Abstract
The primary purpose of sex is reproduction. However, because not all mating events result in fertilization and only a small number of species provide biparental care to their young, successfully reproducing individuals can rarely be identified from behavioral observations alone. Genetic tools permit reliable identification of an individual’s parents and thus of successfully reproducing individuals, because each parent passes on half of their genetic material to their offspring. In cetaceans, genetic tools are required to identify a female’s already weaned offspring and to detect successfully reproducing males due to the absence of paternal care. To date, relatively few studies have investigated variables linked to reproductive success in this taxon, owed to the difficulty of sampling entire cetacean populations. We summarize currently known factors that are linked to successful reproduction in whales, porpoises, and dolphins, as well as in terrestrial mammals with comparable life histories that give birth to single young.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
2.1 Introduction
Sex cannot adequately be studied without considering its consequences. At first glance, it seems obvious that sex may lead to the production of offspring. However, in most species, more mating events take place than fertilizations, raising the question of which matings are actually successful. This question is of particular importance in species where individuals mate with more than a single partner (polygamy), as it is the case in most mammal species (Clutton-Brock 1989; Würsig et al. 2023, this book). The identification of successfully reproducing individuals is of evolutionary significance because only mating events that result in the production of offspring contribute to the next generation’s gene pool and thus to an individual’s evolutionary fitness.
Behavioral observations provide insights into “who mates with whom?”, while genetic tools shed light on “who sires whose offspring?”. Although the answer to these two questions can be the same, research across mammals has shown that in most species, only a subset of individuals that are ready to mate successfully sire offspring (Lukas and Clutton-Brock 2014). Since the 1990s, when genetic tools became readily available to ecologists, multiple studies have explored parentage in natural populations (Flanagan and Jones 2019), while only a limited number of studies involving genetic tools have investigated reproductive success in cetaceans. Marine mammals are generally more difficult to study than terrestrial ones. As well, marine mammals have slow life histories, requiring populations to be studied over long periods of time. In addition, some cetacean species, particularly whales, have a wide distribution with migration routes spanning half the globe (Stern and Friedlaender 2018), increasing the difficulty in sampling populations. In this chapter, we introduce the genetic tools used to investigate reproductive success and provide an overview of what is known to influence reproductive success in terrestrial mammals with high cognitive abilities, slow life histories, and giving birth to single offspring and thus are expected to face similar constraints as cetaceans. We then summarize the studies carried out in cetaceans before drawing comparisons between cetaceans and terrestrial mammals.
2.1.1 The Need for Genetic Tools to Understand Reproductive Success
During the first days (3–5 days in the hooded seal (Cystophora cristata) or years (1.5–3+ years in the sperm whale, Physeter macrocephalus) of their lives, mammals depend on their mothers for milk. Successfully reproducing female mammals can therefore be reliably recognized via behavioral observations of them with dependent offspring. The identification of successfully reproducing males, in contrast, requires genetic tools. The reasons for this necessity are that even in closely monitored populations not all matings are recorded. Furthermore, there is a considerable number of extra-pair matings in monogamous species, extra-group copulations in polygynous populations (one-male multi-female groups), matings with multiple partners in polygamous species such as cetaceans (Würsig et al. 2023, this book), and the lack of paternal care in most mammal species (Kleinman 1977). In cetaceans, the challenge of identifying successfully reproducing males based on behavioral data alone is further exacerbated by copulations occurring below the surface, while behavioral data are mostly collected via boat-based surveys. Furthermore, in long-lived animals, genetic tools can aid in assigning individuals to their mothers once mature, which can prove useful to increase our knowledge on populations where long-term behavioral records are unavailable.
2.2 Genetic Tools for Parentage Analysis
2.2.1 Genetic Sampling
Genetic analyses are based on DNA, the hereditary material of almost all organisms. Most cells of an individual mammal contain two almost identical copies of its full genome. Thus, genetic analyses can be carried out from any source containing an individual’s cells, such as the skin, muscle, or whole blood. To date, most genetic analyses in cetaceans are based on skin samples obtained via biopsy dart (Baker et al. 2018). The biopsy darts, designed to retain the skin’s top layers as well as some of the underlying blubber, are fired from a modified rifle or a crossbow (Fig. 2.1; Lambertsen 1987; Krützen et al. 2002). Wound healing usually progresses well after sampling, with no evidence of infection at the biopsy site (Krützen et al. 2002). Furthermore, there are no known long-term behavioral consequences of collecting biopsies, as individuals resume their activities often within minutes after having been sampled (Barrett-Lennard et al. 1996; Krützen et al. 2002).
A skin sample is collected from a bottlenose dolphin in Shark Bay using a modified rifle (left panel). The biopsy dart penetrates the skin and then bounces free of the animal while retaining a skin sample (middle panel). The dart consists of a steel tip holding the skin sample and a floating polycarbonate body that permits easy sample recovery at sea (right panel). Image credit: Shark Bay Dolphin Project, Svenja Marfurt (left panel), Samuel Wittwer (middle and right panel)
Alternative, less invasive sampling methods have been proposed for cetaceans such as DNA sampling from blow (Frère et al. 2010c), skin swabs (Harlin et al. 1999), or feces (Parsons et al. 2003a). All of these alternatives require close contact to cetaceans for material collection, are more time-consuming compared to biopsy sampling, and do not present a feasible alternative for most studies (Parsons et al. 2003a; Frère et al. 2010c). However, these alternative approaches can yield valuable insights as their collection can supplement genetic information with hormone analyses to measure stress or reproductive status. Over the past years, researchers began to analyze DNA fragments present in aquatic environments as a result of metabolic waste, such as shed dead skin cells (Ruppert et al. 2019). The DNA fragments collected non-invasively from the environment are referred to as environmental DNA (eDNA). A major advantage of eDNA sampling is that no or very few permits are needed for sampling and that sample collection can also occur unmonitored by leaving a passive filtration system in the water (Bessey et al. 2021). To date, eDNA is mainly used to identify the presence of species. However, emerging techniques might soon permit individual-level analysis such as paternity and maternity analyses (Adams et al. 2019).
