Abstract
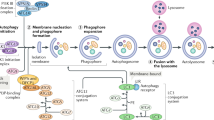
Autophagy is a conserved cellular recycling process that plays critical roles in development, disease, and ageing. During autophagy, cytosolic components are sequestered in double-membrane vesicles that ultimately fuse with lysosomes, where the cargo is degraded and recycled. Intriguingly, genetic and pharmacological experiments in C. elegans have shown that all of the longevity paradigms analysed to date, ranging from reduced insulin/IGF-1 signalling to spermidine supplementation, require autophagy genes for lifespan extension. Moreover, many of the long-lived animals show changes in steady-state levels of autophagy markers and/or display increased transcription of autophagy-related and lysosomal genes via conserved transcription factors such as HLH-30/TFEB. These observations are consistent with the notion that increased autophagy is critical for lifespan extension in C. elegans. Similar genetic links have been reported in other organisms, including flies and mice, where overexpression of certain autophagy-related genes is sufficient to extend lifespan. Although clearance of lipids (lipophagy) and mitochondria (mitophagy) have been proposed as selective types of autophagy with relevance to C. elegans ageing, it is still unclear how long-lived animals may induce autophagy to improve their overall healthspan, or how autophagy is regulated in different tissues during normal ageing. Understanding these mechanisms will be critical for targeting autophagy in higher organisms. This chapter summarizes our current knowledge of the links between autophagy and ageing in C. elegans.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Notes
- 1.
Nomenclature: Yeast genes/proteins are stated first, followed by the mammalian and C. elegans names, if different.
- 2.
Paradoxically, one report has suggested that adult-only RNAi inhibition of several autophagy genes can result in lifespan extension in C. elegans [32]; however, this study was performed on a very small number of animals in the presence of 5-fluoro-2′deoxyuridine, and results were not analysed by survival statistics.
References
Levine B, Kroemer G (2008) Autophagy in the pathogenesis of disease. Cell 132(1):27–42. doi:10.1016/j.cell.2007.12.018
Huang J, Klionsky DJ (2007) Autophagy and human disease. Cell Cycle 6(15):1837–1849
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24(1):24–41. doi:10.1038/cr.2013.168
Chan SN, Tang BL (2013) Location and membrane sources for autophagosome formation – from ER-mitochondria contact sites to Golgi-endosome-derived carriers. Mol Membr Biol 30(8):394–402. doi:10.3109/09687688.2013.850178
Hamasaki M, Shibutani ST, Yoshimori T (2013) Up-to-date membrane biogenesis in the autophagosome formation. Curr Opin Cell Biol 25(4):455–460. doi:10.1016/j.ceb.2013.03.004
Obara K, Ohsumi Y (2008) Dynamics and function of PtdIns(3)P in autophagy. Autophagy 4(7):952–954, doi:6790 [pii]
Simonsen A, Tooze SA (2009) Coordination of membrane events during autophagy by multiple class III PI3-kinase complexes. J Cell Biol 186(6):773–782. doi:jcb.200907014 [pii] 10.1083/jcb.200907014
