Abstract
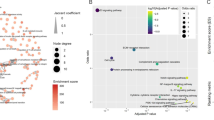
Idiopathic pulmonary fibrosis (IPF) is an interstitial lung disease characterized by chronic progressive pulmonary fibrosis and a poor prognosis. Genetic studies, including transcriptomic and proteomics, have provided new insight into revealing mechanisms of IPF. Herein we provided a novel strategy to identify biomarkers by integrative analysis of transcriptomic and proteomic profiles of IPF patients. We examined the landscape of IPF patients' gene expression in the transcription and translation phases and investigated the expression and functions of two new potential biomarkers. Differentially expressed (DE) mRNAs were mainly enriched in pathways associated with immune system activities and inflammatory responses, while DE proteins are related to extracellular matrix production and wound repair. The upregulated genes in both phases are associated with wound repair and cell differentiation, while the downregulated genes in both phases are associated with reduced immune activities and the damage of the alveolar tissues. On this basis, we identified thirteen potential marker genes. Among them, we validated the expression changes of butyrophilin-like 9 (BTNL9) and plasmolipin (PLLP) and investigated their functional pathways in the IPF mechanism. Both genes are downregulated in the tissues of IPF patients and Bleomycin-induced mice, and co-expression analysis indicates that they have a protective effect by inhibiting extracellular matrix production and promoting wound repair in alveolar epithelial cells.









Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available from the corresponding authors on reasonable request.
Code availability
The authors declare that code for data analysis in this study are available from the corresponding authors on reasonable request.
References
Meyer KC, Nathan SD (2019) Idiopathic pulmonary fibrosis. Humana Press, Cham
Lawson WE, Grant SW, Ambrosini V, Womble KE, Dawson EP, Lane KB et al (2004) Genetic mutations in surfactant protein C are a rare cause of sporadic cases of IPF. Thorax 59:977–980. https://doi.org/10.1136/thx.2004.026336
Wang Y, Kuan PJ, Xing C, Cronkhite JT, Torres F, Rosenblatt RL et al (2009) Genetic defects in surfactant protein A2 are associated with pulmonary fibrosis and lung cancer. Am J Hum Genet 84:52–59. https://doi.org/10.1016/j.ajhg.2008.11.010
Kropski JA, Blackwell TS, Loyd JE (2015) The genetic basis of idiopathic pulmonary fibrosis. Eur Respir J 45:1717–1727. https://doi.org/10.1183/09031936.00163814
Kaur A, Mathai SK, Schwartz DA (2017) Genetics in idiopathic pulmonary fibrosis pathogenesis, prognosis, and treatment. Front Med (Lausanne) 4:154. https://doi.org/10.3389/fmed.2017.00154
Korfei M, Schmitt S, Ruppert C, Henneke I, Markart P, Loeh B et al (2011) Comparative proteomic analysis of lung tissue from patients with idiopathic pulmonary fibrosis (IPF) and lung transplant donor lungs. J Proteome Res 10:2185–2205. https://doi.org/10.1021/pr1009355
Foster MW, Morrison LD, Todd JL, Snyder LD, Thompson JW, Soderblom EJ et al (2015) Quantitative proteomics of bronchoalveolar lavage fluid in idiopathic pulmonary fibrosis. J Proteome Res 14:1238–1249. https://doi.org/10.1021/pr501149m
O’Dwyer DN, Norman KC, Xia M, Huang Y, Gurczynski SJ, Ashley SL et al (2017) The peripheral blood proteome signature of idiopathic pulmonary fibrosis is distinct from normal and is associated with novel immunological processes. Sci Rep 7:46560. https://doi.org/10.1038/srep46560
Niu R, Liu Y, Zhang Y, Zhang Y, Wang H, Wang Y et al (2017) iTRAQ-based proteomics reveals novel biomarkers for idiopathic pulmonary fibrosis. PLoS ONE 12:e0170741. https://doi.org/10.1371/journal.pone.0170741
