Abstract
MicroRNAs (miRs) are non-coding gene transcripts abundantly expressed in both the developing and adult mammalian brain. They act as important modulators of complex gene regulatory networks during neuronal development and plasticity. miR-181c is highly abundant in cerebellar cortex and its expression is increased in autism patients as well as in an animal model of autism. To systematically identify putative targets of miR-181c, we repressed this miR in growing cortical neurons and found over 70 differentially expressed target genes using transcriptome profiling. Pathway analysis showed that the miR-181c-modulated genes converge on signaling cascades relevant to neurite and synapse developmental processes. To experimentally examine the significance of these data, we inhibited miR-181c during rat cortical neuronal maturation in vitro; this loss-of miR-181c function resulted in enhanced neurite sprouting and reduced synaptogenesis. Collectively, our findings suggest that miR-181c is a modulator of gene networks associated with cortical neuronal maturation.






Similar content being viewed by others
References
Buckley SM, Aranda-Orgilles B, Strikoudis A, Apostolou E, Loizou E et al (2012) Regulation of pluripotency and cellular reprogramming by the ubiquitin–proteasome system. Cell Stem Cell 11:783–798
Smart F, Aschrafi A, Atkins A, Owens GC, Pilotte J et al (2007) Two isoforms of the cold-inducible mRNA-binding protein RBM3 localize to dendrites and promote translation. J Neurochem 101:1367–1379
Klann E, Dever TE (2004) Biochemical mechanisms for translational regulation in synaptic plasticity. Nat Rev Neurosci 5:931–942
Huang F, Chotiner JK, Steward O (2005) The mRNA for elongation factor 1alpha is localized in dendrites and translated in response to treatments that induce long-term depression. J Neurosci 25:7199–7209
Aschrafi A, Kar AN, Natera-Naranjo O, Macgibeny MA, Gioio AE et al (2012) MicroRNA-338 regulates the axonal expression of multiple nuclear-encoded mitochondrial mRNAs encoding subunits of the oxidative phosphorylation machinery. Cell Mol Life Sci 69:4017–4027
van Rooij E, Sutherland LB, Qi X, Richardson JA, Hill J et al (2007) Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 316:575–579
Barca-Mayo O, De Pietri Tonelli D (2014) Convergent microRNA actions coordinate neocortical development. Cell Mol Life Sci 71:2975–2995
Bicker S, Lackinger M, Weiss K, Schratt G (2014) MicroRNA-132, -134, and -138: a microRNA troika rules in neuronal dendrites. Cell Mol Life Sci 71:3987–4005
Olde Loohuis NF, Kos A, Martens GJ, Van Bokhoven H, Nadif Kasri N et al (2012) MicroRNA networks direct neuronal development and plasticity. Cell Mol Life Sci 69:89–102
Iyer AN, Bellon A, Baudet ML (2014) microRNAs in axon guidance. Front Cell Neurosci 8:78
McNeill E, Van Vactor D (2012) MicroRNAs shape the neuronal landscape. Neuron 75:363–379
Ghahramani Seno MM, Hu P, Gwadry FG, Pinto D, Marshall CR et al (2011) Gene and miRNA expression profiles in autism spectrum disorders. Brain Res 1380:85–97
Olde Loohuis NF, Kole K, Glennon JC, Karel P, Van der Borg G et al (2015) Elevated microRNA-181c and microRNA-30d levels in the enlarged amygdala of the valproic acid rat model of autism. Neurobiol Dis 80:42–53
Smith B, Treadwell J, Zhang D, Ly D, McKinnell I et al (2010) Large-scale expression analysis reveals distinct microRNA profiles at different stages of human neurodevelopment. PLoS One 5:e11109
He M, Liu Y, Wang X, Zhang MQ, Hannon GJ et al (2012) Cell-type-based analysis of microRNA profiles in the mouse brain. Neuron 73:35–48
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM et al (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773
Chow ML, Pramparo T, Winn ME, Barnes CC, Li HR et al (2012) Age-dependent brain gene expression and copy number anomalies in autism suggest distinct pathological processes at young versus mature ages. PLoS Genet 8:e1002592