Reggiori F, Tucker KA, Stromhaug PE, Klionsky DJ (2004) The Atg1-Atg13 complex regulates Atg9 and Atg23 retrieval transport from the pre-autophagosomal structure. Dev Cell 6(1):79–90, doi:S1534580703004027 [pii]
Mizushima N, Levine B (2010) Autophagy in mammalian development and differentiation. Nat Cell Biol 12(9):823–830. doi:10.1038/ncb0910-823
Manil-Segalen M, Lefebvre C, Jenzer C, Trichet M, Boulogne C, Satiat-Jeunemaitre B, Legouis R (2014) The C. elegans LC3 acts downstream of GABARAP to degrade autophagosomes by interacting with the HOPS subunit VPS39. Dev Cell 28(1):43–55. doi:10.1016/j.devcel.2013.11.022
Zhang H, Chang JT, Guo B, Hansen M, Jia K, Kovacs AL, Kumsta C, Lapierre LR, Legouis R, Lin L, Lu Q, Melendez A, O’Rourke EJ, Sato K, Sato M, Wang X, Wu F (2015) Guidelines for monitoring autophagy in C. elegans. Autophagy 11(1):9–27. doi:10.1080/15548627.2014.1003478
Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H et al (2016) Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12(1):1–222. doi:10.1080/15548627.2015.1100356
Johansen T, Lamark T (2011) Selective autophagy mediated by autophagic adapter proteins. Autophagy 7(3):279–296, doi:14487 [pii]
Khaminets A, Behl C, Dikic I (2016) Ubiquitin-dependent and independent signals in selective autophagy. Trends Cell Biol 26(1):6–16. doi:10.1016/j.tcb.2015.08.010
Tian Y, Li Z, Hu W, Ren H, Tian E, Zhao Y, Lu Q, Huang X, Yang P, Li X, Wang X, Kovacs AL, Yu L, Zhang H (2010) C. elegans screen identifies autophagy genes specific to multicellular organisms. Cell 141(6):1042–1055. doi:10.1016/j.cell.2010.04.034
Palikaras K, Lionaki E, Tavernarakis N (2015) Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans. Nature 521(7553):525–528. doi:10.1038/nature14300
Zhang Y, Yan L, Zhou Z, Yang P, Tian E, Zhang K, Zhao Y, Li Z, Song B, Han J, Miao L, Zhang H (2009) SEPA-1 mediates the specific recognition and degradation of P granule components by autophagy in C. elegans. Cell 136(2):308–321. doi:10.1016/j.cell.2008.12.022
Chen Y, Yu L (2012) Autophagic lysosome reformation. Exp Cell Res 319(2):142–146. doi:S0014-4827(12)00396-5 [pii] 10.1016/j.yexcr.2012.09.004
Russell RC, Yuan HX, Guan KL (2014) Autophagy regulation by nutrient signaling. Cell Res 24(1):42–57. doi:10.1038/cr.2013.166
Lapierre LR, Kumsta C, Sandri M, Ballabio A, Hansen M (2015) Transcriptional and epigenetic regulation of autophagy in aging. Autophagy 11(6):867–880. doi:10.1080/15548627.2015.1034410
Alers S, Loffler AS, Wesselborg S, Stork B (2011) Role of AMPK-mTOR-Ulk1/2 in the regulation of autophagy: cross talk, shortcuts, and feedbacks. Mol Cell Biol 32(1):2–11. doi:MCB.06159-11 [pii] 10.1128/MCB.06159-11
Kapahi P, Chen D, Rogers AN, Katewa SD, Li PW, Thomas EL, Kockel L (2010) With TOR, less is more: a key role for the conserved nutrient-sensing TOR pathway in aging. Cell Metab 11(6):453–465. doi:10.1016/j.cmet.2010.05.001
Apfeld J, O’Connor G, McDonagh T, DiStefano PS, Curtis R (2004) The AMP-activated protein kinase AAK-2 links energy levels and insulin-like signals to lifespan in C. elegans. Genes Dev 18(24):3004–3009
Mair W, Morantte I, Rodrigues AP, Manning G, Montminy M, Shaw RJ, Dillin A (2011) Lifespan extension induced by AMPK and calcineurin is mediated by CRTC-1 and CREB. Nature 470(7334):404–408. doi:10.1038/nature09706