Tian Y, Li H, Gao Y, Liu C, Qiu T, Wu H et al (2019) Quantitative proteomic characterization of lung tissue in idiopathic pulmonary fibrosis. Clin Proteomics 16:6. https://doi.org/10.1186/s12014-019-9226-4
Landi C, Bergantini L, Cameli P, d’Alessandro M, Carleo A, Shaba E et al (2020) Idiopathic pulmonary fibrosis serum proteomic analysis before and after nintedanib therapy. Sci Rep 10:9378. https://doi.org/10.1038/s41598-020-66296-z
Luzina IG, Salcedo MV, Rojas-Pena ML, Wyman AE, Galvin JR, Sachdeva A et al (2018) Transcriptomic evidence of immune activation in macroscopically normal-appearing and scarred lung tissues in idiopathic pulmonary fibrosis. Cell Immunol 325:1–13. https://doi.org/10.1016/j.cellimm.2018.01.002
Sheu CC, Chang WA, Tsai MJ, Liao SH, Chong IW, Kuo PL (2019) Gene expression changes associated with nintedanib treatment in idiopathic pulmonary fibrosis fibroblasts: a next-generation sequencing and bioinformatics study. J Clin Med 8:308. https://doi.org/10.3390/jcm8030308
Sheu CC, Chang WA, Tsai MJ, Liao SH, Chong IW, Kuo PL (2019) Bioinformatic analysis of next-generation sequencing data to identify dysregulated genes in fibroblasts of idiopathic pulmonary fibrosis. Int J Mol Med 43:1643–1656. https://doi.org/10.3892/ijmm.2019.4086
Konigsberg IR, Borie R, Walts AD, Cardwell J, Rojas M, Metzger F et al (2021) Molecular signatures of idiopathic pulmonary fibrosis. Am J Respir Cell Mol Biol. https://doi.org/10.1165/rcmb.2020-0546OC
Raghu G, Remy-Jardin M, Myers JL, Richeldi L, Ryerson CJ, Lederer DJ et al (2018) Diagnosis of idiopathic pulmonary fibrosis. An official ATS/ERS/JRS/ALAT clinical practice guideline. Am J Respir Crit Care Med 198:e44–e68. https://doi.org/10.1164/rccm.201807-1255ST
Group of Interstitial Lung Diseases RDB, Association CM (2016) Consensus of Chinese experts on diagnosis and treatment of idiopathic pulmonary fibrosis. Chin J Tuberc Respir Dis 39:427–432. https://doi.org/10.3760/cma.j.issn.1001-0939.2016.06.005
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Andrews S (2010) FASTQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc. Accessed 26 Apr 2010
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
Anders S, Pyl PT, Huber W (2014) HTSeq—a python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169. https://doi.org/10.1093/bioinformatics/btu638
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong S et al (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:W316-322. https://doi.org/10.1093/nar/gkr483
Wheeler DL, Church DM, Federhen S, Lash AE, Madden TL, Pontius JU et al (2003) Database resources of the National Center for Biotechnology. Nucleic Acids Res 31:28–33. https://doi.org/10.1093/nar/gkg033
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/s0022-2836(05)80360-2
Tafer H, Hofacker IL (2008) RNAplex: a fast tool for RNA-RNA interaction search. Bioinformatics 24:2657–2663. https://doi.org/10.1093/bioinformatics/btn193
Wickham H (2016) ggplot2. Springer International Publishing, Cham
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O et al (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:1523. https://doi.org/10.1038/s41467-019-09234-6
Rohart F, Gautier B, Singh A, Le Cao KA (2017) mixOmics: an R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol 13:e1005752. https://doi.org/10.1371/journal.pcbi.1005752
Singh A, Shannon CP, Gautier B, Rohart F, Vacher M, Tebbutt SJ et al (2019) DIABLO: an integrative approach for identifying key molecular drivers from multi-omics assays. Bioinformatics 35:3055–3062. https://doi.org/10.1093/bioinformatics/bty1054
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M et al (2013) NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res 41:D991-995. https://doi.org/10.1093/nar/gks1193
Nance T, Smith KS, Anaya V, Richardson R, Ho L, Pala M et al (2014) Transcriptome analysis reveals differential splicing events in IPF lung tissue. PLoS ONE 9:e92111. https://doi.org/10.1371/journal.pone.0092111
