels than mRNAs [219] and hence require lower doses of potentially toxic drugs to achieve an equal therapeutic effect; (2) lncRNA expression can be lineage-, tissue-, disease-, or even cell population-specific [220], allowing for more selective therapy with fewer side-effects; (3) the fact that lncRNAs do not need to be translated into protein offers the advantage of greater drug design flexibility and faster response to therapy. As discussed above, the majority of lncRNAs involved in lymphatic-associated diseases constitute oncogenes promoting tumor lymphangiogenesis and lymphatic metastasis. These lncRNAs are therefore the most obvious targets for therapeutic intervention. Several therapeutic strategies to suppress oncogenic lncRNAs or alter their epigenetic effects are outlined in the paragraph below. Targeting lncRNAs could also be a promising therapeutic option for the treatment of other lymphatic vascular pathologies outside the cancer context. For example, downregulation of anti-lymphangiogenic lncRNAs or upregulation of pro-lymphangiogenic lncRNAs may improve delayed wound healing in diabetic patients and provide a therapeutic benefit in primary or secondary lymphedema, including postsurgical lymphedema after lymph node removal.
Since the advantages and limitations of various therapeutic strategies targeting lncRNAs have been extensively reviewed elsewhere [221,222,223,224], we will only briefly reintroduce them to the reader. These strategies include small-molecule inhibitors, RNA interference (RNAi), antisense oligonucleotides (ASOs), ribozymes, and genome editing tools. Small-molecule inhibitors bind to lncRNAs, changing their secondary structures or masking their protein-binding sequences. Alternatively, they may bind to RNA-binding proteins disrupting their interactions with lncRNAs. A variety of methods including small-molecule microarrays, label-based methods, mass spectrometry, dynamic combinatorial chemistry, NMR spectroscopy and virtual screening can be used to identify small molecule lncRNA binders [225]. At first glance, small-molecule lncRNA inhibitors appear to be a promising therapeutic option due to their high tissue penetration ability. However, if such molecules need to be delivered to the lymphatics, their small size becomes an issue because small molecules are readily reabsorbed into the bloodstream instead of accumulating in the lymphatic system [226]. A potential solution to this problem is the conjugation or physical encapsulation of lncRNA-targeting small molecules into various nanocarriers (see the section below on lymphatic delivery). The lncRNA-targeting strategy based on RNAi takes advantage of the natural process that largely occurs in the cytoplasm and employs small interfering RNAs (siRNAs) or short hairpin RNAs (shRNAs) as guides for sequence-specific gene silencing [227]. RNAi has been widely used to knock down predominantly cytoplasmic lncRNAs [228,229,230,231], making it a potential therapeutic approach. The main advantage of RNAi is its simplicity, as most cells already have functional RNAi machinery and do not require additional enzymatic components. On the other hand, dsRNA-induced immune responses, incomplete silencing, and widespread off-target effects limit the use of RNAi in the clinic [232,233,234]. A number of approaches are being explored to address these issues. For example, off-target effects have been minimized by increasing the length of siRNAs from the ‘conventional’ 21–23 nucleotides to 27 nucleotides and reducing siRNA concentrations to picomolar levels [235,236,237]. Furthermore, the ongoing development of novel algorithms and machine learning approaches for the analysis of siRNA-RNA interaction networks offers the potential to circumvent dsRNA-induced immune responses and select the most effective siRNAs for further clinical use [237,238,239]. ASOs are short, chemically synthesized, single-stranded antisense DNA oligonucleotides that bind to target lncRNA through base pairing rules, triggering RNase H-mediated lncRNA degradation [240]. Locked nucleic acid GapmeRs (LNA GapmeRs) are similar to native ASOs except that they are end-modified by LNA to increase nuclease resistance and binding affinity towards complementary lncRNA molecules [241]. Given their greater effectiveness in targeting nuclear transcripts compared to RNAi approaches [242], LNA GapmeRs have become the most widely used means of knocking down lncRNA in the nucleus. In addition, LNA GapmeRs show promise in relieving the repression of mRNAs by their natural antisense lncRNA transcripts (NATs). Commonly termed “antagoNATs”, these LNA GapmeRs inhibit the sense-antisense interaction between mRNAs and complementary NATs, triggering the NAT cleavage by RNase H and subsequent degradation by exonucleases [243]. Mixmers represent yet another type of chemically modified ASOs composed of alternating short stretches of LNA and DNA. In contrast to GapmeRs, mixmers do not trigger RNase H-dependent degradation of lncRNAs, acting instead as steric blockers of their targets [241]. ASOs have several advantages over siRNAs, including independence from the RNAi machinery, lower immunogenicity, and the ability to enter the nucleus more easily due to their small size [222]. Moreover, ASOs have greater specificity than siRNAs and cause fewer off-target effects [221]. Nonetheless, imperfect binding of ASOs to partially complementary regions in RNA, or to proteins, remains an issue, resulting in unintended off-target effects associated with hepato-, renal-, and neurotoxicity [244]. A possible solution to this problem is to train machine learning models to predict ASO toxicity using toxicity-associated sequence features as inputs [245]. Another lncRNA-targeting strategy involves ribozymes and deoxyribozymes, which are nucleic acid molecules with enzymatic activity that can be designed to target lncRNAs through base pairing, catalyzing their cleavage in trans [246, 247]. Trans-cleaving ribozymes, such as the hammerhead ribozyme (hhRz), have a significant advantage over exogenous ribonucleases for lncRNA cleavage because they are less likely to elicit a host immune response and are small enough to be easily incorporated into gene therapy vectors [248]. Furthermore, hhRzs have a low tolerance for even a single nucleotide mismatch with their targets [249], implying that they will have fewer off-target effects than siRNAs or ASOs. Indeed, when hhRz libraries were used for functional gene discovery, they caused less off-target effects than siRNA libraries [237]. The fact that hhRz has reached the stage of clinical trials for HIV-1 infection [250] demonstrates its potential for clinical applications, including lncRNA-targeted therapies. However, as with any technology, hhRzs have limitations. One of the concerns is that, despite their higher specificity, hhRzs have lower suppressive activity compared to siRNAs [237]. In the future, the discovery of more ribozyme motifs and a better understanding of the molecular and structural mechanisms underlying ribozyme action will likely lead to the engineering of new variants of trans-cleaving ribozymes with enhanced therapeutic properties [251]. Finally, genome-editing tools offer a range of opportunities not only for lncRNA inactivation but also for restoring the functions of pathologically downregulated or lost lncRNAs [222, 252]. These tools include, among others, zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), the clustered regularly interspaced short palindromic repeats (CRISPR/Cas9) system and its modifications such as base editing, prime editing, RNA editing with Cas7-11, CRISPR-based epigenetic editing and programmable addition via site-specific targeting elements (PASTE) [253,254,255,256]. The undisputed advantage of genome-editing tools for therapeutic targeting of lncRNAs is their high effectiveness. For example, a ZFN-based approach reduced the expression of the highly abundant lncRNA MALAT1 in human lung cancer cells by more than 1000-fold, which is 50 times greater than the 20-fold reduction achieved with ASOs [257]. However, like any technology, genome-editing tools such as CRISPR/Cas9 come with their own caveats. Potential off-target effects limit their clinical application [258,259,260], albeit to a lesser extent than with traditional technologies like RNAi. The development of computational prediction algorithms and machine learning models [261, 262], as well as the engineering of nucleases with improved specificity [263], are among the approaches taken to address this issue. Furthermore, successful genome editing requires the delivery of a large cargo, such as long CRISPR/Cas9 sequences, into target tissues and cells, which remains a major therapeutic challenge [264]. The development of novel carrier systems for the delivery of large genome-editing agents remains an area of active research [265], and some of the proposed approaches are discussed in the next section. Overall, a number of promising strategies are currently being explored for therapeutic targeting of lncRNAs. Although the field is still developing and no effective therapy has yet been approved for clinical use, lncRNA-based therapeutics hold great potential for the treatment or even prevention of a variety of lymphatic-associated diseases, including metastatic cancer.
Approaches to lymphatic delivery of lncRNA-based therapeutics
The choice of optimal delivery system is one of the most important and challenging problems in the development of therapies targeting lncRNAs in the lymphatic endothelium. Many current drugs that aim to reach the lymphatic system, such as anticancer chemotherapeutic agents, are administered intravenously. However, when administered in this way, these drugs not only show poor uptake into the lymphatics [266, 267] but also cause toxicity upon entering normal, unaffected organs and tissues. Moreover, they are quickly eliminated from the body via renal and hepatobiliary clearance pathways instead of accumulating in the lymphatic system [268]. The two primary reasons for this are the suboptimal route of administration and the small drug size of less than 5‒10 nm, which allows drug reabsorption back into the circulation through blood vessel walls [269, 270]. A more promising approach is to use the lymphatic vasculature as a delivery route and encapsulate drugs in nanocarriers with sizes ranging from 20 to 100 nm, which are thought to be optimal for lymphatic uptake and LN accumulation [268, 271]. Multiple strategies are currently being developed for the targeted delivery of therapeutic agents to the lymphatic vasculature. In these strategies, cargo molecules are packaged into a variety of liposomes, micelles, nanoparticles, and other nanocarrier systems [272] before being injected into subcutaneous or intradermal lymphatic vessels using conventional or microneedle techniques [226]. Though we discuss some nanocarrier delivery strategies in the following paragraphs, the literature on this subject is vast and rapidly growing, and we refer the reader to several comprehensive reviews for more information and references [226, 272,273,274]
As many times before in history, insights from nature may provide important clues for solving the problem of targeted lymphatic delivery. As discussed above, oncogenic lncRNAs can be loaded into lipid-based EVs (e.g., exosomes), secreted by cancer cells and then successfully internalized by other cells such as LECs (Fig. 3). This EV internalization mechanism can be modified to handle therapeutic molecules targeting lncRNAs. There are several reasons why endogenous or artificial EVs hold promise for the development of lncRNA-targeted therapies for various diseases involving the lymphatic vasculature. First, the payload in such vesicles is encapsulated in a lipid bilayer, reducing concerns over its stability. Second, lipid-based EVs are less immunogenic and more stable in vivo than the widely used viral vectors [223]. Third, artificial EVs coated with suitable hydrophilic polymers and/or assembled from several lipid components can achieve increased cellular uptake and circulatory half-life with reduced cytotoxicity [221]. Fourth, the low packaging capacity of endogenous exosomes can be markedly increased through integration with artificial liposomes or lipid nanoparticles. The resulting exosome-liposome hybrids can encapsulate molecules as large as CRISPR/Cas9 expression vectors [275]. However, most importantly, such exosome-liposome hybrids could retain specific ligands naturally targeting them to LECs. It is well known that LECs cannot be efficiently transfected with conventional cationic liposomes such as Lipofectamine 2000 or Oligofectamine, which lack the protein components of endogenous EVs. To overcome this problem, it will be necessary to identify the specific ligands on the surface of secreted EVs that are responsible for targeting them to LECs. One such ligand might be laminin γ2 because its knockdown in cancer cells results in the secretion of dysfunctional EVs which cannot be efficiently internalized by LECs and show a reduced ability to drain into LNs and promote lymphangiogenesis [276]. In addition, several surface determinants for endothelial targeting have been identified in the blood vascular endothelium [277]. It remains to be seen whether some of these determinants can be utilized for the delivery of therapeutic EVs into LECs. The successful identification of LEC-targeting ligands would allow engineering of artificial EVs that can deliver their therapeutic cargo via selective uptake by LECs. One way to achieve this is by expressing the targeting ligand in donor cells, which will then be passed through membrane pores or microfluidic devices [278] to form artificial exosomes with the targeting ligand on their surface. The desired payload, such as molecules selectively interfering with lncRNA expression, can then be loaded into these artificial exosomes via various physical or chemical methods [279]. Another avenue of research is genetic modification of cancer cells that naturally secrete EVs targeted to LECs. The payload of such EVs could be genetically modified or even replaced to serve a therapeutic purpose. The proof of concept for this approach was recently demonstrated with genetically engineered ovarian cancer cells stably overexpressing the tumor-suppressor miR-92b-3p. The engineered cells successfully packaged miR-92b-3p into exosomes and delivered the miRNA to human umbilical vein endothelial cells (HUVECs), exhibiting potent anti-angiogenic and anti-tumor activity in vitro and in vivo [280].