Ulgherait M, Rana A, Rera M, Graniel J, Walker DW (2014) AMPK modulates tissue and organismal aging in a non-cell-autonomous manner. Cell Rep 8(6):1767–1780. doi:10.1016/j.celrep.2014.08.006
Stenesen D, Suh JM, Seo J, Yu K, Lee KS, Kim JS, Min KJ, Graff JM (2013) Adenosine nucleotide biosynthesis and AMPK regulate adult life span and mediate the longevity benefit of caloric restriction in flies. Cell Metab 17(1):101–112. doi:10.1016/j.cmet.2012.12.006
Gelino S, Hansen M (2012) Autophagy – an emerging anti-aging mechanism. J Clin Exp Pathol Suppl 4:pii: 006
Jia K, Levine B (2007) Autophagy is required for dietary restriction-mediated life span extension in C. elegans. Autophagy 3(6):597–599
Hansen M, Chandra A, Mitic LL, Onken B, Driscoll M, Kenyon C (2008) A role for autophagy in the extension of lifespan by dietary restriction in C. elegans. PLoS Genet 4(2):e24. doi:10.1371/journal.pgen.0040024
Tavernarakis N, Pasparaki A, Tasdemir E, Maiuri MC, Kroemer G (2008) The effects of p53 on whole organism longevity are mediated by autophagy. Autophagy 4(7):870–873
Lapierre LR, Gelino S, Melendez A, Hansen M (2011) Autophagy and lipid metabolism coordinately modulate life span in germline-less C. elegans. Curr Biol 21(18):1507–1514. doi:10.1016/j.cub.2011.07.042
Hashimoto Y, Ookuma S, Nishida E (2009) Lifespan extension by suppression of autophagy genes in C. elegans. Genes Cells 14(6):717–726. doi:10.1111/j.1365-2443.2009.01306.x
Kenyon CJ (2010) The genetics of ageing. Nature 464(7288):504–512. doi:10.1038/nature08980
Kenyon C, Chang J, Gensch E, Rudner A, Tabtiang R (1993) A C. elegans mutant that lives twice as long as wild type. Nature 366(6454):461–464
Melendez A, Talloczy Z, Seaman M, Eskelinen EL, Hall DH, Levine B (2003) Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 301(5638):1387–1391
Hars ES, Qi H, Ryazanov AG, Jin S, Cai L, Hu C, Liu LF (2007) Autophagy regulates ageing in C. elegans. Autophagy 3(2):93–95
Egan DF, Shackelford DB, Mihaylova MM, Gelino S, Kohnz RA, Mair W, Vasquez DS, Joshi A, Gwinn DM, Taylor R, Asara JM, Fitzpatrick J, Dillin A, Viollet B, Kundu M, Hansen M, Shaw RJ (2011) Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331(6016):456–461. doi:10.1126/science.1196371
Lapierre LR, De Magalhaes Filho CD, McQuary PR, Chu CC, Visvikis O, Chang JT, Gelino S, Ong B, Davis AE, Irazoqui JE, Dillin A, Hansen M (2013) The TFEB orthologue HLH-30 regulates autophagy and modulates longevity in C. elegans. Nat Commun 4:2267. doi:10.1038/ncomms3267
Laplante M, Sabatini DM (2012) mTOR signaling in growth control and disease. Cell 149(2):274–293. doi:10.1016/j.cell.2012.03.017
Vellai T, Takacs-Vellai K, Zhang Y, Kovacs AL, Orosz L, Muller F (2003) Genetics: influence of TOR kinase on lifespan in C. elegans. Nature 426(6967):620. doi:10.1038/426620a
Depuydt G, Shanmugam N, Rasulova M, Dhondt I, Braeckman BP (2016) Increased protein stability and decreased protein turnover in the C. elegans Ins/IGF-1 daf-2 mutant. J Gerontol A Biol Sci Med Sci. doi:10.1093/gerona/glv221
Mair W, Dillin A (2008) Aging and survival: the genetics of life span extension by dietary restriction. Annu Rev Biochem 77:727–754. doi:10.1146/annurev.biochem.77.061206.171059