2.2.2 Parentage Analysis
Using genetic parentage analysis, an individual’s offspring can be identified because it inherits one half of each parent’s genome. Accordingly, parent-offspring relationships can be resolved using genetic techniques. To date, most genetic parentage analyses in natural animal populations have been conducted by analyzing 10 to 20 highly variable microsatellites (Flanagan and Jones 2019). Microsatellites are fragments in the genome consisting of repeated sequence motifs of one to six DNA base pairs (e.g., GA or TAC as a repeat of a two or three base pair motif, respectively). Individual microsatellite markers have multiple alleles differing in repeat number and thus fragment length. Owed to the elevated mutation rate of microsatellites compared to nuclear DNA (Lynch 2007), they are highly variable, resulting in differing microsatellite “fingerprints” between individuals. Because each parent contributes one half to the genome of their offspring, the genetic microsatellite fingerprint of a descendant matches half their mother’s and half their father’s (Fig. 2.2).
Microsatellite “fingerprints” of a hypothetical mother, her three offspring, and a candidate father. Each offspring shares half of each parents’ alleles. Hence, offspring 1 obtained allele 204 from the mother, while allele 220 must stem from the father. Similarly, offspring 2’s copy of allele 208 must be present in the father because offspring 2 received allele 230 from the mother. Offspring 3 inherited the mother’s copy of allele 204. Because allele 212 is not present in the candidate father, offspring 3 was most likely sired by another male in the population than the candidate father
There are three main approaches to parentage analysis: exclusion, likelihood-based parentage assignment, and Bayesian parentage analysis (Jones et al. 2010). Exclusion is an approach assuming that an individual can be excluded as a parent when none of its alleles matches the offspring under consideration. Although this approach appears compelling, it is rarely used nowadays because it has multiple pitfalls, such as scoring errors that can lead to the true parent being excluded. The currently most used technique is likelihood-based parentage assignment—a method based on likelihood ratios between the two competing hypotheses that two individuals either represent a parent-offspring dyad or are unrelated (Marshall et al. 1998). The widely used software CERVUS permits likelihood-based parentage assignment employed in a user-friendly graphical user interface (GUI, Kalinowski et al. 2007). Bayesian parentage analysis permits the inclusion of information that is thought to influence reproductive success, such as age or dominance rank. This information is then taken into account when calculating the probability that an individual is another’s parent. The incorporation of such information requires profound knowledge of the population and the species under consideration (Flanagan and Jones 2019). Possibly because such information is often unavailable for natural populations, this approach is rarely used. Independent of which method is chosen, parentage analysis is more powerful in cases where mothers are known and genotyped, as it can be inferred which of the offspring’s alleles are derived from the mother and thus which alleles must stem from the father (Huisman 2017).
Sex information of the genotyped individuals can further facilitate parentage analysis as it permits the separation of the genotyped individuals into candidate mothers and fathers. This is valuable in species with low levels of sexual dimorphism as is the case for most delphinids (Mesnick and Ralls 2018a). Genetic sexing has been employed in many studies as a fast and reliable means for sex determination. It is carried out by testing for the presence/absence of sex-chromosomal markers. In mammals, where females are the homogametic sex (XX), only X-chromosomal markers are detected. In contrast, males are the heterogametic sex (XY) and test positive for both X- and Y-chromosomal markers (Fig. 2.3). Sexing in cetaceans is often done by a joint analysis of the X-linked and Y-linked exons of the ZFX and ZFY genes (Bérubé and Palsbøll 1996).
Electrophoresis gel showing PCR products of a reaction amplifying X- and Y-chromosomal markers. Males (samples 993, 995, 999, 1001) have two bands, because they are carriers of both sex chromosomes (XY), while females (samples 994, 996, 997, 998, 1000) can be identified as individuals with single bands (XX) (image credit: Manuela Bizzozzero)
2.2.3 Genetic Marker Systems for Parentage Analysis
Due to their hypervariable nature, microsatellites have long been the most-used genetic marker for parentage analysis (Flanagan and Jones 2019). Across species, microsatellites were the genetic markers of choice to investigate many parameters important in evolution and ecology such as dispersal patterns, migration rates, population size, and kinship (Hodel et al. 2016). However, population geneticists now widely use next-generation sequencing (NGS) approaches. Compared to traditional sequencing approaches, including microsatellite genotyping where only few loci are considered, NGS approaches permit the parallel genotyping of millions of single nucleotide polymorphisms (SNPs). Because these high-resolution SNP data are better suited to address ecological and evolutionary questions, there has been a dramatic decrease of studies using microsatellites over the past decade.
SNPs typically have two different alleles per locus, while microsatellites often have multiple alleles. Compared to a single microsatellite locus, single SNPs are therefore less informative. Reliable parentage assignment can be achieved by analyzing as few as ten highly polymorphic microsatellite markers but requires 100 SNPs (Weng et al. 2021). However, because NGS permits the simultaneous sequencing of millions of SNPs, this requirement is commonly met without difficulty. Like microsatellites, the SNPs used for parentage analysis are inherited in a Mendelian fashion, meaning that the offspring receives one copy from each parent. Thus, the same suite of analytical software can be used. Furthermore, compared to microsatellite data, a large number of SNPs derived from an NGS approach are much better suited to estimate pairwise relatedness, thereby permitting to assign dyads to other relationship categories than parent-offspring.
2.3 Paternity Success in Male Mammals
2.3.1 Variables Influencing Reproductive Success in Terrestrial Male Mammals
In most mammal species, more males are ready to reproduce than females because paternal care is absent in 95%–97% of species (Kleinman 1977) and the production of offspring requires a considerable time and energy investment from females, caused by gestation and lactation. This difference in parental investment causes a conflict between the sexes, where males often compete which each other over access to females. Because some males are better competitors than others, or successfully employ alternative non-competitive strategies (e.g., sneaking fertilizations without the knowledge of other males), the reproductive success among males is highly variable. For example, the variance of male lifetime reproductive success in rhesus macaques (Macaca mulatta) is five times larger compared to females (Dubuc et al. 2014).
Given that reproduction for males mainly consists of mating, male reproductive success is influenced by access to fertile females. Depending on the distribution of females, males employ different strategies (van Schaik and van Hooff 1994). If females are highly dispersed, males are likely to have less control over access to females compared to females aggregated in groups with high site fidelity. Where females can be monopolized, males frequently engage in contest competition, involving aggressive behavior, but also in sperm competition, attempting to outcompete other males that mate with the same female by ejaculating larger sperm quantities. In contrast, in populations where females are more dispersed, males are more likely to employ a roaming strategy (scramble competition), aiming to find and mate with females before others do. Furthermore, females might be more willing to mate with certain males (mate choice competition), potentially such with persuasive courtship behavior. These male mating tactics are not mutually exclusive, requiring males to compete on multiple levels, further complicating a male’s pursuit for a mate.