Lymphatics serve as a critical conduit for transporting dietary lipids from the gastrointestinal tract into systemic circulation. A typical route for intestinal fat absorption is the chylomicron pathway, which transports dietary lipids from enterocytes to lymphatics in the form of large triglyceride-rich lipoprotein particles known as chylomicrons [281, 282]. Another important route that allows particles such as pathogens to reach the intestinal lymphatics is the microfold cell (M cell) pathway. M cells are located within the intestinal epithelial layer in Peyer’s patches, groups of mucosa-associated lymphoid follicles mainly found in the ileum [283, 284]. Their primary function is to capture antigens from the intestinal lumen, actively transport them to the sub-mucosal lymphoid tissues, and present them to immune cells such as dendritic cells and lymphocytes [285, 286]. Therefore, M cells are increasingly viewed as a potential portal for oral drug delivery to the lymphatic system via the lymphatic capillaries surrounding Peyer’s patches [287,288,289]. These two pathways, especially the chylomicron pathway, could be utilized to deliver lipid nanoparticles containing therapeutic cargo to intestinal lymphatics and further to the lymph nodes. This strategy has potential to be developed into a method for the lymphatic delivery of therapeutic molecules targeting lncRNAs. For more details on the chylomicron-based, M cell-based and other lymphatic delivery strategies, we refer the reader to two recent comprehensive reviews [288, 290].
In addition to lipid-based EVs, several other types of delivery vectors have been proposed for lncRNA-targeting therapeutics [252]. They include various polymer-based nanoparticles and micelles, dendrimers, carbon nanotubes, and nanoparticles comprising metals or metalloids [221, 252]. Many of these nanoparticle formulations have already been successfully used for lymphatic delivery [290]. At first glance, these vectors might seem less promising because they lack LEC-specific targeting. However, this problem can be addressed by conjugating nanoparticles with LEC-specific homing molecules such as antibodies. For example, selective targeting to LECs has been achieved by conjugating polyethylene glycol-coated magnetic nanoparticles with a monoclonal antibody to lymphatic vessel endothelial hyaluronan receptor 1 (LYVE-1) [291], a selective marker of the lymphatic endothelium [292]. A similar strategy was recently employed for selective delivery of lipid nanoparticles containing small interfering RNA (siRNA) into LECs. The nanoparticles were conjugated with a monoclonal antibody to the LEC-specific marker podoplanin (PDPN) [293], thus providing an alternative to the above-discussed addition of targeting ligands to the particle surface. Furthermore, antibodies can be directly conjugated to therapeutic oligonucleotides such as siRNAs to facilitate targeted delivery to specific tissues and cell types without the use of a vector [294]. This approach to siRNA delivery into LECs, however, has a major limitation: the siRNA-antibody conjugates are often trapped in the endocytic compartment, thereby reducing their functional activity [295]. Selective targeting of therapeutic oligonucleotides or nanoparticles to LECs can be also achieved by conjugating them to aptamers directed against lymphatic markers such as LYVE-1, PDPN or VEGFR-3. Aptamers, also known as “chemical antibodies”, are single-stranded oligonucleotides selected for their affinity to a protein or other biomolecule of interest by a repetitive in vitro process called systematic evolution of ligands by exponential enrichment (SELEX) [296]. Since aptamers are 15 to 20 times smaller than antibodies, they offer superior normal tissue and tumor penetration [297] and therefore appear more promising than antibodies for the selective delivery of therapeutic molecules to specific sites in the body. Homing peptides represent yet another strategy for selectively targeting therapeutic molecules and nanocarriers to the lymphatics. They have, like aptamers, a much smaller molecular size than antibodies, which allows for better tissue penetration, faster clearance and lower immunogenicity [298]. The cyclic peptide Ly‐1P (CGNKRTRGC), first identified by Laakkonen et al. [299], represents a good example of such a homing peptide. Ly‐1P effectively targets tumor‐associated lymphatics, tumor cells and tumor-associated macrophages [299, 300], underscoring the promising clinical value of this lymphatic-homing peptide for imaging and therapy of tumors [301]. Furthermore, conjugation of Ly‐1P to nanoparticles [302], polymeric micelles [303] and liposomes [304] has been shown to facilitate the successful delivery of these nanocarriers to tumor lymphatics, resulting in the targeted release of anticancer drugs at metastatic tumor sites in vivo [303, 305]. Thus, the conjugation of lymphatic-homing peptides to nanocarriers may be a promising strategy for the lymphatic delivery of lncRNA-targeting therapeutics.
Despite the great potential of the above-discussed nanocarrier systems, viral vectors currently represent the most efficient means of delivering therapeutic nucleic acids into LECs. Several types of viral vectors such as genetically modified adenoviruses, adeno-associated viruses, retroviruses or lentiviruses are suitable for mediating RNA interference (RNAi) in multiple cell types including LECs. Each vector type has its advantages and disadvantages [306, 307] and should be selected in accordance with the therapeutic objective. The successful use of viral vectors to target lncRNAs has already been described in the literature—for example, knockdown of the lncRNA HOTAIR by lentiviral vector-mediated RNAi inhibited proliferation and invasion of endometrial carcinoma cells in vitro and in vivo [308]. Nevertheless, viral vectors are not without limitations. Innate and adaptive immune responses present obstacles to clinical application [309]; furthermore, viral vectors generally lack cell type specificity and may be difficult and costly to mass produce. In summary, a number of delivery systems based on lipid, polymeric or inorganic nanoparticles as well as viral vectors hold promise for the selective delivery of therapeutic molecules targeting lncRNAs to the lymphatic vasculature. Similar delivery systems could be used to target oncogenic lncRNAs in cancer cells, thereby inhibiting tumor lymphangiogenesis and lymphatic metastasis.
Conclusion
To date, our understanding of the molecular mechanisms underlying lymphangiogenesis has been primarily obtained from analysis of various proteins and protein signaling cascades. Recent research has revealed that long non-coding RNAs, a heterogeneous and functionally diverse class of molecules, have a previously unsuspected regulatory role in this process. Two new subclasses of pro-lymphangiogenic and anti-lymphangiogenic lncRNAs are emerging as a novel avenue for manipulating the lymphatic vasculature. In the cancer setting, these two groups of lncRNAs act as oncogenes or tumor suppressors by promoting or inhibiting tumor lymphangiogenesis and lymphatic metastasis. Furthermore, there is now evidence for a third, lineage-specific group of lncRNAs that are specifically expressed in normal, non-cancerous lymphatic endothelium and are critically involved in the regulation of the lymphatic-specific molecular and cellular mechanisms. The discovery of the first lymphatic-specific lncRNA LETR1 not only enhances our understanding of how lymphatic development and function are controlled, but also raises new questions. For example, the fact that LETR1 acts as gatekeeper of the LEC transcriptome leaves us with the important question: how is LETR1 itself regulated and is there a feedback loop between LETR1 and its targets? Another interesting question is whether lymphatic-specific lncRNAs have a role in non-cancerous lymphatic pathologies such as lymphedema. While we have barely scratched the surface of the complex lncRNA-mediated mechanisms governing lymphangiogenesis, we are already beginning to understand the therapeutic and diagnostic potential of lncRNAs in a variety of lymphatic-associated diseases including metastatic cancer. At present, there is only limited experience of therapeutic targeting of lncRNAs and the underlying methodological concepts are still mainly at the laboratory stage. One problem will be to identify the optimal delivery system combining high efficacy with lymphatic selectivity. Yet, despite the many challenges, the next few decades are likely to see substantial advances in the treatment of lymphatic-associated diseases based on highly specific or even individualized lncRNA-targeted therapies.
Data availability
Not applicable.