Greer EL, Brunet A (2009) Different dietary restriction regimens extend lifespan by both independent and overlapping genetic pathways in C. elegans. Aging Cell 8(2):113–127
Lakowski B, Hekimi S (1998) The genetics of caloric restriction in C. elegans. Proc Natl Acad Sci U S A 95(22):13091–13096
Gomez-Amaro RL, Valentine ER, Carretero M, LeBoeuf SE, Rangaraju S, Broaddus CD, Solis GM, Williamson JR, Petrascheck M (2015) Measuring food intake and nutrient absorption in C. elegans. Genetics. doi:10.1534/genetics.115.175851
Toth ML, Sigmond T, Borsos E, Barna J, Erdelyi P, Takacs-Vellai K, Orosz L, Kovacs AL, Csikos G, Sass M, Vellai T (2008) Longevity pathways converge on autophagy genes to regulate life span in C. elegans. Autophagy 4(3):330–338
Morck C, Pilon M (2007) Caloric restriction and autophagy in C. elegans. Autophagy 3(1):51–53
Morselli E, Maiuri MC, Markaki M, Megalou E, Pasparaki A, Palikaras K, Criollo A, Galluzzi L, Malik SA, Vitale I, Michaud M, Madeo F, Tavernarakis N, Kroemer G (2010) Caloric restriction and resveratrol promote longevity through the sirtuin-1-dependent induction of autophagy. Cell Death Dis 1:e10. doi:10.1038/cddis.2009.8
Heestand BN, Shen Y, Liu W, Magner DB, Storm N, Meharg C, Habermann B, Antebi A (2013) Dietary restriction induced longevity is mediated by nuclear receptor NHR-62 in C. elegans. PLoS Genet 9(7):e1003651. doi:10.1371/journal.pgen.1003651
Panowski SH, Wolff S, Aguilaniu H, Durieux J, Dillin A (2007) PHA-4/Foxa mediates diet-restriction-induced longevity of C. elegans. Nature 447(7144):550–555
Pandit A, Jain V, Kumar N, Mukhopadhyay A (2014) PHA-4/FOXA-regulated microRNA feed forward loops during C. elegans dietary restriction. Aging 6(10):835–855
O’Rourke EJ, Kuballa P, Xavier R, Ruvkun G (2013) Omega-6 polyunsaturated fatty acids extend life span through the activation of autophagy. Genes Dev 27(4):429–440. doi:10.1101/gad.205294.112
Hansen M, Taubert S, Crawford D, Libina N, Lee SJ, Kenyon C (2007) Lifespan extension by conditions that inhibit translation in C. elegans. Aging Cell 6(1):95–110
Robida-Stubbs S, Glover-Cutter K, Lamming DW, Mizunuma M, Narasimhan SD, Neumann-Haefelin E, Sabatini DM, Blackwell TK (2012) TOR signaling and rapamycin influence longevity by regulating SKN-1/Nrf and DAF-16/FoxO. Cell Metab 15(5):713–724. doi:10.1016/j.cmet.2012.04.007
McQuary PR, Liao CY, Chang JT, Kumsta C, She X, Davis A, Chu CC, Gelino S, Gomez-Amaro RL, Petrascheck M, Brill LM, Ladiges WC, Kennedy BK, Hansen M (2016) C. elegans S6K mutants require a creatine-kinase-like effector for lifespan extension. Cell Rep 14(9):2059–2067. doi:10.1016/j.celrep.2016.02.012
Hansen M, Flatt T, Aguilaniu H (2013) Reproduction, fat metabolism, and life span: what is the connection? Cell Metab 17(1):10–19. doi:10.1016/j.cmet.2012.12.003
Wang MC, O'Rourke EJ, Ruvkun G (2008) Fat metabolism links germline stem cells and longevity in C. elegans. Science 322(5903):957–960. doi:322/5903/957 [pii] 10.1126/science.1162011
Folick A, Oakley HD, Yu Y, Armstrong EH, Kumari M, Sanor L, Moore DD, Ortlund EA, Zechner R, Wang MC (2015) Aging. Lysosomal signaling molecules regulate longevity in C. elegans. Science 347(6217):83–86. doi:10.1126/science.1258857