In most mammals, females remain in their natal area (Greenwood 1980) and as a result cluster with their female relatives. To avoid inbreeding, males often leave their natal area once mature. To reproduce, males join new groups, where they compete with other males over reproductive opportunities. These opportunities can arise by replacing the breeding male of a polygynous (single-male, multi-female) group. In polygynandrous (multi-male, multi-female) groups, males frequently compete with other males to attain a high rank because dominant males sire more offspring in many species (Moore et al. 1995; Clutton-Brock and Isvaran 2006; Majolo et al. 2012). Male dominance is often established by agonistic interactions. Hence, body size and strength are good predictors of male status and thus reproductive success. Nevertheless, it is rare that a dominant male exclusively sires all offspring in a group (Clutton-Brock and Isvaran 2006). Genetic tests found that over 80% of all offspring can be sired by males other than the alpha male in rhesus macaques, with females adjusting their willingness to mate with subordinates depending on whether other group members were present or not (Overduin-de Vries et al. 2012). Subordinate males therefore appeared to use a different mating tactic, engaging in sneaky copulations which the dominant male does not notice. A male’s ability to monopolize offspring is thus also influenced by his capability to closely guard females (Clutton-Brock and Isvaran 2006). This is also true for single-male, multi-female mating systems. Although the resident male sires, on average, a larger proportion of offspring in this mating system compared to one where multiple adult males are present, genetic tests revealed that a low percentage of paternities are frequently obtained by another male than the group’s single resident adult male (Clutton-Brock and Isvaran 2006).
Body size and strength are not only important in stable single or multi-male-female groups but also in species forming all-female groups. African elephants (Loxodonta africana), for example, form highly mobile groups consisting of a lead female (the matriarch), her offspring, and sometimes the matriarch’s sisters and their offspring (Archie et al. 2006). Female offspring remain in the group, but males leave the group once mature. Female elephants are fertile for a short window of 3 to 6 days every 3 to 9 years (Moss and Poole 1983; Poole and Moss 1989). As a result, male elephants face the challenge of locating an incredibly limited and highly mobile resource while preventing access from other males (Poole 1989; Poole and Moss 1989). Males are expected to be better competitors with increasing size. As a result, male elephants might have been selected to grow throughout their lives (Lee and Moss 1995). Paternity analyses in elephants confirmed that older and hence larger elephants sired more offspring than younger males (Hollister-Smith et al. 2007; Rasmussen et al. 2007). This effect was even more pronounced when males were in largely testosterone-driven musth, a condition where males are more aggressive and sexually active.
In some species, males cooperate to gain access to females or attain a higher rank (Smith 2014), which increases their chances to mate. Such male cooperation mostly occurs in the form of temporary coalitions in which multiple males collaborate to compete against a single or multiple others. Due to the indivisibility of fertilizations, male cooperation poses an evolutionary paradox: although all males get to mate, only a single male succeeds in siring offspring. However, kin selection can resolve this paradox in cases where coalitions or alliances consist of relatives. Genetic studies confirmed that kin selection underlies cooperation in male cheetahs (Acinonyx jubatus, Caro 1990; Caro and Kelly 2019) and some, but not all, coalitions in lions (Panthera leo, Packer et al. 1991; Chakrabarti et al. 2020) and chimpanzees (Pan troglodytes, Mitani et al. 2000; Langergraber et al. 2007).
In cases where alliances and coalitions were not found to be kin-biased, cooperation often occurred among males with close social bonds (Berghänel et al. 2011; Feldblum et al. 2021; Gerber et al. 2022). Social bonds can be defined as affiliative and persisting relationships and are sometimes referred to as “friendships” (Silk 2002; Cords and Thompson 2017; Massen 2017). A study in chimpanzees revealed that males with vast social networks and strong social bonds to others sired more offspring compared to males with few or weak social bonds (Feldblum et al. 2021). In Barbary macaques (Macaca sylvanus), males affiliating in the non-mating season formed coalitions during the mating season (Berghänel et al. 2011); the strong social bonds facilitating coalition formation in this species correlated with future social status and thereby paternity success (Schülke et al. 2010). Although kinship facilitated social bond formation, the majority of social bonds were formed among non-kin (De Moor et al. 2020). Coalition formation thus can increase a male’s direct and indirect fitness.
2.3.2 Variables Contributing to Reproductive Success in Male Cetaceans
Female cetaceans are highly mobile, often dispersed, and have three dimensions to escape mating attempts by males. Thus, cetacean females cannot easily be monopolized, resulting in males having little control over access to females. Because of this, most male cetaceans have to search for receptive females to mate with while outcompeting other males, either by mate guarding, physical fights, or display competition like songs (Mesnick and Ralls 2018a, b).
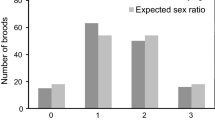
Genetic paternity tests in multiple cetacean species found that paternity skew was low, thereby confirming that males lack control over access to females and thus are likely to employ a roaming approach to find females. In humpback whales (Megaptera novaeangliae), 62 calves were assigned to 51 fathers, indicating that most males who successfully sired an offspring did so only once; no male was identified as the father of more than three calves (Cerchio et al. 2005). Similarly, in Atlantic spotted dolphins (Stenella frontalis), seven males sired ten offspring (Green et al. 2011), all of whom were 18 years or older despite males reaching sexual maturity between 12 and 15 years old, suggesting that older males have higher chances of siring offspring compared to younger ones. In North Atlantic right whales (Eubalaena glacialis) and killer whales (Orcinus orca), genetic analyses found reproductive success to be skewed toward older males (Frasier et al. 2007; Ford et al. 2011). In killer whales, aggressive encounters between males have rarely been observed, implying that the greater reproductive success of older males compared to younger males is because they are preferred by females or due to them having an advantage in sperm competition. In North Atlantic right whales, a single female and 2 to 40 males form mating groups referred to as surface active groups (SAGs), within which males aggressively compete for positions closest to the female (Kraus and Hatch 2001; Parks et al. 2007). Over the course of an average SAG, lasting 1 hour, the female copulates approximately 60 times with multiple males, implying intense sperm competition (Kraus and Hatch 2001). Considering that testes may not yet be fully developed in young adult males engaging in SAGs, older males may indeed have an advantage (Frasier et al. 2007).