Vukmirovic M, Herazo-Maya JD, Blackmon J, Skodric-Trifunovic V, Jovanovic D, Pavlovic S et al (2017) Identification and validation of differentially expressed transcripts by RNA-sequencing of formalin-fixed, paraffin-embedded (FFPE) lung tissue from patients with idiopathic pulmonary fibrosis. BMC Pulm Med 17:15. https://doi.org/10.1186/s12890-016-0356-4
Schafer MJ, White TA, Iijima K, Haak AJ, Ligresti G, Atkinson EJ et al (2017) Cellular senescence mediates fibrotic pulmonary disease. Nat Commun 8:14532. https://doi.org/10.1038/ncomms14532
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21. https://doi.org/10.1093/bioinformatics/bts635
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930. https://doi.org/10.1093/bioinformatics/btt656
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/gr.1239303
Kucera M, Isserlin R, Arkhangorodsky A, Bader GD (2016) AutoAnnotate: a cytoscape app for summarizing networks with semantic annotations. F1000Res 5:1717. https://doi.org/10.12688/f1000research.9090.1
Richeldi L, Collard HR, Jones MG (2017) Idiopathic pulmonary fibrosis. Lancet 389:1941–1952. https://doi.org/10.1016/s0140-6736(17)30866-8
Kulkarni T, de Andrade J, Zhou Y, Luckhardt T, Thannickal VJ (2016) Alveolar epithelial disintegrity in pulmonary fibrosis. Am J Physiol Lung Cell Mol Physiol 311:L185-191. https://doi.org/10.1152/ajplung.00115.2016
Rout-Pitt N, Farrow N, Parsons D, Donnelley M (2018) Epithelial mesenchymal transition (EMT): a universal process in lung diseases with implications for cystic fibrosis pathophysiology. Respir Res 19:136. https://doi.org/10.1186/s12931-018-0834-8
Demetriades C, Doumpas N, Teleman Aurelio A (2014) Regulation of TORC1 in response to amino acid starvation via lysosomal recruitment of TSC2. Cell 156:786–799. https://doi.org/10.1016/j.cell.2014.01.024
Plate M, Guillotin D, Chambers RC (2020) The promise of mTOR as a therapeutic target pathway in idiopathic pulmonary fibrosis. Eur Respir Rev 29:200269. https://doi.org/10.1183/16000617.0269-2020
Romero Y, Bueno M, Ramirez R, Alvarez D, Sembrat JC, Goncharova EA et al (2016) mTORC1 activation decreases autophagy in aging and idiopathic pulmonary fibrosis and contributes to apoptosis resistance in IPF fibroblasts. Aging Cell 15:1103–1112. https://doi.org/10.1111/acel.12514
Hosseinzadeh A, Javad-Moosavi SA, Reiter RJ, Hemati K, Ghaznavi H, Mehrzadi S (2018) Idiopathic pulmonary fibrosis (IPF) signaling pathways and protective roles of melatonin. Life Sci 201:17–29. https://doi.org/10.1016/j.lfs.2018.03.032
Nataraj D, Ernst A, Kalluri R (2010) Idiopathic pulmonary fibrosis is associated with endothelial to mesenchymal transition. Am J Respir Cell Mol Biol 43:129–130. https://doi.org/10.1165/rcmb.2010-0044ED
Hunninghake GW, Gadek JE, Lawley TJ, Crystal RG (1981) Mechanisms of neutrophil accumulation in the lungs of patients with idiopathic pulmonary fibrosis. J Clin Invest 68:259–269. https://doi.org/10.1172/jci110242
Bucala R, Spiegel LA, Chesney J, Hogan M, Cerami A (1994) Circulating fibrocytes define a new leukocyte subpopulation that mediates tissue repair. Mol Med 1:71–81
Beers MF, Morrisey EE (2011) The three R’s of lung health and disease: repair, remodeling, and regeneration. J Clin Invest 121:2065–2073. https://doi.org/10.1172/JCI45961
Chambers RC, Mercer PF (2015) Mechanisms of alveolar epithelial injury, repair, and fibrosis. Ann Am Thorac Soc 12(Suppl 1):S16-20. https://doi.org/10.1513/AnnalsATS.201410-448MG
Li Y, Bao J, Bian Y, Erben U, Wang P, Song K et al (2018) S100A4(+) macrophages are necessary for pulmonary fibrosis by activating lung fibroblasts. Front Immunol 9:1776. https://doi.org/10.3389/fimmu.2018.01776