Lapierre LR, Silvestrini MJ, Nunez L, Ames K, Wong S, Le TT, Hansen M, Melendez A (2013) Autophagy genes are required for normal lipid levels in C. elegans. Autophagy 9(3):278–286. doi:10.4161/auto.22930
O’Rourke EJ, Ruvkun G (2013) MXL-3 and HLH-30 transcriptionally link lipolysis and autophagy to nutrient availability. Nat Cell Biol 15(6):668–676. doi:10.1038/ncb2741
Seah NE, de Magalhaes Filho CD, Petrashen AP, Henderson HR, Laguer J, Gonzalez J, Dillin A, Hansen M, Lapierre LR (2016) Autophagy-mediated longevity is modulated by lipoprotein biogenesis. Autophagy 12(2):261–272. doi:10.1080/15548627.2015.1127464
Durieux J, Dillin A (2007) Mitochondria and aging: dilution is the solution. Cell Metab 6(6):427–429. doi:10.1016/j.cmet.2007.11.008
Ewbank JJ, Barnes TM, Lakowski B, Lussier M, Bussey H, Hekimi S (1997) Structural and functional conservation of the C. elegans timing gene clk-1. Science 275(5302):980–983
Feng J, Bussiere F, Hekimi S (2001) Mitochondrial electron transport is a key determinant of life span in C. elegans. Dev Cell 1(5):633–644
Yang W, Hekimi S (2010) Two modes of mitochondrial dysfunction lead independently to lifespan extension in C. elegans. Aging Cell 9(3):433–447. doi:10.1111/j.1474-9726.2010.00571.x
Schiavi A, Torgovnick A, Kell A, Megalou E, Castelein N, Guccini I, Marzocchella L, Gelino S, Hansen M, Malisan F, Condo I, Bei R, Rea SL, Braeckman BP, Tavernarakis N, Testi R, Ventura N (2013) Autophagy induction extends lifespan and reduces lipid content in response to frataxin silencing in C. elegans. Exp Gerontol 48(2):191–201. doi:10.1016/j.exger.2012.12.002
Schiavi A, Maglioni S, Palikaras K, Shaik A, Strappazzon F, Brinkmann V, Torgovnick A, Castelein N, De Henau S, Braeckman BP, Cecconi F, Tavernarakis N, Ventura N (2015) Iron-starvation-induced mitophagy mediates lifespan extension upon mitochondrial stress in C. elegans. Curr Biol 25(14):1810–1822. doi:10.1016/j.cub.2015.05.059
Arum O, Johnson TE (2007) Reduced expression of the C. elegans p53 ortholog cep-1 results in increased longevity. J Gerontol 62(9):951–959
Pietrocola F, Izzo V, Niso-Santano M, Vacchelli E, Galluzzi L, Maiuri MC, Kroemer G (2013) Regulation of autophagy by stress-responsive transcription factors. Semin Cancer Biol 23(5):310–322. doi:10.1016/j.semcancer.2013.05.008
Tissenbaum HA, Guarente L (2001) Increased dosage of a sir-2 gene extends lifespan in C. elegans. Nature 410(6825):227–230
Viswanathan M, Kim SK, Berdichevsky A, Guarente L (2005) A role for SIR-2.1 regulation of ER stress response genes in determining C. elegans life span. Dev Cell 9(5):605–615. doi:10.1016/j.devcel.2005.09.017
Rogina B, Helfand SL (2004) Sir2 mediates longevity in the fly through a pathway related to calorie restriction. Proc Natl Acad Sci U S A 101(45):15998–16003. doi:10.1073/pnas.0404184101
Burnett C, Valentini S, Cabreiro F, Goss M, Somogyvari M, Piper MD, Hoddinott M, Sutphin GL, Leko V, McElwee JJ, Vazquez-Manrique RP, Orfila AM, Ackerman D, Au C, Vinti G, Riesen M, Howard K, Neri C, Bedalov A, Kaeberlein M, Soti C, Partridge L, Gems D (2011) Absence of effects of Sir2 overexpression on lifespan in C. elegans and Drosophila. Nature 477(7365):482–485. doi:10.1038/nature10296