Bottlenose dolphins (Tursiops spp.) have a wide distribution with distinct morphological and behavioral differences among populations. In some populations where sexual size dimorphism is low, males form cooperative alliances to mate with females (Möller et al. 2001; Parsons et al. 2003b; Whitehead and Connor 2005). Compared to acting alone, multiple cooperating males are believed to be better at preventing females from escaping coerced matings. Additionally, multiple males can outcompete single males. In Indo-Pacific bottlenose dolphins (Tursiops aduncus) in Shark Bay, Australia, for example, non-allied males sired no or very few offspring (Krützen et al. 2004; Gerber et al. 2022). A study on the same species but in a different location (Port Stephens, Australia) found that alliance size correlated with reproductive success, suggesting that larger alliances have higher chances of siring offspring compared to smaller ones (Wiszniewski et al. 2012). However, cooperating with others to gain mating opportunities might be costly because only one male will be able to sire a female’s single offspring per pregnancy.
Multiple studies investigated whether kin selection can explain alliance formation in bottlenose dolphins. In a Tursiops cf. australis population in South Australia and a Tursiops truncatus population in the Bahamas, allied males appeared to be more closely related than expected by chance (Parsons et al. 2003b; Diaz-Aguirre et al. 2018). However, this was not the case for the Tursiops aduncus populations in Shark Bay and Port Stephens, both in Australia (Möller et al. 2001; Gerber et al. 2021). Paternity success in Shark Bay was predicted by social integration; male dolphins with strong social bonds to their alliance partners sired more offspring compared to those with weaker bonds (Gerber et al. 2022). Thus, the differences between these populations might disappear if conducted with more comparable datasets and methods. Bottlenose dolphins are the only cetacean taxon where male reproductive success has been studied in multiple populations over a wide geographic scale. The differing results suggest that males employ different mating tactics, potentially dependent on their ecological and social environments that can differ within a species. Whether this is also the case in other cetacean taxa remains to be investigated.
2.4 Reproductive Success of Female Mammals
2.4.1 Variables Influencing Reproductive Success in Females
The reproductive success of females is influenced by their access to resources and their reproductive timespan as a result of the energetic and temporal demands of gestation and lactation (Clutton-Brock 1989). Young mammals are dependent on their mothers for nutrition and are therefore found in association with their mothers during the first period of their lives. For that reason, genetic tests are rarely required to identify a female’s offspring, at least not in well-monitored long-term study populations. However, genetic tools can be useful to identify whether females embedded in a vast kin network have higher lifetime reproductive success compared to such with few relatives.
Philopatry, defined as an individual’s tendency to remain in the area where it was born (Mayr 2013), increases the chances to have access to kin. In most mammals, females are philopatric, possibly because females gain more benefits from remaining in their natal area than males (Greenwood 1980). Benefits include the avoidance of the energetic demands of dispersal and the maintenance of a familiar diet in a familiar habitat with familiar individuals (Clutton-Brock and Lukas 2012). In group-living individuals, female philopatry results in females being in the same social groups as their relatives (Clutton-Brock and Lukas 2012; van Noordwijk et al. 2012), while in solitary species, female relatives frequently have adjoining habitats, as, for example, observed in Bornean orangutans (Pongo pygmaeus) (van Noordwijk et al. 2012). Using genetic tools, it was found that although related and unrelated female orangutans had similar home-range overlaps, related females spent more time in association and permitted their offspring to play, which was not the case for the offspring of unrelated females (van Noordwijk et al. 2012). Similarly, a study in African elephants found that group fusions were more likely to occur when the matriarchs of the groups were related than unrelated (Archie et al. 2006). Moreover, fissions within a group were influenced by genetic relatedness; female elephants remained in the same group as their relatives (Archie et al. 2006).
Yellow baboons (Papio cynocephalus) live in multi-male with multi-female groups. Females are philopatric. Unlike males, where social status depends on the outcome of aggressive interactions, females inherit the social status of their mothers (Samuels et al. 1987). Compared to low-ranking females, high-ranking females benefit from better access to resources and thus often have large-for-age offspring (Altmann and Alberts 2005). Yet, the influence of dominance rank on female reproductive success is generally low (Altmann and Alberts 2003; Cheney et al. 2004). However, the offspring of females with close social bonds to other females had higher rates of offspring survival and lived longer compared to females with weaker social bonds (Silk et al. 2009). Most social bonds were formed among related females. Nevertheless, females without relatives formed social bonds to non-kin conveying the same fitness benefits (Silk et al. 2009). Thus, social bonds to relatives and non-relatives contribute to female reproductive success. Overall, studies on terrestrial mammals suggest that solitary as well as group-living females benefit from affiliating with their female relatives.
2.4.2 The Influence of Female Relatives on Reproductive Success in Cetaceans
Cetaceans are long-lived mammals with slow life histories; after a gestational period of 9–17 months, females give birth to a single calf (Drinkwater and Branch 2022). Calves are dependent on their mothers for the first period of their lives, leading to long inter-birth intervals ranging from 1 to 7 years (Mesnick and Ralls 2018b). In all cetacean species, calves are born precocial, meaning they can move independently, come to the surface for air, and maintain proximity with their mothers from birth (Whitehead and Mann 2000). However, females benefit from being in association with other females through cooperative hunting, increased vigilance, joint defense of their calves and themselves, and potentially allomaternal care (i.e., temporal care of a calf by a non-mother; Würsig et al. 2023, this book). In sperm whales (Physeter macrocephalus), for example, young individuals are accompanied at the surface by different group members, while other group members, including mothers, forage at depth (Whitehead 1996; Gero et al. 2009).
Female group composition is often influenced by kinship. Killer and sperm whales, for example, form stable matrilineal units consisting of a female and her male and female offspring (Ford 2018). A kinship analysis in sperm whales found that females preferably affiliated with close kin within social units (Konrad et al. 2018). However, maternal relatives also maintain stronger bonds in dynamic fission-fusion societies consisting of multiple matrilines, such as bottlenose dolphins (Frère et al. 2010b).
In the well-studied Indo-Pacific bottlenose dolphins in Shark Bay, Australia, females form loose kin-biased social networks (Frère et al. 2010b). Female dolphins inherit the social network of their mother (Frère et al. 2010a), which affects their reproductive success because calving success (Frère et al. 2010b) and the survival of male offspring (Stanton and Mann 2012) are influenced by a female’s social bonds to others. A potential influence of social bonds on reproductive success was also found in female humpback whales; female pairs that were observed together over multiple years sired the most offspring (Ramp et al. 2010). It is unclear whether these associations were kin-biased or not. However, research on different individuals in the same location found maternally related females more likely to associate than expected by chance (Weinrich et al. 2006), implying that associations in this species might contribute to direct and indirect fitness.