Cui Y, Ji J, Hou J, Tan Y, Han X (2021) Identification of key candidate genes involved in the progression of idiopathic pulmonary fibrosis. Molecules. https://doi.org/10.3390/molecules26041123
Chen L, Tang R-Z, Ruan J, Zhu X-B, Yang Y (2019) Up-regulation of THY1 attenuates interstitial pulmonary fibrosis and promotes lung fibroblast apoptosis during acute interstitial pneumonia by blockade of the WNT signaling pathway. Cell Cycle 18:670–681
Chilosi M, Calio A, Rossi A, Gilioli E, Pedica F, Montagna L et al (2017) Epithelial to mesenchymal transition-related proteins ZEB1, beta-catenin, and beta-tubulin-III in idiopathic pulmonary fibrosis. Mod Pathol 30:26–38. https://doi.org/10.1038/modpathol.2016.147
DePianto DJ, Chandriani S, Abbas AR, Jia G, N’Diaye EN, Caplazi P et al (2015) Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis. Thorax 70:48–56. https://doi.org/10.1136/thoraxjnl-2013-204596
Bauer Y, Tedrow J, de Bernard S, Birker-Robaczewska M, Gibson KF, Guardela BJ et al (2015) A novel genomic signature with translational significance for human idiopathic pulmonary fibrosis. Am J Respir Cell Mol Biol 52:217–231. https://doi.org/10.1165/rcmb.2013-0310OC
Wang F, Li P, Li FS (2020) Integrated analysis of a gene correlation network identifies critical regulation of fibrosis by lncRNAs and TFs in idiopathic pulmonary fibrosis. Biomed Res Int 2020:6537462. https://doi.org/10.1155/2020/6537462
Hsu Y-L, Hung J-Y, Lee Y-L, Chen F-W, Chang K-F, Chang W-A et al (2017) Identification of novel gene expression signature in lung adenocarcinoma by using next-generation sequencing data and bioinformatics analysis. Oncotarget. https://doi.org/10.18632/oncotarget.21022
Ma W, Liang J, Zhang S, Ningdong H, Chen Z, Tian D (2020) Butyrophilin-like 9 regulates immune infiltration and serves as a prognostic marker in lung adenocarcinoma. BMC Cancer. https://doi.org/10.21203/rs.3.rs-113263/v1
Wang Y, Zhao N, Zhang X, Li Z, Liang Z, Yang J et al (2020) Bibliometrics analysis of butyrophilins as immune regulators [1992–2019] and implications for cancer prognosis. Front Immunol. https://doi.org/10.3389/fimmu.2020.01187
Bao Y, Wang L, Shi L, Yun F, Liu X, Chen Y et al (2019) Transcriptome profiling revealed multiple genes and ECM-receptor interaction pathways that may be associated with breast cancer. Cell Mol Biol Lett 24:38. https://doi.org/10.1186/s11658-019-0162-0
Le Guelte A, Macara IG (2015) Plasmolipin—a new player in endocytosis and epithelial development. EMBO J 34:1147–1148. https://doi.org/10.15252/embj.201591448
You J, Corley SM, Wen L, Hodge C, Hollhumer R, Madigan MC et al (2018) RNA-Seq analysis and comparison of corneal epithelium in keratoconus and myopia patients. Sci Rep 8:389. https://doi.org/10.1038/s41598-017-18480-x
Caswell DR, Chuang CH, Ma RK, Winters IP, Snyder EL, Winslow MM (2018) Tumor suppressor activity of Selenbp1, a direct Nkx2-1 target, in lung adenocarcinoma. Mol Cancer Res 16:1737–1749. https://doi.org/10.1158/1541-7786.Mcr-18-0392
Thul PJ, Akesson L, Wiking M, Mahdessian D, Geladaki A, Ait Blal H et al (2017) A subcellular map of the human proteome. Science 356:eaal3321. https://doi.org/10.1126/science.aal3321
Richeldi L, du Bois RM, Raghu G, Azuma A, Brown KK, Costabel U et al (2014) Efficacy and safety of nintedanib in idiopathic pulmonary fibrosis. N Engl J Med 370:2071–2082. https://doi.org/10.1056/NEJMoa1402584
King TE Jr, Bradford WZ, Castro-Bernardini S, Fagan EA, Glaspole I, Glassberg MK et al (2014) A phase 3 trial of pirfenidone in patients with idiopathic pulmonary fibrosis. N Engl J Med 370:2083–2092. https://doi.org/10.1056/NEJMoa1402582
Tzouvelekis A, Kaminski N (2015) Epigenetics in idiopathic pulmonary fibrosis. Biochem Cell Biol 93:159–170. https://doi.org/10.1139/bcb-2014-0126
Huang C, Yang Y, Liu L (2015) Interaction of long noncoding RNAs and microRNAs in the pathogenesis of idiopathic pulmonary fibrosis. Physiol Genom 47:463–469. https://doi.org/10.1152/physiolgenomics.00064.2015