References
Djebali S, Davis CA, Merkel A et al (2012) Landscape of transcription in human cells. Nature 489:101–108. https://doi.org/10.1038/nature11233
Pertea M, Shumate A, Pertea G et al (2018) CHESS: a new human gene catalog curated from thousands of large-scale RNA sequencing experiments reveals extensive transcriptional noise. Genome Biol 19:208. https://doi.org/10.1186/s13059-018-1590-2
Dhamija S, Menon MB (2018) Non-coding transcript variants of protein-coding genes—what are they good for? RNA Biol 15:1025–1031. https://doi.org/10.1080/15476286.2018.1511675
Mattick JS, Amaral PP, Carninci P et al (2023) Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat Rev Mol Cell Biol. https://doi.org/10.1038/s41580-022-00566-8
Mercer TR, Mattick JS (2013) Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol 20:300–307. https://doi.org/10.1038/nsmb.2480
Bunch H (2018) Gene regulation of mammalian long non-coding RNA. Mol Genet Genom 293:1–15. https://doi.org/10.1007/s00438-017-1370-9
Geisler S, Coller J (2013) RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol 14:699–712. https://doi.org/10.1038/nrm3679
Schmitz SU, Grote P, Herrmann BG (2016) Mechanisms of long noncoding RNA function in development and disease. Cell Mol Life Sci 73:2491–2509. https://doi.org/10.1007/s00018-016-2174-5
Statello L, Guo CJ, Chen LL, Huarte M (2021) Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol 22:96–118. https://doi.org/10.1038/s41580-020-00315-9
Karakas D, Ozpolat B (2021) The role of LncRNAs in translation. Noncoding RNA. https://doi.org/10.3390/ncrna7010016
Ali T, Grote P (2020) Beyond the RNA-dependent function of LncRNA genes. Elife. https://doi.org/10.7554/eLife.60583
Oo JA, Brandes RP, Leisegang MS (2022) Long non-coding RNAs: novel regulators of cellular physiology and function. Pflugers Arch 474:191–204. https://doi.org/10.1007/s00424-021-02641-z
Grote P, Wittler L, Hendrix D et al (2013) The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell 24:206–214. https://doi.org/10.1016/j.devcel.2012.12.012
Allou L, Balzano S, Magg A et al (2021) Non-coding deletions identify Maenli lncRNA as a limb-specific En1 regulator. Nature 592:93–98. https://doi.org/10.1038/s41586-021-03208-9
Kitagawa M, Kitagawa K, Kotake Y, Niida H, Ohhata T (2013) Cell cycle regulation by long non-coding RNAs. Cell Mol Life Sci 70:4785–4794. https://doi.org/10.1007/s00018-013-1423-0
Ballarino M, Morlando M, Fatica A, Bozzoni I (2016) Non-coding RNAs in muscle differentiation and musculoskeletal disease. J Clin Investig 126:2021–2030. https://doi.org/10.1172/JCI84419
Sun X, Wong D (2016) Long non-coding RNA-mediated regulation of glucose homeostasis and diabetes. Am J Cardiovasc Dis 6:17–25
Lewandowski JP, Lee JC, Hwang T et al (2019) The Firre locus produces a trans-acting RNA molecule that functions in hematopoiesis. Nat Commun 10:5137. https://doi.org/10.1038/s41467-019-12970-4
Loda A, Heard E (2019) Xist RNA in action: Past, present, and future. PLoS Genet 15:e1008333. https://doi.org/10.1371/journal.pgen.1008333
Clemson CM, Hutchinson JN, Sara SA, Ensminger AW, Fox AH, Chess A, Lawrence JB (2009) An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell 33:717–726. https://doi.org/10.1016/j.molcel.2009.01.026
Trembinski DJ, Bink DI, Theodorou K et al (2020) Aging-regulated anti-apoptotic long non-coding RNA Sarrah augments recovery from acute myocardial infarction. Nat Commun 11:2039. https://doi.org/10.1038/s41467-020-15995-2
Erdogan I, Sweef O, Akgul B (2022) Long noncoding RNAs in human cancer and apoptosis. Curr Pharm Biotechnol. https://doi.org/10.2174/1389201023666220624094950
Atianand MK, Hu W, Satpathy AT et al (2016) A long noncoding RNA lincRNA-EPS acts as a transcriptional brake to restrain inflammation. Cell 165:1672–1685. https://doi.org/10.1016/j.cell.2016.05.075
Wang Y, Huang L, Wang Y et al (2020) Single-cell RNA-sequencing analysis identifies host long noncoding RNA MAMDC2-AS1 as a co-factor for HSV-1 nuclear transport. Int J Biol Sci 16:1586–1603. https://doi.org/10.7150/ijbs.42556
Schmitt AM, Chang HY (2016) Long noncoding RNAs in cancer pathways. Cancer Cell 29:452–463. https://doi.org/10.1016/j.ccell.2016.03.010
Winkler L, Dimitrova N (2022) A mechanistic view of long noncoding RNAs in cancer. Wiley Interdiscip Rev RNA 13:e1699. https://doi.org/10.1002/wrna.1699
Bridges MC, Daulagala AC, Kourtidis A (2021) LNCcation: lncRNA localization and function. J Cell Biol. https://doi.org/10.1083/jcb.202009045
Saxena A, Carninci P (2011) Long non-coding RNA modifies chromatin: epigenetic silencing by long non-coding RNAs. BioEssays 33:830–839. https://doi.org/10.1002/bies.201100084
Mele M, Rinn JL (2016) “Cat’s Cradling” the 3D genome by the act of LncRNA transcription. Mol Cell 62:657–664. https://doi.org/10.1016/j.molcel.2016.05.011
Goldfarb KC, Cech TR (2017) Targeted CRISPR disruption reveals a role for RNase MRP RNA in human preribosomal RNA processing. Genes Dev 31:59–71. https://doi.org/10.1101/gad.286963.116
Tripathi V, Ellis JD, Shen Z et al (2010) The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell 39:925–938. https://doi.org/10.1016/j.molcel.2010.08.011
Gong C, Maquat LE (2011) lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470:284–288. https://doi.org/10.1038/nature09701
Carrieri C, Cimatti L, Biagioli M et al (2012) Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature 491:454–457. https://doi.org/10.1038/nature11508
Yoon JH, Abdelmohsen K, Srikantan S et al (2012) LincRNA-p21 suppresses target mRNA translation. Mol Cell 47:648–655. https://doi.org/10.1016/j.molcel.2012.06.027
Cesana M, Cacchiarelli D, Legnini I et al (2011) A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 147:358–369. https://doi.org/10.1016/j.cell.2011.09.028
Du Z, Sun T, Hacisuleyman E et al (2016) Integrative analyses reveal a long noncoding RNA-mediated sponge regulatory network in prostate cancer. Nat Commun 7:10982. https://doi.org/10.1038/ncomms10982
Lee S, Kopp F, Chang TC et al (2016) Noncoding RNA NORAD regulates genomic stability by sequestering PUMILIO proteins. Cell 164:69–80. https://doi.org/10.1016/j.cell.2015.12.017
Ribeiro DM, Zanzoni A, Cipriano A et al (2018) Protein complex scaffolding predicted as a prevalent function of long non-coding RNAs. Nucleic Acids Res 46:917–928. https://doi.org/10.1093/nar/gkx1169
Lin A, Li C, Xing Z et al (2016) The LINK-A lncRNA activates normoxic HIF1alpha signalling in triple-negative breast cancer. Nat Cell Biol 18:213–224. https://doi.org/10.1038/ncb3295
Aznaourova M, Janga H, Sefried S et al (2020) Noncoding RNA MaIL1 is an integral component of the TLR4-TRIF pathway. Proc Natl Acad Sci USA 117:9042–9053. https://doi.org/10.1073/pnas.1920393117
Gomez JA, Wapinski OL, Yang YW et al (2013) The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-gamma locus. Cell 152:743–754. https://doi.org/10.1016/j.cell.2013.01.015
Collier SP, Henderson MA, Tossberg JT, Aune TM (2014) Regulation of the Th1 genomic locus from Ifng through Tmevpg1 by T-bet. J Immunol 193:3959–3965. https://doi.org/10.4049/jimmunol.1401099
Man HSJ, Sukumar AN, Lam GC et al (2018) Angiogenic patterning by STEEL, an endothelial-enriched long noncoding RNA. Proc Natl Acad Sci USA 115:2401–2406. https://doi.org/10.1073/pnas.1715182115
Zhang X, Hamblin MH, Yin KJ (2017) The long noncoding RNA Malat 1: Its physiological and pathophysiological functions. RNA Biol 14:1705–1714. https://doi.org/10.1080/15476286.2017.1358347
McPherson R, Pertsemlidis A, Kavaslar N et al (2007) A common allele on chromosome 9 associated with coronary heart disease. Science 316:1488–1491. https://doi.org/10.1126/science.1142447
Neumann P, Jae N, Knau A et al (2018) The lncRNA GATA6-AS epigenetically regulates endothelial gene expression via interaction with LOXL2. Nat Commun 9:237. https://doi.org/10.1038/s41467-017-02431-1
Stritt S, Koltowska K, Makinen T (2021) Homeostatic maintenance of the lymphatic vasculature. Trends Mol Med 27:955–970. https://doi.org/10.1016/j.molmed.2021.07.003
Randolph GJ, Ivanov S, Zinselmeyer BH, Scallan JP (2017) The lymphatic system: integral roles in immunity. Annu Rev Immunol 35:31–52. https://doi.org/10.1146/annurev-immunol-041015-055354
Petrova TV, Koh GY (2020) Biological functions of lymphatic vessels. Science. https://doi.org/10.1126/science.aax4063
Oliver G, Kipnis J, Randolph GJ, Harvey NL (2020) The lymphatic vasculature in the 21(st) century: novel functional roles in homeostasis and disease. Cell 182:270–296. https://doi.org/10.1016/j.cell.2020.06.039
Bernier-Latmani J, Petrova TV (2017) Intestinal lymphatic vasculature: structure, mechanisms and functions. Nat Rev Gastroenterol Hepatol 14:510–526. https://doi.org/10.1038/nrgastro.2017.79
Cserr HF, Harling-Berg CJ, Knopf PM (1992) Drainage of brain extracellular fluid into blood and deep cervical lymph and its immunological significance. Brain Pathol 2:269–276. https://doi.org/10.1111/j.1750-3639.1992.tb00703.x
Aspelund A, Antila S, Proulx ST et al (2015) A dural lymphatic vascular system that drains brain interstitial fluid and macromolecules. J Exp Med 212:991–999. https://doi.org/10.1084/jem.20142290
Ma Q, Ineichen BV, Detmar M, Proulx ST (2017) Outflow of cerebrospinal fluid is predominantly through lymphatic vessels and is reduced in aged mice. Nat Commun 8:1434. https://doi.org/10.1038/s41467-017-01484-6
Ma Q, Decker Y, Muller A, Ineichen BV, Proulx ST (2019) Clearance of cerebrospinal fluid from the sacral spine through lymphatic vessels. J Exp Med 216:2492–2502. https://doi.org/10.1084/jem.20190351
Jacob L, Boisserand LSB, Geraldo LHM et al (2019) Anatomy and function of the vertebral column lymphatic network in mice. Nat Commun 10:4594. https://doi.org/10.1038/s41467-019-12568-w
Rockson SG (2001) Lymphedema. Am J Med 110:288–295. https://doi.org/10.1016/s0002-9343(00)00727-0
Bittar S, Simman R, Lurie F (2020) Lymphedema: a practical approach and clinical update. Wounds 32:86–92
Harvey NL, Srinivasan RS, Dillard ME et al (2005) Lymphatic vascular defects promoted by Prox1 haploinsufficiency cause adult-onset obesity. Nat Genet 37:1072–1081. https://doi.org/10.1038/ng1642
Escobedo N, Oliver G (2017) The lymphatic vasculature: its role in adipose metabolism and obesity. Cell Metab 26:598–609. https://doi.org/10.1016/j.cmet.2017.07.020