Garbett K, Ebert PJ, Mitchell A, Lintas C, Manzi B et al (2008) Immune transcriptome alterations in the temporal cortex of subjects with autism. Neurobiol Dis 30:303–311
Gupta S, Ellis SE, Ashar FN, Moes A, Bader JS et al (2014) Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism. Nat Commun 5:5748
Sasaki Y, Gross C, Xing L, Goshima Y, Bassell GJ (2014) Identification of axon-enriched microRNAs localized to growth cones of cortical neurons. Dev Neurobiol 74:397–406
Hancock ML, Preitner N, Quan J, Flanagan JG (2014) MicroRNA-132 is enriched in developing axons, locally regulates Rasa1 mRNA, and promotes axon extension. J Neurosci 34:66–78
Natera-Naranjo O, Aschrafi A, Gioio AE, Kaplan BB (2010) Identification and quantitative analyses of microRNAs located in the distal axons of sympathetic neurons. RNA 16:1516–1529
Dajas-Bailador F, Bonev B, Garcez P, Stanley P, Guillemot F et al (2012) microRNA-9 regulates axon extension and branching by targeting Map1b in mouse cortical neurons. Nat Neurosci. doi:10.1038/nn.3082
Aschrafi A, Schwechter AD, Mameza MG, Natera-Naranjo O, Gioio AE et al (2008) MicroRNA-338 regulates local cytochrome c oxidase IV mRNA levels and oxidative phosphorylation in the axons of sympathetic neurons. J Neurosci 28:12581–12590
Park JW, Vahidi B, Taylor AM, Rhee SW, Jeon NL (2006) Microfluidic culture platform for neuroscience research. Nat Protoc 1:2128–2136
Marchetto MC, Carromeu C, Acab A, Yu D, Yeo GW et al (2010) A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell 143:527–539
Brennand KJ, Simone A, Jou J, Gelboin-Burkhart C, Tran N et al (2011) Modelling schizophrenia using human induced pluripotent stem cells. Nature 473:221–225
Gallagher D, Voronova A, Zander MA, Cancino GI, Bramall A et al (2015) Ankrd11 is a chromatin regulator involved in autism that is essential for neural development. Dev Cell 32:31–42
Belmonte MK, Allen G, Beckel-Mitchener A, Boulanger LM, Carper RA et al (2004) Autism and abnormal development of brain connectivity. J Neurosci 24:9228–9231
Belmonte MK, Bourgeron T (2006) Fragile X syndrome and autism at the intersection of genetic and neural networks. Nat Neurosci 9:1221–1225
Ouyang YB, Lu Y, Yue S, Giffard RG (2012) miR-181 targets multiple Bcl-2 family members and influences apoptosis and mitochondrial function in astrocytes. Mitochondrion 12:213–219
Ouyang YB, Lu Y, Yue S, Xu LJ, Xiong XX et al (2012) miR-181 regulates GRP78 and influences outcome from cerebral ischemia in vitro and in vivo. Neurobiol Dis 45:555–563
Helwak A, Kudla G, Dudnakova T, Tollervey D (2013) Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 153:654–665
Wang WX, Wilfred BR, Xie K, Jennings MH, Hu YH et al (2010) Individual microRNAs (miRNAs) display distinct mRNA targeting “rules”. RNA Biol 7:373–380
Head BP, Peart JN, Panneerselvam M, Yokoyama T, Pearn ML et al (2010) Loss of caveolin-1 accelerates neurodegeneration and aging. PLoS One 5:e15697
Head BP, Hu Y, Finley JC, Saldana MD, Bonds JA et al (2011) Neuron-targeted caveolin-1 protein enhances signaling and promotes arborization of primary neurons. J Biol Chem 286:33310–33321
Malik AR, Urbanska M, Gozdz A, Swiech LJ, Nagalski A et al (2013) Cyr61, a matricellular protein, is needed for dendritic arborization of hippocampal neurons. J Biol Chem 288:8544–8559
Lu Y, Tian QB, Endo S, Suzuki T (2007) A role for LRP4 in neuronal cell viability is related to apoE-binding. Brain Res 1177:19–28
Yumoto N, Kim N, Burden SJ (2012) Lrp4 is a retrograde signal for presynaptic differentiation at neuromuscular synapses. Nature 489:438–442
Mei Y, Wang Z, Zhang L, Zhang Y, Li X et al (2012) Regulation of neuroblastoma differentiation by forkhead transcription factors FOXO1/3/4 through the receptor tyrosine kinase PDGFRA. Proc Natl Acad Sci USA 109:4898–4903