Molyneaux PL, Willis-Owen SAG, Cox MJ, James P, Cowman S, Loebinger M et al (2017) Host-microbial interactions in idiopathic pulmonary fibrosis. Am J Respir Crit Care Med 195:1640–1650. https://doi.org/10.1164/rccm.201607-1408OC
Han MK, Zhou Y, Murray S, Tayob N, Noth I, Lama VN et al (2014) Lung microbiome and disease progression in idiopathic pulmonary fibrosis: an analysis of the COMET study. Lancet Respir Med 2:548–556. https://doi.org/10.1016/S2213-2600(14)70069-4
Pandit KV, Milosevic J (2015) MicroRNA regulatory networks in idiopathic pulmonary fibrosis. Biochem Cell Biol 93:129–137. https://doi.org/10.1139/bcb-2014-0101
Mizuno K, Mataki H, Seki N, Kumamoto T, Kamikawaji K, Inoue H (2017) MicroRNAs in non-small cell lung cancer and idiopathic pulmonary fibrosis. J Hum Genet 62:57–65. https://doi.org/10.1038/jhg.2016.98
Yang IV, Schwartz DA (2015) Epigenetics of idiopathic pulmonary fibrosis. Transl Res 165:48–60. https://doi.org/10.1016/j.trsl.2014.03.011
Stelzer G, Rosen N, Plaschkes I, Zimmerman S, Twik M, Fishilevich S et al (2016) The genecards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinform. https://doi.org/10.1002/cpbi.5
Li D, Liu Y, Wang B (2020) Identification of transcriptomic markers for developing idiopathic pulmonary fibrosis: an integrative analysis of gene expression profiles. Int J Clin Exp Pathol 13:1698–1706. https://www.ncbi.nlm.nih.gov/pubmed/32782692. Accessed 15 July 2020
Wang Z, Zhu J, Chen F, Ma L (2019) Weighted gene coexpression network analysis identifies key genes and pathways associated with idiopathic pulmonary fibrosis. Med Sci Monit 25:4285–4304. https://doi.org/10.12659/MSM.916828
Furusawa H, Cardwell JH, Okamoto T, Walts AD, Konigsberg IR, Kurche JS et al (2020) Chronic hypersensitivity pneumonitis, an interstitial lung disease with distinct molecular signatures. Am J Respir Crit Care Med 202:1430–1444. https://doi.org/10.1164/rccm.202001-0134OC
Fischer I, Durrie R, Sapirstein VS (1994) Plasmolipin: the other myelin proteolipid. A review of studies on its structure, expression, and function. Neurochem Res 19:959–966. https://doi.org/10.1007/bf00968705
Rodríguez-Fraticelli AE, Bagwell J, Bosch-Fortea M, Boncompain G, Reglero-Real N, García-León MJ et al (2015) Developmental regulation of apical endocytosis controls epithelial patterning in vertebrate tubular organs. Nat Cell Biol 17:241–250. https://doi.org/10.1038/ncb3106
Zhang Y-H, Hoopmann MR, Castaldi PJ, Simonsen KA, Midha MK, Cho MH et al (2021) Lung proteomic biomarkers associated with chronic obstructive pulmonary disease. medRxiv. https://doi.org/10.1101/2021.04.07.21255030
Mohning MP, Swigris JJ, Olson AL (2019) Idiopathic pulmonary fibrosis: the epidemiology and natural history of disease. In: Meyer KC, Nathan SD (eds) Idiopathic pulmonary fibrosis. Springer International Publishing, Cham, pp 11–35 (Respiratory Medicine)
Funding
This work was supported by the National Natural Science Foundation of China (Project No. 81871736), Bureau of traditional Chinese Medicine Scientific Research Project of Guangdong (Project No. 20192048), The First Affiliated Hospital Of Guangzhou Medical University (ZH201915) (funds from GMU), Guangzhou Institute of Respiratory Health Open Project (Funds provided by China Evergrande Group, Project No. 2020GIRHHMS04), the Zhongnanshan Medical Foundation of Guangdong Province (Project No: ZNSA-2021005 and Project No: ZNSA-2020001), the University of Macau (grant numbers: FHS-CRDA-029-002-2017, and MYRG2018-00071-FHS), the Science and Technology Development Fund, Macau SAR (File No. 0004/2019/AFJ and 0011/2019/AKP), and Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (Project No. 2018PT31048). Funding for open access charge: National Natural Science Foundation of China.