In at least some killer whale populations, females form “pods” consisting of a matriarch and her sons and daughters. The calves of older matriarchs suffer from higher mortality rates compared to their daughter’s offspring in the same group (Croft et al. 2017). Furthermore, the presence of a post-reproductive mother increased survival of her older sons (Foster et al. 2012). With increasing age, the indirect fitness benefits gained from helping offspring might therefore outweigh the direct fitness benefits gained from reproducing. This might have contributed toward the evolution of reproductive senescence (menopause) in this species. The evolutionary fitness of female cetaceans can thereafter not simply be understood as a by-product of resource availability but depends on a species’ social structure and the availability of kin therein.
2.5 Comparison Between Terrestrial and Marine Mammals
The transition from terrestrial into marine habitats by the predecessor of marine mammals was facilitated by morphological, physiological, and behavioral adaptations. However, despite large morphological differences, marine and terrestrial mammals with slow life histories and singleton births face similar constraints resulting in analogies. Cetaceans and long-lived terrestrial species, such as primates and elephants, thus bear striking behavioral similarities; all three possess high cognitive skills and have the ability for social learning (Lee and Moss 1999; Whiten and van de Waal 2017; Whitehead and Rendell 2021). Primates, cetaceans, and elephants belong to different taxonomic orders (primates, Primata; elephants, Proboscidea; cetaceans, Artiodactyla or Cetartiodactyla). Thus, these shared traits are the result of convergent evolution (i.e., they have evolved independently). Genetic studies in marine and terrestrial mammals established that analogies among marine and terrestrial mammals can also be observed as regards their reproduction; the monopolization potential of females affects male reproductive success, while females benefit from being in association with relatives. However, there are also differences among the species inhabiting land and sea.
Like most mammals, cetaceans are either polygynous or polygynandrous. However, reproductive skew in marine mammals is much lower compared to terrestrial mammals (Frasier et al. 2007), possibly because males have less control over access to females in aquatic species where females can move in three dimensions or because paternity data are still scarce even in the most-studied populations. If multiple males cooperate, females are less likely to outmaneuver males, which might have contributed to the evolution of male alliances in species that are able to move in three dimensions such as chimpanzees with remarkable climbing skills (Watts 1998), some birds (e.g., long-tailed manakins (Chiroxiphia linearis), McDonald and Potts 1994), and bottlenose dolphins (Connor and Krützen 2015).
Social status has a profound effect on male reproductive success in a multitude of mammalian species. Yet, little is known of the existence of dominance hierarchies in cetaceans (Tyack 2018). Although the lack of supporting evidence for dominance hierarchies in cetaceans does not mean that they are non-existent, it is likely that dominance hierarchies do not govern inter-individual interactions to the same extent as in terrestrial species. Compared to females in terrestrial mammal species, female cetaceans can move in three dimensions and thus might have increased abilities to avoid matings with undesired males. Furthermore, marine food sources such as fish and krill are widely distributed and cannot easily be monopolized by social groups, resulting in vast overlapping home ranges or migratory lifestyles. Lack of controlled access to females and of clustered resources may have contributed to the (apparent) lack of social hierarchies.
The lack of social hierarchies, however, does not mean that social interactions are of less importance in marine compared to terrestrial mammals. The presently most complex social system known outside of humans is in male bottlenose dolphins in Shark Bay, Australia, that cooperate in multi-level alliances over access to females (Connor and Krützen 2015). Similar to humans (Snyder-Mackler et al. 2020) and chimpanzees (Feldblum et al. 2021), same-sex social bonds positively contributed to the evolutionary fitness of male and female bottlenose dolphins in Shark Bay (Frère et al. 2010b; Gerber et al. 2022). In females but not males, social bonds are often biased toward relatives (Frère et al. 2010b; Gerber et al. 2021).
In African elephants, a matriarch’s ability to assess threats from predators increases with age (McComb et al. 2011). In killer whales, old females lead their matrilines to alternative feeding grounds when prey abundance at their current site is low, thereby ensuring the survival and health of their relatives, in particular of their adult sons (Brent et al. 2015). The indirect fitness benefits gained from assisting relatives, combined with the increased mortality rates of their own offspring with age, might have contributed to the evolution of reproductive senescence in killer whales. This is similar to humans, where grandmothers increase their inclusive fitness by caring and providing for their daughter’s children (Shanley et al. 2007). Mothers can also positively influence the reproductive success of their sons. In bonobos (Pan paniscus), males that live in the same groups as their mothers sire more offspring compared to males without their mothers (Surbeck et al. 2019). The influence of maternal presence on male reproductive success in cetaceans is largely unexplored. However, a female killer whale cooperated with her adult son in killing an unrelated female’s calf (Towers et al. 2018), potentially to increase his own reproduction. In order to aid their sons, females may hinder other males from mating or bring their sons in proximity to estrus females as observed in bonobos (Surbeck et al. 2011). In cetaceans, mothers could positively influence the fitness of their sons where both sexes remain in their natal area and sexual dimorphism is low, such as for some bottlenose dolphin populations or other delphinids (Mesnick and Ralls 2018a).
2.6 Conclusions and Future Directions
Genetic advances over the past two to four decades have confirmed what scientists, dating back to the theories of Darwin, already suspected: factors improving a male’s access to females increase male reproductive success while female reproductive success is positively affected by variables influencing their own and their offspring’s survival. The large diversity in reproductive strategies and tactics across mammals exemplifies that there are often multiple ways that reproductive success can be maximized. Similarities occur between terrestrial and marine species, while in each realm there is large diversity; this implies that reproductive strategies are often the result of convergent evolution and that somewhat similar selective pressures are experienced on land and in the sea.
Due to the slow life histories of cetaceans, paternity studies require that populations are monitored over a long time, and such studies are rare. Nevertheless, the results from long-term investments provide unique insights into mating strategies and tactics, and are invaluable to increase our understanding of how individuals maximize individual (and as a by-product, evolutionary) fitness. Novel molecular techniques might decrease the large amount of time dedicated to sampling and monitoring populations required for parentage analyses; passive eDNA collection might permit the collection of population-wide samples within a few weeks. Furthermore, epigenetic clocks produce reliable age estimates for cetaceans including bottlenose dolphins (Peters et al. 2023), beluga whales (Delphinapterus leucas, Bors et al. 2021), and humpback whales (Horvath et al. 2022). Using epigenetic clocks in populations where individual ages are unknown will greatly facilitate parentage analyses because the direction of a parent-offspring relationship will be known (i.e., the older individual will be assigned as parent of the younger one and not vice versa). In the next decade, advances in molecular biology will permit the ability to fill some of the numerous gaps of knowledge on cetacean reproductive success, thereby learning more about what variables contribute to direct fitness in the marine realm.