Dong MQ, Venable JD, Au N, Xu T, Park SK, Cociorva D, Johnson JR, Dillin A, Yates JR 3rd (2007) Quantitative mass spectrometry identifies insulin signaling targets in C. elegans. Science 317(5838):660–663
Dwivedi M, Song HO, Ahnn J (2009) Autophagy genes mediate the effect of calcineurin on life span in C. elegans. Autophagy 5(5):604–607
Medina DL, Di Paola S, Peluso I, Armani A, De Stefani D, Venditti R, Montefusco S, Scotto-Rosato A, Prezioso C, Forrester A, Settembre C, Wang W, Gao Q, Xu H, Sandri M, Rizzuto R, De Matteis MA, Ballabio A (2015) Lysosomal calcium signalling regulates autophagy through calcineurin and TFEB. Nat Cell Biol 17(3):288–299. doi:10.1038/ncb3114
Yang J, Chen D, He Y, Melendez A, Feng Z, Hong Q, Bai X, Li Q, Cai G, Wang J, Chen X (2013) MiR-34 modulates C. elegans lifespan via repressing the autophagy gene atg9. Age (Dordr) 35(1):11–22. doi:10.1007/s11357-011-9324-3
Mosbech MB, Kruse R, Harvald EB, Olsen AS, Gallego SF, Hannibal-Bach HK, Ejsing CS, Faergeman NJ (2013) Functional loss of two ceramide synthases elicits autophagy-dependent lifespan extension in C. elegans. PLoS ONE 8(7):e70087. doi:10.1371/journal.pone.0070087
Denzel MS, Storm NJ, Gutschmidt A, Baddi R, Hinze Y, Jarosch E, Sommer T, Hoppe T, Antebi A (2014) Hexosamine pathway metabolites enhance protein quality control and prolong life. Cell 156(6):1167–1178. doi:10.1016/j.cell.2014.01.061
Shintani T, Yamazaki F, Katoh T, Umekawa M, Matahira Y, Hori S, Kakizuka A, Totani K, Yamamoto K, Ashida H (2010) Glucosamine induces autophagy via an mTOR-independent pathway. Biochem Biophys Res Commun 391(4):1775–1779. doi:10.1016/j.bbrc.2009.12.154
Carames B, Kiosses WB, Akasaki Y, Brinson DC, Eap W, Koziol J, Lotz MK (2013) Glucosamine activates autophagy in vitro and in vivo. Arthritis Rheum 65(7):1843–1852. doi:10.1002/art.37977
Eisenberg T, Knauer H, Schauer A, Buttner S, Ruckenstuhl C, Carmona-Gutierrez D, Ring J, Schroeder S, Magnes C, Antonacci L, Fussi H, Deszcz L, Hartl R, Schraml E, Criollo A, Megalou E, Weiskopf D, Laun P, Heeren G, Breitenbach M, Grubeck-Loebenstein B, Herker E, Fahrenkrog B, Frohlich KU, Sinner F, Tavernarakis N, Minois N, Kroemer G, Madeo F (2009) Induction of autophagy by spermidine promotes longevity. Nat Cell Biol 11(11):1305–1314. doi:10.1038/ncb1975
Morselli E, Marino G, Bennetzen MV, Eisenberg T, Megalou E, Schroeder S, Cabrera S, Benit P, Rustin P, Criollo A, Kepp O, Galluzzi L, Shen S, Malik SA, Maiuri MC, Horio Y, Lopez-Otin C, Andersen JS, Tavernarakis N, Madeo F, Kroemer G (2011) Spermidine and resveratrol induce autophagy by distinct pathways converging on the acetylproteome. J Cell Biol 192(4):615–629. doi:10.1083/jcb.201008167
Gupta VK, Scheunemann L, Eisenberg T, Mertel S, Bhukel A, Koemans TS, Kramer JM, Liu KS, Schroeder S, Stunnenberg HG, Sinner F, Magnes C, Pieber TR, Dipt S, Fiala A, Schenck A, Schwaerzel M, Madeo F, Sigrist SJ (2013) Restoring polyamines protects from age-induced memory impairment in an autophagy-dependent manner. Nat Neurosci 16(10):1453–1460. doi:10.1038/nn.3512