Feliciano P, Andrade R, Bykhovskaia M (2013) Synapsin II and Rab3a cooperate in the regulation of epileptic and synaptic activity in the CA1 region of the hippocampus. J Neurosci 33:18319–18330
Takao-Rikitsu E, Mochida S, Inoue E, Deguchi-Tawarada M, Inoue M et al (2004) Physical and functional interaction of the active zone proteins, CAST, RIM1, and Bassoon, in neurotransmitter release. J Cell Biol 164:301–311
Yuen EY, Wei J, Liu W, Zhong P, Li X et al (2012) Repeated stress causes cognitive impairment by suppressing glutamate receptor expression and function in prefrontal cortex. Neuron 73:962–977
Saba R, Storchel PH, Aksoy-Aksel A, Kepura F, Lippi G et al (2012) Dopamine-regulated microRNA MiR-181a controls GluA2 surface expression in hippocampal neurons. Mol Cell Biol 32:619–632
Beveridge NJ, Tooney PA, Carroll AP, Gardiner E, Bowden N et al (2008) Dysregulation of miRNA 181b in the temporal cortex in schizophrenia. Hum Mol Genet 17:1156–1168
Beveridge NJ, Gardiner E, Carroll AP, Tooney PA, Cairns MJ (2010) Schizophrenia is associated with an increase in cortical microRNA biogenesis. Mol Psychiatry 15:1176–1189
Kamanu TK, Radovanovic A, Archer JA, Bajic VB (2013) Exploration of miRNA families for hypotheses generation. Sci Rep 3:2940
Sun X, Sit A, Feinberg MW (2014) Role of miR-181 family in regulating vascular inflammation and immunity. Trends Cardiovasc Med 24:105–112
Issler O, Haramati S, Paul ED, Maeno H, Navon I et al (2014) MicroRNA 135 is essential for chronic stress resiliency, antidepressant efficacy, and intact serotonergic activity. Neuron 83:344–360
Abbott AL, Alvarez-Saavedra E, Miska EA, Lau NC, Bartel DP et al (2005) The let-7 MicroRNA family members mir-48, mir-84, and mir-241 function together to regulate developmental timing in Caenorhabditis elegans. Dev Cell 9:403–414
Xu B, Hsu PK, Stark KL, Karayiorgou M, Gogos JA (2013) Derepression of a neuronal inhibitor due to miRNA dysregulation in a schizophrenia-related microdeletion. Cell 152:262–275
Willemsen MH, Valles A, Kirkels LA, Mastebroek M, Olde Loohuis N et al (2011) Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J Med Genet 48:810–818
Perkins DO, Jeffries CD, Jarskog LF, Thomson JM, Woods K et al (2007) microRNA expression in the prefrontal cortex of individuals with schizophrenia and schizoaffective disorder. Genome Biol 8:R27
Xu B, Karayiorgou M, Gogos JA (2010) MicroRNAs in psychiatric and neurodevelopmental disorders. Brain Res 1338:78–88
Stark KL, Xu B, Bagchi A, Lai WS, Liu H et al (2008) Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat Genet 40:751–760
Mellios N, Sugihara H, Castro J, Banerjee A, Le C et al (2011) miR-132, an experience-dependent microRNA, is essential for visual cortex plasticity. Nat Neurosci 14:1240–1242
Lin Q, Wei W, Coelho CM, Li X, Baker-Andresen D et al (2011) The brain-specific microRNA miR-128b regulates the formation of fear-extinction memory. Nat Neurosci 14:1115–1117
Zhang F, Gradinaru V, Adamantidis AR, Durand R, Airan RD et al (2010) Optogenetic interrogation of neural circuits: technology for probing mammalian brain structures. Nat Protoc 5:439–456
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N et al (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:RESEARCH0034
Dweep H, Sticht C, Pandey P, Gretz N (2011) miRWalk—database: prediction of possible miRNA binding sites by “walking” the genes of three genomes. J Biomed Inform 44:839–847
Meijering E, Jacob M, Sarria JC, Steiner P, Hirling H et al (2004) Design and validation of a tool for neurite tracing and analysis in fluorescence microscopy images. Cytometry A 58:167–176
Rodriguez A, Ehlenberger DB, Dickstein DL, Hof PR, Wearne SL (2008) Automated three-dimensional detection and shape classification of dendritic spines from fluorescence microscopy images. PLoS One 3:e1997