Haemmerle M, Keller T, Egger G et al (2013) Enhanced lymph vessel density, remodeling, and inflammation are reflected by gene expression signatures in dermal lymphatic endothelial cells in type 2 diabetes. Diabetes 62:2509–2529. https://doi.org/10.2337/db12-0844
Jiang X, Tian W, Nicolls MR, Rockson SG (2019) The lymphatic system in obesity, insulin resistance, and cardiovascular diseases. Front Physiol 10:1402. https://doi.org/10.3389/fphys.2019.01402
Baluk P, Tammela T, Ator E et al (2005) Pathogenesis of persistent lymphatic vessel hyperplasia in chronic airway inflammation. J Clin Investig 115:247–257. https://doi.org/10.1172/JCI22037
Schwager S, Detmar M (2019) Inflammation and lymphatic function. Front Immunol 10:308. https://doi.org/10.3389/fimmu.2019.00308
Aspelund A, Robciuc MR, Karaman S, Makinen T, Alitalo K (2016) Lymphatic system in cardiovascular medicine. Circ Res 118:515–530. https://doi.org/10.1161/CIRCRESAHA.115.306544
Brakenhielm E, Alitalo K (2019) Cardiac lymphatics in health and disease. Nat Rev Cardiol 16:56–68. https://doi.org/10.1038/s41569-018-0087-8
Lim HY, Thiam CH, Yeo KP et al (2013) Lymphatic vessels are essential for the removal of cholesterol from peripheral tissues by SR-BI-mediated transport of HDL. Cell Metab 17:671–684. https://doi.org/10.1016/j.cmet.2013.04.002
Rademakers T, van der Vorst EP, Daissormont IT et al (2017) Adventitial lymphatic capillary expansion impacts on plaque T cell accumulation in atherosclerosis. Sci Rep 7:45263. https://doi.org/10.1038/srep45263
Milasan A, Smaani A, Martel C (2019) Early rescue of lymphatic function limits atherosclerosis progression in Ldlr(−/−) mice. Atherosclerosis 283:106–119. https://doi.org/10.1016/j.atherosclerosis.2019.01.031
Da Mesquita S, Louveau A, Vaccari A et al (2018) Functional aspects of meningeal lymphatics in ageing and Alzheimer’s disease. Nature 560:185–191. https://doi.org/10.1038/s41586-018-0368-8
Patel TK, Habimana-Griffin L, Gao X et al (2019) Dural lymphatics regulate clearance of extracellular tau from the CNS. Mol Neurodegener 14:11. https://doi.org/10.1186/s13024-019-0312-x
Zou W, Pu T, Feng W et al (2019) Blocking meningeal lymphatic drainage aggravates Parkinson’s disease-like pathology in mice overexpressing mutated alpha-synuclein. Transl Neurodegener 8:7. https://doi.org/10.1186/s40035-019-0147-y
Louveau A, Herz J, Alme MN et al (2018) CNS lymphatic drainage and neuroinflammation are regulated by meningeal lymphatic vasculature. Nat Neurosci 21:1380–1391. https://doi.org/10.1038/s41593-018-0227-9
Thomson BR, Heinen S, Jeansson M et al (2014) A lymphatic defect causes ocular hypertension and glaucoma in mice. J Clin Investig 124:4320–4324. https://doi.org/10.1172/JCI77162
Kim J, Park DY, Bae H et al (2017) Impaired angiopoietin/Tie2 signaling compromises Schlemm’s canal integrity and induces glaucoma. J Clin Investig 127:3877–3896. https://doi.org/10.1172/JCI94668
Tammela T, Alitalo K (2010) Lymphangiogenesis: molecular mechanisms and future promise. Cell 140:460–476. https://doi.org/10.1016/j.cell.2010.01.045
Karaman S, Detmar M (2014) Mechanisms of lymphatic metastasis. J Clin Investig 124:922–928. https://doi.org/10.1172/JCI71606
Stacker SA, Williams SP, Karnezis T, Shayan R, Fox SB, Achen MG (2014) Lymphangiogenesis and lymphatic vessel remodelling in cancer. Nat Rev Cancer 14:159–172. https://doi.org/10.1038/nrc3677
Rezzola S, Sigmund EC, Halin C, Ronca R (2022) The lymphatic vasculature: an active and dynamic player in cancer progression. Med Res Rev 42:576–614. https://doi.org/10.1002/med.21855
Jha SK, Rauniyar K, Jeltsch M (2018) Key molecules in lymphatic development, function, and identification. Ann Anat 219:25–34. https://doi.org/10.1016/j.aanat.2018.05.003
Gonzalez-Loyola A, Petrova TV (2021) Development and aging of the lymphatic vascular system. Adv Drug Deliv Rev 169:63–78. https://doi.org/10.1016/j.addr.2020.12.005
Ducoli L, Detmar M (2021) Beyond PROX1: transcriptional, epigenetic, and noncoding RNA regulation of lymphatic identity and function. Dev Cell 56:406–426. https://doi.org/10.1016/j.devcel.2021.01.018
Ducoli L, Agrawal S, Sibler E et al (2021) LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C. Nat Commun 12:925. https://doi.org/10.1038/s41467-021-21217-0
Evans PM, Liu C (2008) Roles of Krupel-like factor 4 in normal homeostasis, cancer and stem cells. Acta Biochim Biophys Sin (Shanghai) 40:554–564. https://doi.org/10.1111/j.1745-7270.2008.00439.x
Tamagnone L, Comoglio PM (2004) To move or not to move? Semaphorin signalling in cell migration. EMBO Rep 5:356–361. https://doi.org/10.1038/sj.embor.7400114
Banu N, Teichman J, Dunlap-Brown M, Villegas G, Tufro A (2006) Semaphorin 3C regulates endothelial cell function by increasing integrin activity. FASEB J 20:2150–2152. https://doi.org/10.1096/fj.05-5698fje
Laugesen A, Helin K (2014) Chromatin repressive complexes in stem cells, development, and cancer. Cell Stem Cell 14:735–751. https://doi.org/10.1016/j.stem.2014.05.006
Piunti A, Shilatifard A (2021) The roles of Polycomb repressive complexes in mammalian development and cancer. Nat Rev Mol Cell Biol 22:326–345. https://doi.org/10.1038/s41580-021-00341-1
Rauniyar K, Jha SK, Jeltsch M (2018) Biology of vascular endothelial growth factor C in the morphogenesis of lymphatic vessels. Front Bioeng Biotechnol 6:7. https://doi.org/10.3389/fbioe.2018.00007
Joukov V, Pajusola K, Kaipainen A et al (1996) A novel vascular endothelial growth factor, VEGF-C, is a ligand for the Flt4 (VEGFR-3) and KDR (VEGFR-2) receptor tyrosine kinases. EMBO J 15:290–298
Alitalo K (2011) The lymphatic vasculature in disease. Nat Med 17:1371–1380. https://doi.org/10.1038/nm.2545
Joukov V, Sorsa T, Kumar V et al (1997) Proteolytic processing regulates receptor specificity and activity of VEGF-C. EMBO J 16:3898–3911. https://doi.org/10.1093/emboj/16.13.3898
Kunnapuu J, Bokharaie H, Jeltsch M (2021) Proteolytic cleavages in the VEGF family: generating diversity among angiogenic VEGFs, essential for the activation of lymphangiogenic VEGFs. Biology (Basel). https://doi.org/10.3390/biology10020167
Karkkainen MJ, Haiko P, Sainio K et al (2004) Vascular endothelial growth factor C is required for sprouting of the first lymphatic vessels from embryonic veins. Nat Immunol 5:74–80. https://doi.org/10.1038/ni1013
Karaman S, Leppanen VM, Alitalo K (2018) Vascular endothelial growth factor signaling in development and disease. Development. https://doi.org/10.1242/dev.151019
Skobe M, Hawighorst T, Jackson DG et al (2001) Induction of tumor lymphangiogenesis by VEGF-C promotes breast cancer metastasis. Nat Med 7:192–198. https://doi.org/10.1038/84643
Mandriota SJ, Jussila L, Jeltsch M et al (2001) Vascular endothelial growth factor-C-mediated lymphangiogenesis promotes tumour metastasis. EMBO J 20:672–682. https://doi.org/10.1093/emboj/20.4.672
Hirakawa S, Brown LF, Kodama S, Paavonen K, Alitalo K, Detmar M (2007) VEGF-C-induced lymphangiogenesis in sentinel lymph nodes promotes tumor metastasis to distant sites. Blood 109:1010–1017. https://doi.org/10.1182/blood-2006-05-021758
Alitalo A, Detmar M (2012) Interaction of tumor cells and lymphatic vessels in cancer progression. Oncogene 31:4499–4508. https://doi.org/10.1038/onc.2011.602
Mushimiyimana I, Tomas Bosch V, Niskanen H et al (2021) Genomic landscapes of noncoding RNAs regulating VEGFA and VEGFC expression in endothelial cells. Mol Cell Biol 41:e0059420. https://doi.org/10.1128/MCB.00594-20
Seitz AK, Christensen LL, Christensen E et al (2017) Profiling of long non-coding RNAs identifies LINC00958 and LINC01296 as candidate oncogenes in bladder cancer. Sci Rep 7:395. https://doi.org/10.1038/s41598-017-00327-0
He W, Zhong G, Jiang N et al (2018) Long noncoding RNA BLACAT2 promotes bladder cancer-associated lymphangiogenesis and lymphatic metastasis. J Clin Investig 128:861–875. https://doi.org/10.1172/JCI96218
Guarnaccia AD, Tansey WP (2018) Moonlighting with WDR5: a cellular multitasker. J Clin Med. https://doi.org/10.3390/jcm7020021
Chen C, He W, Huang J et al (2018) LNMAT1 promotes lymphatic metastasis of bladder cancer via CCL2 dependent macrophage recruitment. Nat Commun 9:3826. https://doi.org/10.1038/s41467-018-06152-x
Farnsworth RH, Karnezis T, Maciburko SJ, Mueller SN, Stacker SA (2019) The interplay between lymphatic vessels and chemokines. Front Immunol 10:518. https://doi.org/10.3389/fimmu.2019.00518
Zheng H, Chen C, Luo Y et al (2021) Tumor-derived exosomal BCYRN1 activates WNT5A/VEGF-C/VEGFR3 feedforward loop to drive lymphatic metastasis of bladder cancer. Clin Transl Med 11:e497. https://doi.org/10.1002/ctm2.497
Asem MS, Buechler S, Wates RB, Miller DL, Stack MS (2016) Wnt5a signaling in cancer. Cancers (Basel). https://doi.org/10.3390/cancers8090079
Gore AV, Swift MR, Cha YR et al (2011) Rspo1/Wnt signaling promotes angiogenesis via Vegfc/Vegfr3. Development 138:4875–4886. https://doi.org/10.1242/dev.068460
Qu B, Liu BR, Du YJ, Chen J, Cheng YQ, Xu W, Wang XH (2014) Wnt/beta-catenin signaling pathway may regulate the expression of angiogenic growth factors in hepatocellular carcinoma. Oncol Lett 7:1175–1178. https://doi.org/10.3892/ol.2014.1828
Wang Y, Zhang W, Liu W et al (2021) Long noncoding RNA VESTAR regulates lymphangiogenesis and lymph node metastasis of esophageal squamous cell carcinoma by enhancing VEGFC mRNA stability. Cancer Res 81:3187–3199. https://doi.org/10.1158/0008-5472.CAN-20-1713
Muller-McNicoll M, Botti V, de Jesus Domingues AM et al (2016) SR proteins are NXF1 adaptors that link alternative RNA processing to mRNA export. Genes Dev 30:553–566. https://doi.org/10.1101/gad.276477.115
Dufies M, Giuliano S, Ambrosetti D et al (2017) Sunitinib stimulates expression of VEGFC by tumor cells and promotes lymphangiogenesis in clear cell renal cell carcinomas. Cancer Res 77:1212–1226. https://doi.org/10.1158/0008-5472.CAN-16-3088
Zhao M, Mishra L, Deng CX (2018) The role of TGF-beta/SMAD4 signaling in cancer. Int J Biol Sci 14:111–123. https://doi.org/10.7150/ijbs.23230
Li X, Lv X, Li Z et al (2019) Long noncoding RNA ASLNC07322 functions in VEGF-C expression regulated by Smad4 during colon cancer metastasis. Mol Ther Nucleic Acids 18:851–862. https://doi.org/10.1016/j.omtn.2019.10.012