Park S, Mori R, Shimokawa I (2013) Do sirtuins promote mammalian longevity? A critical review on its relevance to the longevity effect induced by calorie restriction. Mol Cells 35(6):474–480. doi:10.1007/s10059-013-0130-x
Pyo JO, Yoo SM, Ahn HH, Nah J, Hong SH, Kam TI, Jung S, Jung YK (2013) Overexpression of Atg5 in mice activates autophagy and extends lifespan. Nat Commun 4:2300. doi:ncomms3300 [pii] 10.1038/ncomms3300
Simonsen A, Cumming RC, Brech A, Isakson P, Schubert DR, Finley KD (2008) Promoting basal levels of autophagy in the nervous system enhances longevity and oxidant resistance in adult Drosophila. Autophagy 4(2):176–184, doi:5269 [pii]
Bai H, Kang P, Hernandez AM, Tatar M (2013) Activin signaling targeted by insulin/dFOXO regulates aging and muscle proteostasis in Drosophila. PLoS Genet 9(11):e1003941. doi:10.1371/journal.pgen.1003941 PGENETICS-D-13-01286 [pii]
Demontis F, Perrimon N (2010) FOXO/4E-BP signaling in Drosophila muscles regulates organism-wide proteostasis during aging. Cell 143(5):813–825. doi:10.1016/j.cell.2010.10.007
Ling D, Salvaterra PM (2009) A central role for autophagy in Alzheimer-type neurodegeneration. Autophagy 5(5):738–740, doi:8626 [pii]
Cuervo AM, Dice JF (2000) Age-related decline in chaperone-mediated autophagy. J Biol Chem 275(40):31505–31513. doi:10.1074/jbc.M002102200 M002102200 [pii]
Vittorini S, Paradiso C, Donati A, Cavallini G, Masini M, Gori Z, Pollera M, Bergamini E (1999) The age-related accumulation of protein carbonyl in rat liver correlates with the age-related decline in liver proteolytic activities. J Gerontol A Biol Sci Med Sci 54(8):B318–B323
Ye W, Xu K, Huang D, Liang A, Peng Y, Zhu W, Li C (2011) Age-related increases of macroautophagy and chaperone-mediated autophagy in rat nucleus pulposus. Connect Tissue Res 52(6):472–478. doi:10.3109/03008207.2011.564336
Kaushik S, Arias E, Kwon H, Lopez NM, Athonvarangkul D, Sahu S, Schwartz GJ, Pessin JE, Singh R (2012) Loss of autophagy in hypothalamic POMC neurons impairs lipolysis. EMBO Rep 13(3):258–265. doi:10.1038/embor.2011.260
Sarkis GJ, Ashcom JD, Hawdon JM, Jacobson LA (1988) Decline in protease activities with age in the nematode C. elegans. Mech Ageing Dev 45(3):191–201
Del Roso A, Vittorini S, Cavallini G, Donati A, Gori Z, Masini M, Pollera M, Bergamini E (2003) Ageing-related changes in the in vivo function of rat liver macroautophagy and proteolysis. Exp Gerontol 38(5):519–527
Donati A, Cavallini G, Paradiso C, Vittorini S, Pollera M, Gori Z, Bergamini E (2001) Age-related changes in the regulation of autophagic proteolysis in rat isolated hepatocytes. J Gerontol A Biol Sci Med Sci 56(7):B288–B293
Cavallini G, Donati A, Gori Z, Pollera M, Bergamini E (2001) The protection of rat liver autophagic proteolysis from the age-related decline co-varies with the duration of anti-ageing food restriction. Exp Gerontol 36(3):497–506
Donati A, Cavallini G, Paradiso C, Vittorini S, Pollera M, Gori Z, Bergamini E (2001) Age-related changes in the autophagic proteolysis of rat isolated liver cells: effects of antiaging dietary restrictions. J Gerontol A Biol Sci Med Sci 56(9):B375–B383
Chapin HC, Okada M, Merz AJ, Miller DL (2015) Tissue-specific autophagy responses to aging and stress in C. elegans. Aging 7(6):419–434