Edbauer D, Neilson JR, Foster KA, Wang CF, Seeburg DP et al (2010) Regulation of synaptic structure and function by FMRP-associated microRNAs miR-125b and miR-132. Neuron 65:373–384
Ebert MS, Neilson JR, Sharp PA (2007) MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat Methods 4:721–726
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
18_2016_2179_MOESM1_ESM.jpg
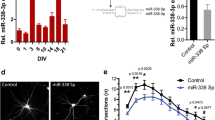
Supplement Figure 1: MiR-181c transduction efficiency into primary cortical neurons. A. Schematic overview of the lentiviral miR-181c-sponge construct designed to continuously and specifically sequester miR-181c. A concatemer of four complementary miR-181c binding sites is placed within the 3′UTR of the eGFP gene. As example, one binding site of the sponge with the mature miR-181c is shown, the central mismatch is included to enhance efficiency [56,63,64]. B. Micrographs of primary cortical neurons infected with the miR-181c sponge lentivirus. The left image shows cell nuclei visualized with DAPI (bleu), the middle panel shows GFP from the miR-181c sponge (green) and on the right is the DAPI and GFP overlay. C. Relative expression of miR-181c in DIV 9-old primary cortical neurons expressing the GFP control or miR-181c sponge construct normalized to control. D. Representative images of primary cortical neurons transfected with siGLO transfection indicator. The micrographs from left to right show the nuclear DAPI staining (blue), siGLO (red) and a DAPI with siGLO overlay. E. Relative expression of miR-181c in DIV 6 primary cortical neurons transfected with NT control or miR-181c mimic, levels were normalized to control. F. Relative expression of miR-181c in DIV 6 primary cortical neurons transfected with NT control or anti-miR-181c, levels were normalized to control. Data are shown as mean ± SEM; p values are determined by two-tailed unpaired Student’s t test. *p ≤ 0.01 and ** p ≤ 0.001 (JPEG 1349 kb)
18_2016_2179_MOESM2_ESM.jpg
Supplement Figure 2: Growing pure axonal fractions in microfluidic chambers. A. Schematic representation of a microfluidic chamber with the soma side (gray) and the axon side (green) connected with microgrooves in the middle. Neurons are shown in red, which grow their axons through the microgrooves to the axonal side. B. An example immunostaining of cortical neurons cultured in microfluidic devices. The DIC image shows the microgrooves of the microfluidic chamber, MAP2 highlights the dendrites in green and Tau1 for visualizing axons in red. The image on the bottom right corner shows an overlay of DIC, MAP2 and Tau1 (JPEG 403 kb)
18_2016_2179_MOESM3_ESM.jpg
Supplement Figure 3: Mature cortical neurons transduced with miR-181c mimic have a necrotic phenotype. Representative micrographs of DIV16 primary cortical neurons transfected with 30 nM NT control (upper panels) or miR-181c mimic (lower panels) and 20 nM SiGlo fluorescent transfection indicator at DIV 3. The panels depict micrographs of DIV16 neurons stained with DAPI (blue), PSD-95 (green) and the SiGlo transfection indicator (red). The necrotic phenotype (condensed nuclei) in miR-181c transfected neurons was observed for around 90% of the cells; this experiment was repeated three times, with similar outcomes (JPEG 853 kb)
18_2016_2179_MOESM4_ESM.pdf
Supplement Table 1: List of genes functioning in behavior, cell-to-cell signaling and interaction, nervous system development and function (PDF 100 kb)
Rights and permissions
About this article
Cite this article
Kos, A., Olde Loohuis, N., Meinhardt, J. et al. MicroRNA-181 promotes synaptogenesis and attenuates axonal outgrowth in cortical neurons. Cell. Mol. Life Sci. 73, 3555–3567 (2016). https://doi.org/10.1007/s00018-016-2179-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-016-2179-0