Zhang F, Li J, Xiao H, Zou Y, Liu Y, Huang W (2018) AFAP1-AS1: a novel oncogenic long non-coding RNA in human cancers. Cell Prolif. https://doi.org/10.1111/cpr.12397
Xia M, Duan LJ, Lu BN, Pang YZ, Pang ZR (2021) LncRNA AFAP1-AS1/miR-27b-3p/VEGF-C axis modulates stemness characteristics in cervical cancer cells. Chin Med J (Engl) 134:2091–2101. https://doi.org/10.1097/CM9.0000000000001665
Liang H, Zhang C, Guan H, Liu J, Cui Y (2019) LncRNA DANCR promotes cervical cancer progression by upregulating ROCK1 via sponging miR-335-5p. J Cell Physiol 234:7266–7278. https://doi.org/10.1002/jcp.27484
Luo Y, Wang Q, Teng L et al (2020) LncRNA DANCR promotes proliferation and metastasis in pancreatic cancer by regulating miRNA-33b. FEBS Open Bio 10:18–27. https://doi.org/10.1002/2211-5463.12732
Zhan Y, Chen Z, Li Y et al (2018) Long non-coding RNA DANCR promotes malignant phenotypes of bladder cancer cells by modulating the miR-149/MSI2 axis as a ceRNA. J Exp Clin Cancer Res 37:273. https://doi.org/10.1186/s13046-018-0921-1
Ping Q, Shi Y, Yang M et al (2021) LncRNA DANCR regulates lymphatic metastasis of bladder cancer via the miR-335/VEGF-C axis. Transl Androl Urol 10:1743–1753. https://doi.org/10.21037/tau-21-226
Liu X, Zhang C, Wang X et al (2023) Long non-coding RNA MFSD4A-AS1 promotes lymphangiogenesis and lymphatic metastasis of papillary thyroid cancer. Endocr Relat Cancer. https://doi.org/10.1530/ERC-22-0221
Liu Y, Zhao F, Tan F et al (2022) HNF1A-AS1: a tumor-associated long non-coding RNA. Curr Pharm Des 28:1720–1729. https://doi.org/10.2174/1381612828666220520113846
Liu HT, Ma RR, Lv BB et al (2020) LncRNA-HNF1A-AS1 functions as a competing endogenous RNA to activate PI3K/AKT signalling pathway by sponging miR-30b-3p in gastric cancer. Br J Cancer 122:1825–1836. https://doi.org/10.1038/s41416-020-0836-4
Janku F, Yap TA, Meric-Bernstam F (2018) Targeting the PI3K pathway in cancer: are we making headway? Nat Rev Clin Oncol 15:273–291. https://doi.org/10.1038/nrclinonc.2018.28
Zhu C, Qi X, Chen Y, Sun B, Dai Y, Gu Y (2011) PI3K/Akt and MAPK/ERK1/2 signaling pathways are involved in IGF-1-induced VEGF-C upregulation in breast cancer. J Cancer Res Clin Oncol 137:1587–1594. https://doi.org/10.1007/s00432-011-1049-2
Xenou L, Papakonstanti EA (2020) p110delta PI3K as a therapeutic target of solid tumours. Clin Sci (Lond) 134:1377–1397. https://doi.org/10.1042/CS20190772
Kong Y, Li Y, Luo Y et al (2020) circNFIB1 inhibits lymphangiogenesis and lymphatic metastasis via the miR-486-5p/PIK3R1/VEGF-C axis in pancreatic cancer. Mol Cancer 19:82. https://doi.org/10.1186/s12943-020-01205-6
Greene J, Baird AM, Brady L, Lim M, Gray SG, McDermott R, Finn SP (2017) Circular RNAs: biogenesis, function and role in human diseases. Front Mol Biosci 4:38. https://doi.org/10.3389/fmolb.2017.00038
Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J (2019) The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet 20:675–691. https://doi.org/10.1038/s41576-019-0158-7
Xu C, Zhang J (2021) Mammalian circular RNAs result largely from splicing errors. Cell Rep 36:109439. https://doi.org/10.1016/j.celrep.2021.109439
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495:384–388. https://doi.org/10.1038/nature11993
Panda AC (2018) Circular RNAs act as miRNA sponges. Adv Exp Med Biol 1087:67–79. https://doi.org/10.1007/978-981-13-1426-1_6
Liu P, Wang Z, Ou X et al (2022) The FUS/circEZH2/KLF5/feedback loop contributes to CXCR4-induced liver metastasis of breast cancer by enhancing epithelial-mesenchymal transition. Mol Cancer 21:198. https://doi.org/10.1186/s12943-022-01653-2
Zheng S, Yang L, Zou Y et al (2020) Long non-coding RNA HUMT hypomethylation promotes lymphangiogenesis and metastasis via activating FOXK1 transcription in triple-negative breast cancer. J Hematol Oncol 13:17. https://doi.org/10.1186/s13045-020-00852-y
Dent R, Trudeau M, Pritchard KI et al (2007) Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res 13:4429–4434. https://doi.org/10.1158/1078-0432.CCR-06-3045
Evans MK, Matsui Y, Xu B et al (2020) Ybx1 fine-tunes PRC2 activities to control embryonic brain development. Nat Commun 11:4060. https://doi.org/10.1038/s41467-020-17878-y
He L, Gomes AP, Wang X et al (2018) mTORC1 promotes metabolic reprogramming by the suppression of GSK3-dependent Foxk1 phosphorylation. Mol Cell 70:949 e944-960 e944. https://doi.org/10.1016/j.molcel.2018.04.024
Manalo DJ, Rowan A, Lavoie T et al (2005) Transcriptional regulation of vascular endothelial cell responses to hypoxia by HIF-1. Blood 105:659–669. https://doi.org/10.1182/blood-2004-07-2958
Liang X, Yang D, Hu J, Hao X, Gao J, Mao Z (2008) Hypoxia inducible factor-alpha expression correlates with vascular endothelial growth factor-C expression and lymphangiogenesis/angiogenesis in oral squamous cell carcinoma. Anticancer Res 28:1659–1666
Ni X, Zhao Y, Ma J et al (2013) Hypoxia-induced factor-1 alpha upregulates vascular endothelial growth factor C to promote lymphangiogenesis and angiogenesis in breast cancer patients. J Biomed Res 27:478–485. https://doi.org/10.7555/JBR.27.20130021
Apte RS, Chen DS, Ferrara N (2019) VEGF in Signaling and disease: beyond discovery and development. Cell 176:1248–1264. https://doi.org/10.1016/j.cell.2019.01.021
Shi Y, Yang X, Xue X et al (2019) HANR promotes lymphangiogenesis of hepatocellular carcinoma via secreting miR-296 exosome and regulating EAG1/VEGFA signaling in HDLEC cells. J Cell Biochem 120:17699–17708. https://doi.org/10.1002/jcb.29036
Downie BR, Sanchez A, Knotgen H et al (2008) Eag1 expression interferes with hypoxia homeostasis and induces angiogenesis in tumors. J Biol Chem 283:36234–36240. https://doi.org/10.1074/jbc.M801830200
Gu J, Zhang B, An R et al (2022) Molecular interactions of the long noncoding RNA NEAT1 in cancer. Cancers (Basel). https://doi.org/10.3390/cancers14164009
Dong P, Xiong Y, Yue J, Hanley SJB, Kobayashi N, Todo Y, Watari H (2018) Long non-coding RNA NEAT1: a novel target for diagnosis and therapy in human tumors. Front Genet 9:471. https://doi.org/10.3389/fgene.2018.00471
Godet AC, Roussel E, David F et al (2022) Long non-coding RNA Neat1 and paraspeckle components are translational regulators in hypoxia. Elife. https://doi.org/10.7554/eLife.69162
Shin JW, Min M, Larrieu-Lahargue F et al (2006) Prox1 promotes lineage-specific expression of fibroblast growth factor (FGF) receptor-3 in lymphatic endothelium: a role for FGF signaling in lymphangiogenesis. Mol Biol Cell 17:576–584. https://doi.org/10.1091/mbc.e05-04-0368
Zhang H, Yu S, Fei K, Huang Z, Deng S, Xu H (2022) NEAT1 promotes the malignant development of bladder cancer by regulating the miR-101/VEGF-C pathway in vitro and in vivo. BMC Urol 22:193. https://doi.org/10.1186/s12894-022-01151-z
Chen C, Shen N, Chen Y et al (2021) LncCCLM inhibits lymphatic metastasis of cervical cancer by promoting STAU1-mediated IGF-1 mRNA degradation. Cancer Lett 518:169–179. https://doi.org/10.1016/j.canlet.2021.07.005
Li ZJ, Ying XJ, Chen HL et al (2013) Insulin-like growth factor-1 induces lymphangiogenesis and facilitates lymphatic metastasis in colorectal cancer. World J Gastroenterol 19:7788–7794. https://doi.org/10.3748/wjg.v19.i43.7788
Wilting J, Papoutsi M, Christ B et al (2002) The transcription factor Prox1 is a marker for lymphatic endothelial cells in normal and diseased human tissues. FASEB J 16:1271–1273. https://doi.org/10.1096/fj.01-1010fje
Horra A, Salazar J, Ferre R et al (2009) Prox-1 and FOXC2 gene expression in adipose tissue: a potential contributory role of the lymphatic system to familial combined hyperlipidaemia. Atherosclerosis 206:343–345. https://doi.org/10.1016/j.atherosclerosis.2009.02.026
Kretowski A, Adamska E, Maliszewska K et al (2015) The rs340874 PROX1 type 2 diabetes mellitus risk variant is associated with visceral fat accumulation and alterations in postprandial glucose and lipid metabolism. Genes Nutr 10:4. https://doi.org/10.1007/s12263-015-0454-6
Elsir T, Smits A, Lindstrom MS, Nister M (2012) Transcription factor PROX1: its role in development and cancer. Cancer Metastasis Rev 31:793–805. https://doi.org/10.1007/s10555-012-9390-8
Petrova TV, Nykanen A, Norrmen C et al (2008) Transcription factor PROX1 induces colon cancer progression by promoting the transition from benign to highly dysplastic phenotype. Cancer Cell 13:407–419. https://doi.org/10.1016/j.ccr.2008.02.020
Elsir T, Qu M, Berntsson SG et al (2011) PROX1 is a predictor of survival for gliomas WHO grade II. Br J Cancer 104:1747–1754. https://doi.org/10.1038/bjc.2011.162
Skog M, Bono P, Lundin M et al (2011) Expression and prognostic value of transcription factor PROX1 in colorectal cancer. Br J Cancer 105:1346–1351. https://doi.org/10.1038/bjc.2011.297
Ragusa S, Cheng J, Ivanov KI et al (2014) PROX1 promotes metabolic adaptation and fuels outgrowth of Wnt(high) metastatic colon cancer cells. Cell Rep 8:1957–1973. https://doi.org/10.1016/j.celrep.2014.08.041
Shimoda M, Takahashi M, Yoshimoto T, Kono T, Ikai I, Kubo H (2006) A homeobox protein, prox1, is involved in the differentiation, proliferation, and prognosis in hepatocellular carcinoma. Clin Cancer Res 12:6005–6011. https://doi.org/10.1158/1078-0432.CCR-06-0712
Laerm A, Helmbold P, Goldberg M, Dammann R, Holzhausen HJ, Ballhausen WG (2007) Prospero-related homeobox 1 (PROX1) is frequently inactivated by genomic deletions and epigenetic silencing in carcinomas of the bilary system. J Hepatol 46:89–97. https://doi.org/10.1016/j.jhep.2006.07.033
Chang TM, Hung WC (2012) Transcriptional repression of TWIST1 gene by Prospero-related homeobox 1 inhibits invasiveness of hepatocellular carcinoma cells. FEBS Lett 586:3746–3752. https://doi.org/10.1016/j.febslet.2012.08.034
Laitinen A, Bockelman C, Hagstrom J, Kokkola A, Kallio P, Haglund C (2017) High PROX1 expression in gastric cancer predicts better survival. PLoS ONE 12:e0183868. https://doi.org/10.1371/journal.pone.0183868
Pasmant E, Laurendeau I, Heron D, Vidaud M, Vidaud D, Bieche I (2007) Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense noncoding RNA whose expression coclusters with ARF. Cancer Res 67:3963–3969. https://doi.org/10.1158/0008-5472.CAN-06-2004