References
Adams CIM, Knapp M, Gemmell NJ, Jeunen G-J, Bunce M, Lamare MD, Taylor HR (2019) Beyond biodiversity: can environmental DNA (eDNA) cut it as a population genetics tool? Genes 10(3):192
Altmann J, Alberts S (2003) Offspring: the biodemography of fertility and family behavior. The National Academies Press, Washington, DC
Altmann J, Alberts SC (2005) Growth rates in a wild primate population: ecological influences and maternal effects. Behav Ecol Sociobiol 57(5):490–501. https://doi.org/10.1007/s00265-004-0870-x
Archie EA, Moss CJ, Alberts SC (2006) The ties that bind: genetic relatedness predicts the fission and fusion of social groups in wild African elephants. Proc R Soc Lond B 273(1586):513–522. https://doi.org/10.1098/rspb.2005.3361
Baker CS, Steel D, Nieukirk S, Klinck H (2018) Environmental DNA (eDNA) from the wake of the whales: droplet digital PCR for detection and species identification. Front Mar Sci 5. https://doi.org/10.3389/fmars.2018.00133
Barrett-Lennard L, Smith TG, Ellis GM (1996) A cetacean biopsy system using lightweight pneumatic darts, and its effect on the behavior of killer whales. Mar Mamm Sci 12(1):14–27. https://doi.org/10.1111/j.1748-7692.1996.tb00302.x
Berghänel A, Ostner J, Schröder U, Schülke O (2011) Social bonds predict future cooperation in male barbary macaques, Macaca sylvanus. Anim Behav 81(6):1109–1116. https://doi.org/10.1016/j.anbehav.2011.02.009
Bérubé M, Palsbøll P (1996) Identification of sex in cetaceans by multiplexing with three ZFX and ZFY specific primers. Mol Ecol 5(2):283–287. https://doi.org/10.1111/j.1365-294x.1996.tb00315.x
Bessey C, Neil Jarman S, Simpson T, Miller H, Stewart T, Kenneth Keesing J, Berry O (2021) Passive eDNA collection enhances aquatic biodiversity analysis. Commun Biol 4(1):236. https://doi.org/10.1038/s42003-021-01760-8
Bors EK, Baker CS, Wade PR, O'Neill KB, Shelden KEW, Thompson MJ, Fei Z, Jarman S, Horvath S (2021) An epigenetic clock to estimate the age of living beluga whales. Evol Appl 14(5):1263–1273. https://doi.org/10.1111/eva.13195
Brent LJN, Franks DW, Foster EA, Balcomb KC, Cant MA, Croft DP (2015) Ecological knowledge, leadership, and the evolution of menopause in killer whales. Curr Biol 25(6):746–750. https://doi.org/10.1016/j.cub.2015.01.037
Caro TM (1990) Cheetah mothers bias parental investment in favour of cooperating sons. Ethol Ecol Evol 2(4):381–395. https://doi.org/10.1080/08927014.1990.9525399
Caro TM, Kelly MJ (2019) Cheetahs and their mating system. In: Lee Alan D (ed) Model systems in behavioral ecology. Princeton University Press, Princeton, NJ, pp 512–532. https://doi.org/10.1515/9780691207247-027
Cerchio S, Jacobsen JK, Cholewiak DM, Falcone EA, Merriwether DA (2005) Paternity in humpback whales, Megaptera novaeangliae: assessing polygyny and skew in male reproductive success. Anim Behav 70(2):267–277. https://doi.org/10.1016/j.anbehav.2004.10.028
Chakrabarti S, Kolipakam V, Bump JK, Jhala YV (2020) The role of kinship and demography in shaping cooperation amongst male lions. Sci Rep 10(1):17527. https://doi.org/10.1038/s41598-020-74247-x
Cheney DL, Seyfarth RM, Fischer J, Beehner J, Bergman T, Johnson SE, Kitchen DM, Palombit RA, Rendall D, Silk JB (2004) Factors affecting reproduction and mortality among baboons in the Okavango Delta. Botswana. Int J Primat 25(2):401–428. https://doi.org/10.1023/B:IJOP.0000019159.75573.13
Clutton-Brock TH (1989) Review lecture: mammalian mating systems. Proc R Soc Lond B 236(1285):339–372. https://doi.org/10.1098/rspb.1989.0027
Clutton-Brock TH, Isvaran K (2006) Paternity loss in contrasting mammalian societies. Biol Lett 2(4):513–516. https://doi.org/10.1098/rsbl.2006.0531
Clutton-Brock TH, Lukas D (2012) The evolution of social philopatry and dispersal in female mammals. Mol Ecol 21(3):472–492. https://doi.org/10.1111/j.1365-294X.2011.05232.x
Connor RC, Krützen M (2015) Male dolphin alliances in Shark Bay: changing perspectives in a 30-year study. Anim Behav 103:223–235. https://doi.org/10.1016/j.anbehav.2015.02.019
Cords M, Thompson NA (2017) Friendships, coalitions, and alliances. In: APA handbook of comparative psychology: basic concepts, methods, neural substrate, and behavior, APA handbooks in psychology®, vol 1. American Psychological Association, Washington, DC, pp 899–913. https://doi.org/10.1037/0000011-043
Croft DP, Johnstone RA, Ellis S, Nattrass S, Franks DW, Brent LJN, Mazzi S, Balcomb KC, Ford JKB, Cant MA (2017) Reproductive conflict and the evolution of menopause in killer whales. Curr Biol 27(2):298–304. https://doi.org/10.1016/j.cub.2016.12.015
De Moor D, Roos C, Ostner J, Schülke O (2020) Bonds of bros and brothers: kinship and social bonding in postdispersal male macaques. Mol Ecol 29(17):3346–3360. https://doi.org/10.1111/mec.15560
Diaz-Aguirre F, Parra GJ, Passadore C, Möller L (2018) Kinship influences social bonds among male southern Australian bottlenose dolphins (Tursiops cf. australis). Behav Ecol Sociobiol 72(12):190. https://doi.org/10.1007/s00265-018-2621-4
Drinkwater RW, Branch TA (2022) Estimating proportions of identical twins and twin survival rates in cetaceans using fetal data. Mar Mamm Sci 38(4):1–11. https://doi.org/10.1111/mms.12929
Dubuc C, Ruiz-Lambides A, Widdig A (2014) Variance in male lifetime reproductive success and estimation of the degree of polygyny in a primate. Behav Ecol 25(4):878–889. https://doi.org/10.1093/beheco/aru052