Author information
Authors and Affiliations
Contributions
All authors participated in the study design, the interpretation of the results, and the drafting and revision of the manuscript. PZ and SS conceived the idea and conducted the experiments, JW collected the specimens and conducted the experiments, SS analyzed the data and drafted the first version of the manuscript, XDZ and BS supervised the research. All authors reviewed and commented on the manuscript and approved the final draft. The authors would like to express their sincere thanks to Prof. Yingying Gu and Dr. Zhucheng Chen from The First Affiliated Hospital of Guangzhou Medical University for her support in determining the HRCT and histological patterns of the patients and confirming the IPF diagnosis. Besides, we would like to thank all members of Professor Sun’s lab and Professor Zhang’s lab for their kind help in conducting the experiments and preparing this manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
All procedures in this study involving human participants and animals were performed with the approval and according to the guidelines of the ethics committee of The First Affiliated Hospital of Guangzhou Medical University. All procedures were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2000. Informed consent was obtained from all individuals for being included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
18_2021_4094_MOESM1_ESM.pdf
Supplementary file1 Quality control results of RNA sequencing. A, B, and C are QC plots for raw sequences, and D, E, and F are QC plots for filtered sequences. A and D: Percentage of base content along position in reads. The fluctuations around 150th of the read position are caused by the read length limitation. B and E: error rates of bases along the position in the reads. C and F: the quality score of bases along the position in the reads. The quality score was calculated as -10×log10 (error P) (PDF 468 KB)
18_2021_4094_MOESM2_ESM.pdf
Supplementary file2 Distribution of mapped reads. A. Saturation map of all samples. It displays the increase of mapped genes along with the percentage of mapped reads. “Con” stands for control samples, and “Exp” stands for IPF samples. B. Mapped reads distribution. C. Reads density of a sample in the chromosomes. Red stands for forward strands, blue stands for reverse strands (PDF 842 KB)
18_2021_4094_MOESM3_ESM.pdf
Supplementary file3 Quality control of proteome. A. Density distribution of precursors. CV: coefficient of variation. “C” stands for control group and “D” stands for IPF group. B. Cumulative recovery plot for identified proteins. C. Completeness plot of the identified proteins. D. Identified proteins in each sample. C.1–C.5 represent the five control samples, D.1–D.6 represent the six IPF samples. E. Heatmap of the intensity of all quantified proteins (PDF 560 KB)
18_2021_4094_MOESM4_ESM.png
Supplementary file4 IHC results in human lung tissues. This figure displays the IHC staining of BTNL9 and PLLP in healthy and IPF lung tissues. The hematoxylin-stained nucleus is blue, and the DAB-positive area is brownish yellow. “×100” represents images captured through a 100 times lens, while “×400” represents images captured through a 400 times lens. BTNL9 stained nuclei were observed in AEC I cells of healthy lung tissue, while not observed in those of IPF lung tissues. PLLP staining was observed in AEC I cell membrane of healthy lung tissue, while not observed in those of IPF lung tissues (PNG 7897 KB)
18_2021_4094_MOESM5_ESM.png
Supplementary file5 IHC results in mouse lung tissues. This figure displays the IHC staining of BTNL9 and PLLP in lung tissues from BLM-induced mice and healthy ones. The hematoxylin-stained nucleus is blue, and the DAB-positive area is brownish yellow. “×100” represents images captured through a 100 times lens, while “×400” represents images captured through a 400 times lens. The comparison displays the destruction of the lung tissues in BLM-induced mice; both BTNL9 and PLLP showed decreased expression in the cytosol of lung bronchioles epithelial cells, and they increased expressed in alveoli cells (PNG 8832 KB)
Rights and permissions
About this article
Cite this article
Zheng, P., Sun, S., Wang, J. et al. Integrative omics analysis identifies biomarkers of idiopathic pulmonary fibrosis. Cell. Mol. Life Sci. 79, 66 (2022). https://doi.org/10.1007/s00018-021-04094-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-021-04094-0