Cunnington MS, Santibanez Koref M, Mayosi BM, Burn J, Keavney B (2010) Chromosome 9p21 SNPs associated with multiple disease phenotypes correlate with ANRIL expression. PLoS Genet 6:e1000899. https://doi.org/10.1371/journal.pgen.1000899
Sun Z, Ou C, Ren W, Xie X, Li X, Li G (2016) Downregulation of long non-coding RNA ANRIL suppresses lymphangiogenesis and lymphatic metastasis in colorectal cancer. Oncotarget 7:47536–47555. https://doi.org/10.18632/oncotarget.9868
Maruyama K, Asai J, Ii M, Thorne T, Losordo DW, D’Amore PA (2007) Decreased macrophage number and activation lead to reduced lymphatic vessel formation and contribute to impaired diabetic wound healing. Am J Pathol 170:1178–1191. https://doi.org/10.2353/ajpath.2007.060018
He ZY, Wei TH, Zhang PH, Zhou J, Huang XY (2019) Long noncoding RNA-antisense noncoding RNA in the INK4 locus accelerates wound healing in diabetes by promoting lymphangiogenesis via regulating miR-181a/Prox1 axis. J Cell Physiol 234:4627–4640. https://doi.org/10.1002/jcp.27260
Kazenwadel J, Michael MZ, Harvey NL (2010) Prox1 expression is negatively regulated by miR-181 in endothelial cells. Blood 116:2395–2401. https://doi.org/10.1182/blood-2009-12-256297
Ma C, Shi X, Zhu Q, Li Q, Liu Y, Yao Y, Song Y (2016) The growth arrest-specific transcript 5 (GAS5): a pivotal tumor suppressor long noncoding RNA in human cancers. Tumour Biol 37:1437–1444. https://doi.org/10.1007/s13277-015-4521-9
Yang X, Xie Z, Lei X, Gan R (2020) Long non-coding RNA GAS5 in human cancer. Oncol Lett 20:2587–2594. https://doi.org/10.3892/ol.2020.11809
Sawaya AP, Pastar I, Stojadinovic O et al (2018) Topical mevastatin promotes wound healing by inhibiting the transcription factor c-Myc via the glucocorticoid receptor and the long non-coding RNA Gas5. J Biol Chem 293:1439–1449. https://doi.org/10.1074/jbc.M117.811240
He ZY, Huang MT, Cui X, Zhou ST, Wu Y, Zhang PH, Zhou J (2021) Long noncoding RNA GAS5 accelerates diabetic wound healing and promotes lymphangiogenesis via miR-217/Prox1 axis. Mol Cell Endocrinol 532:111283. https://doi.org/10.1016/j.mce.2021.111283
Srinivasan RS, Escobedo N, Yang Y et al (2014) The Prox1-Vegfr3 feedback loop maintains the identity and the number of lymphatic endothelial cell progenitors. Genes Dev 28:2175–2187. https://doi.org/10.1101/gad.216226.113
Sun C, Huang L, Li Z, Leng K, Xu Y, Jiang X, Cui Y (2018) Long non-coding RNA MIAT in development and disease: a new player in an old game. J Biomed Sci 25:23. https://doi.org/10.1186/s12929-018-0427-3
Wang H, Ding XG, Yang JJ et al (2018) LncRNA MIAT facilitated BM-MSCs differentiation into endothelial cells and restored erectile dysfunction via targeting miR-200a in a rat model of erectile dysfunction. Eur J Cell Biol 97:180–189. https://doi.org/10.1016/j.ejcb.2018.02.001
Dai XW, Luo W, Lv CL (2021) lncRNAMIAT facilitates the differentiation of adiposederived mesenchymal stem cells into lymphatic endothelial cells via the miR495/Prox1 axis. Mol Med Rep. https://doi.org/10.3892/mmr.2021.11962
Hwang JH, Kim IG, Lee JY et al (2011) Therapeutic lymphangiogenesis using stem cell and VEGF-C hydrogel. Biomaterials 32:4415–4423. https://doi.org/10.1016/j.biomaterials.2011.02.051
Yan A, Avraham T, Zampell JC, Haviv YS, Weitman E, Mehrara BJ (2011) Adipose-derived stem cells promote lymphangiogenesis in response to VEGF-C stimulation or TGF-beta1 inhibition. Future Oncol 7:1457–1473. https://doi.org/10.2217/fon.11.121
Toyserkani NM, Jensen CH, Sheikh SP, Sorensen JA (2016) Cell-assisted lipotransfer using autologous adipose-derived stromal cells for alleviation of breast cancer-related lymphedema. Stem Cells Transl Med 5:857–859. https://doi.org/10.5966/sctm.2015-0357
Chen C, Luo Y, He W et al (2020) Exosomal long noncoding RNA LNMAT2 promotes lymphatic metastasis in bladder cancer. J Clin Investig 130:404–421. https://doi.org/10.1172/JCI130892
Zu X, Tang Z, Li Y, Gao N, Ding J, Qi L (2006) Vascular endothelial growth factor-C expression in bladder transitional cell cancer and its relationship to lymph node metastasis. BJU Int 98:1090–1093. https://doi.org/10.1111/j.1464-410X.2006.06446.x
Suzuki K, Morita T, Tokue A (2005) Vascular endothelial growth factor-C (VEGF-C) expression predicts lymph node metastasis of transitional cell carcinoma of the bladder. Int J Urol 12:152–158. https://doi.org/10.1111/j.1442-2042.2005.01010.x
Rudzinska M, Czarnecka-Chrebelska KH, Kuznetsova EB et al (2021) Long non-coding PROX1-AS1 expression correlates with renal cell carcinoma metastasis and aggressiveness. Noncoding RNA. https://doi.org/10.3390/ncrna7020025
Zhao Q, Zhang B, Li Z, Tang W, Du L, Sang H (2022) Effects of IncRNA PROX1-AS1 on proliferation, migration, invasion and apoptosis of lung cancer cells by regulating MiR-1305. J Healthc Eng 2022:9570900. https://doi.org/10.1155/2022/9570900
Francois M, Caprini A, Hosking B et al (2008) Sox18 induces development of the lymphatic vasculature in mice. Nature 456:643–647. https://doi.org/10.1038/nature07391
Srinivasan RS, Geng X, Yang Y et al (2010) The nuclear hormone receptor Coup-TFII is required for the initiation and early maintenance of Prox1 expression in lymphatic endothelial cells. Genes Dev 24:696–707. https://doi.org/10.1101/gad.1859310
Chen C, Zheng H, Luo Y et al (2021) SUMOylation promotes extracellular vesicle-mediated transmission of lncRNA ELNAT1 and lymph node metastasis in bladder cancer. J Clin Investig. https://doi.org/10.1172/JCI146431
Misra JR, Irvine KD (2018) The Hippo signaling network and its biological functions. Annu Rev Genet 52:65–87. https://doi.org/10.1146/annurev-genet-120417-031621
Ma S, Meng Z, Chen R, Guan KL (2019) The Hippo pathway: biology and pathophysiology. Annu Rev Biochem 88:577–604. https://doi.org/10.1146/annurev-biochem-013118-111829
Zanconato F, Cordenonsi M, Piccolo S (2016) YAP/TAZ at the roots of cancer. Cancer Cell 29:783–803. https://doi.org/10.1016/j.ccell.2016.05.005
Cho H, Kim J, Ahn JH, Hong YK, Makinen T, Lim DS, Koh GY (2019) YAP and TAZ negatively regulate Prox1 during developmental and pathologic lymphangiogenesis. Circ Res 124:225–242. https://doi.org/10.1161/CIRCRESAHA.118.313707
Cha B, Moon S, Kim W (2021) A novel role of Hippo-Yap/TAZ signaling pathway in lymphatic vascular development. BMB Rep 54:285–294. https://doi.org/10.5483/BMBRep.2021.54.6.020
Liu X, Fu Q, Bian X et al (2021) Long non-coding RNA MAPK8IP1P2 inhibits lymphatic metastasis of thyroid cancer by activating Hippo signaling via sponging miR-146b-3p. Front Oncol. https://doi.org/10.3389/fonc.2020.600927
Garcia-Rendueles ME, Ricarte-Filho JC, Untch BR et al (2015) NF2 loss promotes oncogenic RAS-induced thyroid cancers via YAP-dependent transactivation of RAS proteins and sensitizes them to MEK inhibition. Cancer Discov 5:1178–1193. https://doi.org/10.1158/2159-8290.CD-15-0330
Wu J, Minikes AM, Gao M et al (2019) Intercellular interaction dictates cancer cell ferroptosis via NF2-YAP signalling. Nature 572:402–406. https://doi.org/10.1038/s41586-019-1426-6
Hamaratoglu F, Willecke M, Kango-Singh M et al (2006) The tumour-suppressor genes NF2/Merlin and expanded act through Hippo signalling to regulate cell proliferation and apoptosis. Nat Cell Biol 8:27–36. https://doi.org/10.1038/ncb1339
Zhang N, Bai H, David KK et al (2010) The Merlin/NF2 tumor suppressor functions through the YAP oncoprotein to regulate tissue homeostasis in mammals. Dev Cell 19:27–38. https://doi.org/10.1016/j.devcel.2010.06.015
Praskova M, Khoklatchev A, Ortiz-Vega S, Avruch J (2004) Regulation of the MST1 kinase by autophosphorylation, by the growth inhibitory proteins, RASSF1 and NORE1, and by Ras. Biochem J 381:453–462. https://doi.org/10.1042/BJ20040025
Matallanas D, Romano D, Yee K et al (2007) RASSF1A elicits apoptosis through an MST2 pathway directing proapoptotic transcription by the p73 tumor suppressor protein. Mol Cell 27:962–975. https://doi.org/10.1016/j.molcel.2007.08.008
Dhanaraman T, Singh S, Killoran RC, Singh A, Xu X, Shifman JM, Smith MJ (2020) RASSF effectors couple diverse RAS subfamily GTPases to the Hippo pathway. Sci Signal. https://doi.org/10.1126/scisignal.abb4778
Petrova TV, Karpanen T, Norrmén C et al (2004) Defective valves and abnormal mural cell recruitment underlie lymphatic vascular failure in lymphedema distichiasis. Nat Med 10:974–981. https://doi.org/10.1038/nm1094
Sabine A, Bovay E, Demir CS et al (2015) FOXC2 and fluid shear stress stabilize postnatal lymphatic vasculature. J Clin Investig 125:3861–3877. https://doi.org/10.1172/JCI80454
Norrmén C, Vandevelde W, Ny A et al (2010) Liprin (beta)1 is highly expressed in lymphatic vasculature and is important for lymphatic vessel integrity. Blood 115:906–909. https://doi.org/10.1182/blood-2009-03-212274
Brice G, Mansour S, Bell R et al (2002) Analysis of the phenotypic abnormalities in lymphoedema-distichiasis syndrome in 74 patients with FOXC2 mutations or linkage to 16q24. J Med Genet 39:478–483
Fang J, Dagenais SL, Erickson RP et al (2000) Mutations in FOXC2 (MFH-1), a forkhead family transcription factor, are responsible for the hereditary lymphedema-distichiasis syndrome. Am J Hum Genet 67:1382–1388. https://doi.org/10.1086/316915
Wang T, Zheng L, Wang Q, Hu YW (2018) Emerging roles and mechanisms of FOXC2 in cancer. Clin Chim Acta 479:84–93. https://doi.org/10.1016/j.cca.2018.01.019
Zhu KP, Zhang CL, Shen GQ, Zhu ZS (2015) Long noncoding RNA expression profiles of the doxorubicin-resistant human osteosarcoma cell line MG63/DXR and its parental cell line MG63 as ascertained by microarray analysis. Int J Clin Exp Pathol 8:8754–8773
Zhang CL, Zhu KP, Ma XL (2017) Antisense lncRNA FOXC2-AS1 promotes doxorubicin resistance in osteosarcoma by increasing the expression of FOXC2. Cancer Lett 396:66–75. https://doi.org/10.1016/j.canlet.2017.03.018
Yang H, Chen T, Xu S, Zhang S, Zhang M (2019) Long noncoding RNA FOXC2-AS1 predicts poor survival in breast cancer patients and promotes cell proliferation. Oncol Res 27:219–226. https://doi.org/10.3727/096504018X15213126075068
Sun Z, He C, Xiao M et al (2019) LncRNA FOXC2 antisense transcript accelerates non-small-cell lung cancer tumorigenesis via silencing p15. Am J Transl Res 11:4552–4560