Feldblum JT, Krupenye C, Bray J, Pusey AE, Gilby IC (2021) Social bonds provide multiple pathways to reproductive success in wild male chimpanzees. iScience 24(8):102864. https://doi.org/10.1016/j.isci.2021.102864
Flanagan SP, Jones AG (2019) The future of parentage analysis: from microsatellites to SNPs and beyond. Mol Ecol 28(3):544–567. https://doi.org/10.1111/mec.14988
Ford JKB (2018) Killer whales. In: Würsig B, Thewissen JGM, Kovacs KM (eds) Encyclopedia of marine mammals, 3rd edn. Academic Press, London, pp 531–537
Ford MJ, Hanson MB, Hempelmann JA, Ayres KL, Emmons CK, Schorr GS, Baird RW, Balcomb KC, Wasser SK, Parsons KM, Balcomb-Bartok K (2011) Inferred paternity and male reproductive success in a killer whale (Orcinus orca) population. J Hered 102(5):537–553. https://doi.org/10.1093/jhered/esr067
Foster EA, Franks DW, Mazzi S, Darden SK, Balcomb KC, Ford JKB, Croft DP (2012) Adaptive prolonged postreproductive life span in killer whales. Science 337(6100):1313–1313. https://doi.org/10.1126/science.1224198
Frasier TR, Hamilton PK, Brown MW, Conger LA, Knowlton AR, Marx MK, Slay CK, Kraus SD, White BN (2007) Patterns of male reproductive success in a highly promiscuous whale species: the endangered North Atlantic right whale. Mol Ecol 16(24):5277–5293. https://doi.org/10.1111/j.1365-294X.2007.03570.x
Frère CH, Krützen M, Mann J, Connor RC, Bejder L, Sherwin WB (2010a) Social and genetic interactions drive fitness variation in a free-living dolphin population. Proc Natl Acad Sci 107(46):19949–19954. https://doi.org/10.1073/pnas.1007997107
Frère CH, Krützen M, Mann J, Watson-Capps JJ, Tsai YJ, Patterson EM, Connor R, Bejder L, Sherwin WB (2010b) Home range overlap, matrilineal and biparental kinship drive female associations in bottlenose dolphins. Anim Behav 80(3):481–486. https://doi.org/10.1016/j.anbehav.2010.06.007
Frère CH, Krzyszczyk E, Patterson EM, Hunter S, Ginsburg A, Mann J (2010c) Thar she blows! A novel method for DNA collection from cetacean blow. PLoS One 5(8):e12299. https://doi.org/10.1371/journal.pone.0012299
Gerber L, Wittwer S, Allen SJ, Holmes KG, King SL, Sherwin WB, Wild S, Willems EP, Connor RC, Krützen M (2021) Cooperative partner choice in multi-level male dolphin alliances. Sci Rep 11(1):6901. https://doi.org/10.1038/s41598-021-85583-x
Gerber L, Connor RC, Allen SJ, Horlacher K, King SL, Sherwin WB, Willems EP, Wittwer S, Krützen M (2022) Social integration influences fitness in allied male dolphins. Curr Biol 32(7):1664–1669. https://doi.org/10.1016/j.cub.2022.03.027
Gero S, Engelhaupt D, Rendell L, Whitehead H (2009) Who cares? Between-group variation in alloparental caregiving in sperm whales. Behav Ecol 20(4):838–843. https://doi.org/10.1093/beheco/arp068
Green ML, Herzing DL, Baldwin JD (2011) Reproductive success of male Atlantic spotted dolphins (Stenella frontalis) revealed by noninvasive genetic analysis of paternity. Can J Zool 89(3):239–253. https://doi.org/10.1139/z10-111
Greenwood PJ (1980) Mating systems, philopatry and dispersal in birds and mammals. Anim Behav 28(4):1140–1162
Harlin AD, Würsig B, Baker CS, Markowitz TM (1999) Skin swabbing for genetic analysis: application to dusky dolphins (Lagenorhynchus obscurus). Mar Mamm Sci 15(2):409–425. https://doi.org/10.1111/j.1748-7692.1999.tb00810.x
Hodel RGJ, Segovia-Salcedo MC, Landis JB, Crowl AA, Sun M, Liu X, Gitzendanner MA, Douglas NA, Germain-Aubrey CC, Chen S, Soltis DE, Soltis PS (2016) The report of my death was an exaggeration: a review for researchers using microsatellites in the 21st century. Appl Plant Sci 4(6):1600025. https://doi.org/10.3732/apps.1600025
Hollister-Smith JA, Poole JH, Archie EA, Vance EA, Georgiadis NJ, Moss CJ, Alberts SC (2007) Age, musth and paternity success in wild male African elephants, Loxodonta africana. Anim Behav 74(2):287–296. https://doi.org/10.1016/j.anbehav.2006.12.008
Horvath S, Haghani A, Zoller JA, Fei Z, Bérubé M, Robbins J (2022) DNA methylation age studies of humpback whales. bioRxiv 2022:503952. https://doi.org/10.1101/2022.08.15.503952
Huisman J (2017) Pedigree reconstruction from SNP data: parentage assignment, sibship clustering and beyond. Mol Ecol Res 17(5):1009–1024. https://doi.org/10.1111/1755-0998.12665
Jones AG, Small CM, Paczolt KA, Ratterman NL (2010) A practical guide to methods of parentage analysis. Mol Ecol Res 10(1):6–30. https://doi.org/10.1111/j.1755-0998.2009.02778.x
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol Ecol 16(5):1099–1106
Kleinman DG (1977) Monogamy in mammals. Q Rev Biol 52(1):39–69. https://doi.org/10.1086/409721
Konrad CM, Gero S, Frasier T, Whitehead H (2018) Kinship influences sperm whale social organization within, but generally not among, social units. R Soc Open Sci 5(8):180914. https://doi.org/10.1098/rsos.180914
Kraus SD, Hatch JJ (2001) Mating strategies in the North Atlantic right whale (Eubalaena glacialis). J Cetacean Res Manag Spec Issue 2:237–244
Krützen M, Barré LM, Möller LM, Heithaus MR, Simms C, Sherwin WB (2002) A biopsy system for small cetaceans: darting success and wound healing in Tursiops spp. Mar Mamm Sci 18(4):863–878. https://doi.org/10.1111/j.1748-7692.2002.tb01078.x