Saha S, Ash PE, Gowda V, Liu L, Shirihai O, Wolozin B (2015) Mutations in LRRK2 potentiate age-related impairment of autophagic flux. Mol Neurodegener 10:26. doi:10.1186/s13024-015-0022-y
Wilkinson DS, Jariwala JS, Anderson E, Mitra K, Meisenhelder J, Chang JT, Ideker T, Hunter T, Nizet V, Dillin A, Hansen M (2015) Phosphorylation of LC3 by the Hippo kinases STK3/STK4 is essential for autophagy. Mol Cell 57(1):55–68. doi:10.1016/j.molcel.2014.11.019
Lehtinen MK, Yuan Z, Boag PR, Yang Y, Villen J, Becker EB, DiBacco S, de la Iglesia N, Gygi S, Blackwell TK, Bonni A (2006) A conserved MST-FOXO signaling pathway mediates oxidative-stress responses and extends life span. Cell 125(5):987–1001. doi:10.1016/j.cell.2006.03.046
Altintas O, Park S, Song HK (2016) The role of insulin/IGF-1 signaling in the longevity of model invertebrates, C. elegans and D. melanogaster. BMB Rep 49(2):81–92
Rera M, Azizi MJ, Walker DW (2013) Organ-specific mediation of lifespan extension: more than a gut feeling? Ageing Res Rev 12(1):436–444. doi:10.1016/j.arr.2012.05.003
Kaletsky R, Lakhina V, Arey R, Williams A, Landis J, Ashraf J, Murphy CT (2016) The C. elegans adult neuronal IIS/FOXO transcriptome reveals adult phenotype regulators. Nature 529(7584):92–96. doi:10.1038/nature16483
Dillin A, Crawford DK, Kenyon C (2002) Timing requirements for insulin/IGF-1 signaling in C. elegans. Science 298(5594):830–834
Visvikis O, Ihuegbu N, Labed SA, Luhachack LG, Alves AM, Wollenberg AC, Stuart LM, Stormo GD, Irazoqui JE (2014) Innate host defense requires TFEB-mediated transcription of cytoprotective and antimicrobial genes. Immunity 40(6):896–909. doi:10.1016/j.immuni.2014.05.002
Lin L, Yang P, Huang X, Zhang H, Lu Q, Zhang H (2013) The scaffold protein EPG-7 links cargo-receptor complexes with the autophagic assembly machinery. J Cell Biol 201(1):113–129. doi:10.1083/jcb.201209098
McColl G, Rogers AN, Alavez S, Hubbard AE, Melov S, Link CD, Bush AI, Kapahi P, Lithgow GJ (2010) Insulin-like signaling determines survival during stress via posttranscriptional mechanisms in C. elegans. Cell Metab 12(3):260–272. doi:10.1016/j.cmet.2010.08.004
Pan KZ, Palter JE, Rogers AN, Olsen A, Chen D, Lithgow GJ, Kapahi P (2007) Inhibition of mRNA translation extends lifespan in C. elegans. Aging Cell 6(1):111–119. doi:ACE266 [pii] 10.1111/j.1474-9726.2006.00266.x
Acknowledgments
I wish to acknowledge Hansen lab members and Dr. Anne O’Rourke for feedback on the manuscript, and Dr. Caroline Kumsta for help with Table 15.2. MH was supported by NIH/NIA (R01 AG038664 and R01 AG039756) and a Julie Martin Mid-Career Award in Aging Research supported by The Ellison Medical Foundation and AFAR.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Hansen, M. (2017). Autophagy and Ageing. In: Olsen, A., Gill, M. (eds) Ageing: Lessons from C. elegans. Healthy Ageing and Longevity. Springer, Cham. https://doi.org/10.1007/978-3-319-44703-2_15
Download citation
DOI: https://doi.org/10.1007/978-3-319-44703-2_15
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-44701-8
Online ISBN: 978-3-319-44703-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)