Xu DF, Tao XH, Yu Y, Teng Y, Huang YM, Ma JW, Fan YB (2020) LncRNA FOXC2-AS1 stimulates proliferation of melanoma via silencing p15 by recruiting EZH2. Eur Rev Med Pharmacol Sci 24:8940–8946. https://doi.org/10.26355/eurrev_202009_22835
Pan K, Xie Y (2020) LncRNA FOXC2-AS1 enhances FOXC2 mRNA stability to promote colorectal cancer progression via activation of Ca(2+)-FAK signal pathway. Cell Death Dis 11:434. https://doi.org/10.1038/s41419-020-2633-7
Yan J, Liu J, Huang Z, Huang W, Lv J (2021) FOXC2-AS1 stabilizes FOXC2 mRNA via association with NSUN2 in gastric cancer cells. Hum Cell 34:1755–1764. https://doi.org/10.1007/s13577-021-00583-3
Chen Y, Gu M, Liu C, Wan X, Shi Q, Chen Q, Wang Z (2019) Long noncoding RNA FOXC2-AS1 facilitates the proliferation and progression of prostate cancer via targeting miR-1253/EZH2. Gene 686:37–42. https://doi.org/10.1016/j.gene.2018.10.085
Tan JZ, Yan Y, Wang XX, Jiang Y, Xu HE (2014) EZH2: biology, disease, and structure-based drug discovery. Acta Pharmacol Sin 35:161–174. https://doi.org/10.1038/aps.2013.161
Missaglia S, Tavian D, Michelini S, Maltese PE, Bonanomi A, Bertelli M (2021) Imbalance between expression of FOXC2 and its lncRNA in lymphedema-distichiasis caused by frameshift mutations. Genes (Basel). https://doi.org/10.3390/genes12050650
Groskopf J, Aubin SM, Deras IL et al (2006) APTIMA PCA3 molecular urine test: development of a method to aid in the diagnosis of prostate cancer. Clin Chem 52:1089–1095. https://doi.org/10.1373/clinchem.2005.063289
Auprich M, Bjartell A, Chun FK et al (2011) Contemporary role of prostate cancer antigen 3 in the management of prostate cancer. Eur Urol 60:1045–1054. https://doi.org/10.1016/j.eururo.2011.08.003
Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL (2011) Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25:1915–1927. https://doi.org/10.1101/gad.17446611
Zhao X, Lan Y, Chen D (2022) Exploring long non-coding RNA networks from single cell omics data. Comput Struct Biotechnol J 20:4381–4389. https://doi.org/10.1016/j.csbj.2022.08.003
Slaby O, Laga R, Sedlacek O (2017) Therapeutic targeting of non-coding RNAs in cancer. Biochem J 474:4219–4251. https://doi.org/10.1042/BCJ20170079
Fathi Dizaji B (2020) Strategies to target long non-coding RNAs in cancer treatment: progress and challenges. Egypt J Med Hum Genet 21:41. https://doi.org/10.1186/s43042-020-00074-4
Chen Y, Li Z, Chen X, Zhang S (2021) Long non-coding RNAs: from disease code to drug role. Acta Pharm Sin B 11:340–354. https://doi.org/10.1016/j.apsb.2020.10.001
Winkle M, El-Daly SM, Fabbri M, Calin GA (2021) Noncoding RNA therapeutics—challenges and potential solutions. Nat Rev Drug Discov 20:629–651. https://doi.org/10.1038/s41573-021-00219-z
Haniff HS, Knerr L, Chen JL, Disney MD, Lightfoot HL (2020) Target-directed approaches for screening small molecules against RNA targets. SLAS Discov 25:869–894. https://doi.org/10.1177/2472555220922802
Cote B, Rao D, Alany RG, Kwon GS, Alani AWG (2019) Lymphatic changes in cancer and drug delivery to the lymphatics in solid tumors. Adv Drug Deliv Rev 144:16–34. https://doi.org/10.1016/j.addr.2019.08.009
Setten RL, Rossi JJ, Han SP (2019) The current state and future directions of RNAi-based therapeutics. Nat Rev Drug Discov 18:421–446. https://doi.org/10.1038/s41573-019-0017-4
Kim K, Jutooru I, Chadalapaka G et al (2013) HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene 32:1616–1625. https://doi.org/10.1038/onc.2012.193
Sun R, Qin C, Jiang B et al (2016) Down-regulation of MALAT1 inhibits cervical cancer cell invasion and metastasis by inhibition of epithelial-mesenchymal transition. Mol Biosyst 12:952–962. https://doi.org/10.1039/c5mb00685f
Thorenoor N, Faltejskova-Vychytilova P, Hombach S, Mlcochova J, Kretz M, Svoboda M, Slaby O (2016) Long non-coding RNA ZFAS1 interacts with CDK1 and is involved in p53-dependent cell cycle control and apoptosis in colorectal cancer. Oncotarget 7:622–637. https://doi.org/10.18632/oncotarget.5807
Youn YH, Byun HJ, Yoon JH, Park CH, Lee SK (2019) Long noncoding RNA N-BLR upregulates the migration and invasion of gastric adenocarcinoma. Gut Liver 13:421–429. https://doi.org/10.5009/gnl18408
Jackson AL, Linsley PS (2010) Recognizing and avoiding siRNA off-target effects for target identification and therapeutic application. Nat Rev Drug Discov 9:57–67. https://doi.org/10.1038/nrd3010
Sledz CA, Holko M, de Veer MJ, Silverman RH, Williams BR (2003) Activation of the interferon system by short-interfering RNAs. Nat Cell Biol 5:834–839. https://doi.org/10.1038/ncb1038
Bridge AJ, Pebernard S, Ducraux A, Nicoulaz AL, Iggo R (2003) Induction of an interferon response by RNAi vectors in mammalian cells. Nat Genet 34:263–264. https://doi.org/10.1038/ng1173
Kim DH, Behlke MA, Rose SD, Chang MS, Choi S, Rossi JJ (2005) Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat Biotechnol 23:222–226. https://doi.org/10.1038/nbt1051
Siolas D, Lerner C, Burchard J et al (2005) Synthetic shRNAs as potent RNAi triggers. Nat Biotechnol 23:227–231. https://doi.org/10.1038/nbt1052
Akashi H, Matsumoto S, Taira K (2005) Gene discovery by ribozyme and siRNA libraries. Nat Rev Mol Cell Biol 6:413–422. https://doi.org/10.1038/nrm1646
Han Y, He F, Chen Y, Liu Y, Yu H (2018) SiRNA silencing efficacy prediction based on a deep architecture. BMC Genom 19:669. https://doi.org/10.1186/s12864-018-5028-8
La Rosa M, Fiannaca A, La Paglia L, Urso A (2022) A graph neural network approach for the analysis of siRNA-target biological networks. Int J Mol Sci. https://doi.org/10.3390/ijms232214211
Crooke ST (2017) Molecular mechanisms of antisense oligonucleotides. Nucleic Acid Ther 27:70–77. https://doi.org/10.1089/nat.2016.0656
Kurreck J, Wyszko E, Gillen C, Erdmann VA (2002) Design of antisense oligonucleotides stabilized by locked nucleic acids. Nucleic Acids Res 30:1911–1918. https://doi.org/10.1093/nar/30.9.1911
Lennox KA, Behlke MA (2016) Cellular localization of long non-coding RNAs affects silencing by RNAi more than by antisense oligonucleotides. Nucleic Acids Res 44:863–877. https://doi.org/10.1093/nar/gkv1206
Wahlestedt C (2013) Targeting long non-coding RNA to therapeutically upregulate gene expression. Nat Rev Drug Discov 12:433–446. https://doi.org/10.1038/nrd4018
Hagedorn PH, Brown JM, Easton A et al (2022) Acute neurotoxicity of antisense oligonucleotides after intracerebroventricular injection into mouse brain can be predicted from sequence features. Nucleic Acid Ther 32:151–162. https://doi.org/10.1089/nat.2021.0071
Hagedorn PH, Yakimov V, Ottosen S et al (2013) Hepatotoxic potential of therapeutic oligonucleotides can be predicted from their sequence and modification pattern. Nucleic Acid Ther 23:302–310. https://doi.org/10.1089/nat.2013.0436
Schubert S, Kurreck J (2004) Ribozyme- and deoxyribozyme-strategies for medical applications. Curr Drug Targets 5:667–681. https://doi.org/10.2174/1389450043345092
Huang X, Zhao Y, Pu Q et al (2019) Intracellular selection of trans-cleaving hammerhead ribozymes. Nucleic Acids Res 47:2514–2522. https://doi.org/10.1093/nar/gkz018
Phylactou LA, Kilpatrick MW, Wood MJ (1998) Ribozymes as therapeutic tools for genetic disease. Hum Mol Genet 7:1649–1653. https://doi.org/10.1093/hmg/7.10.1649
Phylactou LA, Tsipouras P, Kilpatrick MW (1998) Hammerhead ribozymes targeted to the FBN1 mRNA can discriminate a single base mismatch between ribozyme and target. Biochem Biophys Res Commun 249:804–810. https://doi.org/10.1006/bbrc.1998.9241
Burnett JC, Rossi JJ (2009) Stem cells, ribozymes and HIV. Gene Ther 16:1178–1179. https://doi.org/10.1038/gt.2009.86
Peng H, Latifi B, Muller S, Luptak A, Chen IA (2021) Self-cleaving ribozymes: substrate specificity and synthetic biology applications. RSC Chem Biol 2:1370–1383. https://doi.org/10.1039/d0cb00207k
Jiang MC, Ni JJ, Cui WY, Wang BY, Zhuo W (2019) Emerging roles of lncRNA in cancer and therapeutic opportunities. Am J Cancer Res 9:1354–1366
Urnov FD, Miller JC, Lee YL et al (2005) Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651. https://doi.org/10.1038/nature03556
Bedell VM, Wang Y, Campbell JM et al (2012) In vivo genome editing using a high-efficiency TALEN system. Nature 491:114–118. https://doi.org/10.1038/nature11537
Ioannidi EI, Yarnall MTN, Schmitt-Ulms C et al (2021) Drag-and-drop genome insertion without DNA cleavage with CRISPR-directed integrases. bioRxiv 2021.2011.2001.466786. https://doi.org/10.1101/2021.11.01.466786
Liu G, Lin Q, Jin S, Gao C (2022) The CRISPR–Cas toolbox and gene editing technologies. Mol Cell 82:333–347. https://doi.org/10.1016/j.molcel.2021.12.002
Gutschner T, Hammerle M, Eissmann M et al (2013) The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res 73:1180–1189. https://doi.org/10.1158/0008-5472.CAN-12-2850
Fu Y, Foden JA, Khayter C, Maeder ML, Reyon D, Joung JK, Sander JD (2013) High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol 31:822–826. https://doi.org/10.1038/nbt.2623
Kuscu C, Arslan S, Singh R, Thorpe J, Adli M (2014) Genome-wide analysis reveals characteristics of off-target sites bound by the Cas9 endonuclease. Nat Biotechnol 32:677–683. https://doi.org/10.1038/nbt.2916
Boettcher M, McManus MT (2015) Choosing the right tool for the job: RNAi, TALEN, or CRISPR. Mol Cell 58:575–585. https://doi.org/10.1016/j.molcel.2015.04.028
Li C, Chu W, Gill RA et al (2022) Computational tools and resources for CRISPR/Cas genome editing. Genom Proteom Bioinform. https://doi.org/10.1016/j.gpb.2022.02.006
Bao XR, Pan Y, Lee CM, Davis TH, Bao G (2021) Tools for experimental and computational analyses of off-target editing by programmable nucleases. Nat Protoc 16:10–26. https://doi.org/10.1038/s41596-020-00431-y
Anzalone AV, Koblan LW, Liu DR (2020) Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat Biotechnol 38:824–844. https://doi.org/10.1038/s41587-020-0561-9
Ates I, Rathbone T, Stuart C, Bridges PH, Cottle RN (2020) Delivery approaches for therapeutic genome editing and challenges. Genes (Basel). https://doi.org/10.3390/genes11101113
Raguram A, Banskota S, Liu DR (2022) Therapeutic in vivo delivery of gene editing agents. Cell 185:2806–2827. https://doi.org/10.1016/j.cell.2022.03.045
Chen J, Wang L, Yao Q, Ling R, Li K, Wang H (2004) Drug concentrations in axillary lymph nodes after lymphatic chemotherapy on patients with breast cancer. Breast Cancer Res 6:R474-477. https://doi.org/10.1186/bcr819
Wilson KD, Raney SG, Sekirov L, Chikh G, deJong SD, Cullis PR, Tam YK (2007) Effects of intravenous and subcutaneous administration on the pharmacokinetics, biodistribution, cellular uptake and immunostimulatory activity of CpG ODN encapsulated in liposomal nanoparticles. Int Immunopharmacol 7:1064–1075. https://doi.org/10.1016/j.intimp.2007.04.002
Trevaskis NL, Kaminskas LM, Porter CJ (2015) From sewer to saviour—targeting the lymphatic system to promote drug exposure and activity. Nat Rev Drug Discov 14:781–803. https://doi.org/10.1038/nrd4608
Oussoren C, Zuidema J, Crommelin DJ, Storm G (1997) Lymphatic uptake and biodistribution of liposomes after subcutaneous injection. II. Influence of liposomal size, lipid compostion and lipid dose. Biochim Biophys Acta 1328:261–272. https://doi.org/10.1016/s0005-2736(97)00122-3
Sarin H (2010) Physiologic upper limits of pore size of different blood capillary types and another perspective on the dual pore theory of microvascular permeability. J Angiogenes Res 2:14. https://doi.org/10.1186/2040-2384-2-14
Rohner NA, Thomas SN (2017) Flexible macromolecule versus rigid particle retention in the injected skin and accumulation in draining lymph nodes are differentially influenced by hydrodynamic size. ACS Biomater Sci Eng 3:153–159. https://doi.org/10.1021/acsbiomaterials.6b00438
Cheng Z, Que H, Chen L, Sun Q, Wei X (2022) Nanomaterial-based drug delivery system targeting lymph nodes. Pharmaceutics. https://doi.org/10.3390/pharmaceutics14071372
Schudel A, Francis DM, Thomas SN (2019) Material design for lymph node drug delivery. Nat Rev Mater 4:415–428. https://doi.org/10.1038/s41578-019-0110-7
Manspeaker MP, Thomas SN (2020) Lymphatic immunomodulation using engineered drug delivery systems for cancer immunotherapy. Adv Drug Deliv Rev 160:19–35. https://doi.org/10.1016/j.addr.2020.10.004
Lin Y, Wu J, Gu W, Huang Y, Tong Z, Huang L, Tan J (2018) Exosome-liposome hybrid nanoparticles deliver CRISPR/Cas9 system in MSCs. Adv Sci (Weinh) 5:1700611. https://doi.org/10.1002/advs.201700611
Wang SH, Liou GG, Liu SH et al (2019) Laminin gamma2-enriched extracellular vesicles of oral squamous cell carcinoma cells enhance in vitro lymphangiogenesis via integrin alpha3-dependent uptake by lymphatic endothelial cells. Int J Cancer 144:2795–2810. https://doi.org/10.1002/ijc.32027
Glassman PM, Myerson JW, Ferguson LT, Kiseleva RY, Shuvaev VV, Brenner JS, Muzykantov VR (2020) Targeting drug delivery in the vascular system: focus on endothelium. Adv Drug Deliv Rev 157:96–117. https://doi.org/10.1016/j.addr.2020.06.013
Li YJ, Wu JY, Liu J et al (2021) Artificial exosomes for translational nanomedicine. J Nanobiotechnol 19:242. https://doi.org/10.1186/s12951-021-00986-2
Fu S, Wang Y, Xia X, Zheng JC (2020) Exosome engineering: current progress in cargo loading and targeted delivery. Nanoimpact 20:100261
Wang J, Wang C, Li Y et al (2021) Potential of peptide-engineered exosomes with overexpressed miR-92b-3p in anti-angiogenic therapy of ovarian cancer. Clin Transl Med 11:e425. https://doi.org/10.1002/ctm2.425
Julve J, Martin-Campos JM, Escola-Gil JC, Blanco-Vaca F (2016) Chylomicrons: advances in biology, pathology, laboratory testing, and therapeutics. Clin Chim Acta 455:134–148. https://doi.org/10.1016/j.cca.2016.02.004
Xiao C, Stahel P, Lewis GF (2019) Regulation of chylomicron secretion: focus on post-assembly mechanisms. Cell Mol Gastroenterol Hepatol 7:487–501. https://doi.org/10.1016/j.jcmgh.2018.10.015
Hussain N, Jaitley V, Florence AT (2001) Recent advances in the understanding of uptake of microparticulates across the gastrointestinal lymphatics. Adv Drug Deliv Rev 50:107–142. https://doi.org/10.1016/s0169-409x(01)00152-1
Brayden DJ, Jepson MA, Baird AW (2005) Keynote review: intestinal Peyer’s patch M cells and oral vaccine targeting. Drug Discov Today 10:1145–1157. https://doi.org/10.1016/S1359-6446(05)03536-1
Ohno H (2016) Intestinal M cells. J Biochem 159:151–160. https://doi.org/10.1093/jb/mvv121
Mabbott NA, Donaldson DS, Ohno H, Williams IR, Mahajan A (2013) Microfold (M) cells: important immunosurveillance posts in the intestinal epithelium. Mucosal Immunol 6:666–677. https://doi.org/10.1038/mi.2013.30
Lundquist P, Artursson P (2016) Oral absorption of peptides and nanoparticles across the human intestine: opportunities, limitations and studies in human tissues. Adv Drug Deliv Rev 106:256–276. https://doi.org/10.1016/j.addr.2016.07.007
Zhang Z, Lu Y, Qi J, Wu W (2021) An update on oral drug delivery via intestinal lymphatic transport. Acta Pharm Sin B 11:2449–2468. https://doi.org/10.1016/j.apsb.2020.12.022
Zhang Q, Li L, Hao S et al (2022) A lymphatic route for a hyperbranched heteroglycan from Radix Astragali to trigger immune responses after oral dosing. Carbohydr Polym 292:119653. https://doi.org/10.1016/j.carbpol.2022.119653
McCright J, Naiknavare R, Yarmovsky J, Maisel K (2022) Targeting lymphatics for nanoparticle drug delivery. Front Pharmacol 13:887402. https://doi.org/10.3389/fphar.2022.887402
Guo Q, Liu Y, Xu K, Ren K, Sun W (2013) Mouse lymphatic endothelial cell targeted probes: anti-LYVE-1 antibody-based magnetic nanoparticles. Int J Nanomed 8:2273–2284. https://doi.org/10.2147/IJN.S45817
Banerji S, Ni J, Wang SX et al (1999) LYVE-1, a new homologue of the CD44 glycoprotein, is a lymph-specific receptor for hyaluronan. J Cell Biol 144:789–801. https://doi.org/10.1083/jcb.144.4.789
Sakurai Y, Abe N, Yoshikawa K et al (2022) Targeted delivery of lipid nanoparticle to lymphatic endothelial cells via anti-podoplanin antibody. J Control Release. https://doi.org/10.1016/j.jconrel.2022.06.052
Sugo T, Terada M, Oikawa T et al (2016) Development of antibody-siRNA conjugate targeted to cardiac and skeletal muscles. J Control Release 237:1–13. https://doi.org/10.1016/j.jconrel.2016.06.036
Cuellar TL, Barnes D, Nelson C et al (2015) Systematic evaluation of antibody-mediated siRNA delivery using an industrial platform of THIOMAB-siRNA conjugates. Nucleic Acids Res 43:1189–1203. https://doi.org/10.1093/nar/gku1362
Nimjee SM, White RR, Becker RC, Sullenger BA (2017) Aptamers as therapeutics. Annu Rev Pharmacol Toxicol 57:61–79. https://doi.org/10.1146/annurev-pharmtox-010716-104558
Xiang D, Zheng C, Zhou SF et al (2015) Superior performance of aptamer in tumor penetration over antibody: implication of aptamer-based theranostics in solid tumors. Theranostics 5:1083–1097. https://doi.org/10.7150/thno.11711
Zhang F, Niu G, Lin X et al (2012) Imaging tumor-induced sentinel lymph node lymphangiogenesis with LyP-1 peptide. Amino Acids 42:2343–2351. https://doi.org/10.1007/s00726-011-0976-1
Laakkonen P, Porkka K, Hoffman JA, Ruoslahti E (2002) A tumor-homing peptide with a targeting specificity related to lymphatic vessels. Nat Med 8:751–755. https://doi.org/10.1038/nm720
Enback J, Laakkonen P (2007) Tumour-homing peptides: tools for targeting, imaging and destruction. Biochem Soc Trans 35:780–783. https://doi.org/10.1042/BST0350780
Song N, Zhao L, Zhu M, Zhao J (2019) Recent progress in LyP-1-based strategies for targeted imaging and therapy. Drug Deliv 26:363–375. https://doi.org/10.1080/10717544.2019.1587047
Luo G, Yu X, Jin C et al (2010) LyP-1-conjugated nanoparticles for targeting drug delivery to lymphatic metastatic tumors. Int J Pharm 385:150–156. https://doi.org/10.1016/j.ijpharm.2009.10.014
Wang Z, Yu Y, Ma J et al (2012) LyP-1 modification to enhance delivery of artemisinin or fluorescent probe loaded polymeric micelles to highly metastatic tumor and its lymphatics. Mol Pharm 9:2646–2657. https://doi.org/10.1021/mp3002107
Yan Z, Zhan C, Wen Z et al (2011) LyP-1-conjugated doxorubicin-loaded liposomes suppress lymphatic metastasis by inhibiting lymph node metastases and destroying tumor lymphatics. Nanotechnology 22:415103. https://doi.org/10.1088/0957-4484/22/41/415103
Yan Z, Wang F, Wen Z et al (2012) LyP-1-conjugated PEGylated liposomes: a carrier system for targeted therapy of lymphatic metastatic tumor. J Control Release 157:118–125. https://doi.org/10.1016/j.jconrel.2011.07.034
Couto LB, High KA (2010) Viral vector-mediated RNA interference. Curr Opin Pharmacol 10:534–542. https://doi.org/10.1016/j.coph.2010.06.007
Schott JW, Morgan M, Galla M, Schambach A (2016) Viral and synthetic RNA vector technologies and applications. Mol Ther 24:1513–1527. https://doi.org/10.1038/mt.2016.143
Huang J, Ke P, Guo L, Wang W, Tan H, Liang Y, Yao S (2014) Lentivirus-mediated RNA interference targeting the long noncoding RNA HOTAIR inhibits proliferation and invasion of endometrial carcinoma cells in vitro and in vivo. Int J Gynecol Cancer 24:635–642. https://doi.org/10.1097/IGC.0000000000000121
Shirley JL, de Jong YP, Terhorst C, Herzog RW (2020) Immune responses to viral gene therapy vectors. Mol Ther 28:709–722. https://doi.org/10.1016/j.ymthe.2020.01.001
Acknowledgements
We thank Prof. Tatiana V. Petrova, Department of Oncology and Ludwig Institute for Cancer Research, University of Lausanne, Switzerland, for critical reading of the manuscript and helpful comments. We also thank Katerina Burgess (Eurasian Linguistic Services Ltd.) for assistance with language editing. We apologize to colleagues whose work we could not cite due to space constraints or otherwise.
Funding
Open Access funding provided by the University of Helsinki, including the Helsinki University Central Hospital. This work was supported by the Russian Science Foundation (Grant # 21-75-30020).
Author information
Authors and Affiliations
Contributions
All authors substantially contributed to the conception and design of the manuscript and interpreting the literature data. The first draft of the manuscript was written by KI and all authors revised it critically for important intellectual content. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ivanov, K.I., Samuilova, O.V. & Zamyatnin, A.A. The emerging roles of long noncoding RNAs in lymphatic vascular development and disease. Cell. Mol. Life Sci. 80, 197 (2023). https://doi.org/10.1007/s00018-023-04842-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-023-04842-4