Krützen M, Barré LM, Connor RC, Mann J, Sherwin WB (2004) ‘O father: where art thou?’— paternity assessment in an open fission–fusion society of wild bottlenose dolphins (Tursiops sp.) in Shark Bay, Western Australia. Mol Ecol 13(7):1975–1990. https://doi.org/10.1111/j.1365-294X.2004.02192.x
Lambertsen RH (1987) A biopsy system for large whales and its use for cytogenetics. J Mamm 68(2):443–445. https://doi.org/10.2307/1381495
Langergraber KE, Mitani JC, Vigilant L (2007) The limited impact of kinship on cooperation in wild chimpanzees. Proc Natl Acad Sci 104(19):7786–7790. https://doi.org/10.1073/pnas.0611449104
Lee PC, Moss CJ (1995) Statural growth in known-age African elephants (Loxodonta africana). J Zool 236(1):29–41
Lee PC, Moss CJ (1999) The social context for learning and behavioural development among wild African elephants. In: Box HO, Gibson KR (eds) Mammalian social learning: comparative and ecological. Cambridge University Press, Cambridge, pp 102–125
Lukas D, Clutton-Brock T (2014) Costs of mating competition limit male lifetime breeding success in polygynous mammals. Proc R Soc Lond B 281(1786):20140418. https://doi.org/10.1098/rspb.2014.0418
Lynch M (2007) The origins of genome architecture. Sinauer Associates, Sunderland, MA
Majolo B, Lehmann J, de Bortoli VA, Schino G (2012) Fitness-related benefits of dominance in primates. Am J Phys Anthropol 147(4):652–660. https://doi.org/10.1002/ajpa.22031
Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7(5):639–655. https://doi.org/10.1046/j.1365-294x.1998.00374.x
Massen JJM (2017) Friendships in animals. In: Vonk J, Shackelford T (eds) Encyclopedia of animal cognition and behavior. Springer International Publishing, Cham, pp 1–6. https://doi.org/10.1007/978-3-319-47829-6_1899-1
Mayr E (2013) Animal species and evolution. Harvard University Press, Cambridge, MA. https://doi.org/10.4159/harvard.9780674865327
McComb K, Shannon G, Durant SM, Sayialel K, Slotow R, Poole J, Moss C (2011) Leadership in elephants: the adaptive value of age. Proc R Soc Lond B 278(1722):3270–3276. https://doi.org/10.1098/rspb.2011.0168
McDonald DB, Potts WK (1994) Cooperative display and relatedness among males in a lek-mating bird. Science 266(5187):1030–1032. https://doi.org/10.1126/science.7973654
Mesnick SL, Ralls K (2018a) Sexual dimorphism. In: Würsig B, Thewissen JGM, Kovacs KM (eds) Encyclopedia of marine mammals, 3rd edn. Academic Press, London, pp 848–853
Mesnick SL, Ralls K (2018b) Mating systems. In: Würsig B, Thewissen JGM, Kovacs KM (eds) Encyclopedia of marine mammals. Academic Press, London, pp 586–592
Mitani JC, Merriwether DA, Zhang C (2000) Male affiliation, cooperation and kinship in wild chimpanzees. Anim Behav 59(4):885–893
Möller LM, Beheregaray LB, Harcourt RG, Krützen M (2001) Alliance membership and kinship in wild male bottlenose dolphins (Tursiops aduncus) of southeastern Australia. Proc R Soc Lond B 268(1479):1941–1947. https://doi.org/10.1098/rspb.2001.1756
Moore NP, Kelly PF, Cahill JP, Hayden TJ (1995) Mating strategies and mating success of fallow (Dama dama) bucks in a non-lekking population. Behav Ecol Sociobiol 36(2):91–100. https://doi.org/10.1007/bf00170713
Moss CJ, Poole JH (1983) Relationships and social structure of African elephants. In: Hinde RA, Berman CM (eds) Primate social relationships: an integrated approach, vol 315. Sinauer Associates, Sunderland, p 325
Overduin-de Vries AM, Massen JJM, Spruijt BM, Sterck EHM (2012) Sneaky monkeys: an audience effect of male rhesus macaques (Macaca mulatta) on sexual behavior. Am J Primatol 74(3):217–228. https://doi.org/10.1002/ajp.21988
Packer C, Gilbert DA, Pusey AE, O'Brieni SJ (1991) A molecular genetic analysis of kinship and cooperation in African lions. Nature 351(6327):562–565
Parks SE, Brown MW, Conger LA, Hamilton PK, Knowlton AR, Kraus SD, Slay CK, Tyack PL (2007) Occurrence, composition, and potential functions of North Atlantic right whale (Eubalaena glacialis) surface active groups. Mar Mamm Sci 23(4):868–887. https://doi.org/10.1111/j.1748-7692.2007.00154.x
Parsons KM, Durban JW, Claridge DE (2003a) Comparing two alternative methods for sampling small cetaceans for molecular analysis. Mar Mamm Sci 19(1):224–231. https://doi.org/10.1111/j.1748-7692.2003.tb01104.x
Parsons KM, Durban JW, Claridge DE, Balcomb KC, Noble LR, Thompson PM (2003b) Kinship as a basis for alliance formation between male bottlenose dolphins, Tursiops truncatus, in The Bahamas. Anim Behav 66(1):185–194. https://doi.org/10.1006/anbe.2003.2186
Peters KJ, Gerber L, Scheu L, Cicciarella R, Zoller JA, Fei Z, Horvath S, Allen SJ, King SL, Connor RC, Rollins LA, Krützen M (2023) An epigenetic DNA methylation clock for age estimates in Indo-Pacific bottlenose dolphins (Tursiops aduncus ). Evol Appl 16:126–133. https://doi.org/10.1111/eva.13516
Poole JH (1989) Mate guarding, reproductive success and female choice in African elephants. Anim Behav 37:842–849
Poole JH, Moss CJ (1989) Elephant mate searching: group dynamics and vocal and olfactory communication. Symp Zool Soc Lond:111–125
Ramp C, Hagen W, Palsbøll P, Bérubé M, Sears R (2010) Age-related multi-year associations in female humpback whales (Megaptera novaeangliae). Behav Ecol Sociobiol 64(10):1563–1576. https://doi.org/10.1007/s00265-010-0970-8
Rasmussen HB, Okello JBA, Wittemyer G, Siegismund HR, Arctander P, Vollrath F, Douglas-Hamilton I (2007) Age- and tactic-related paternity success in male African elephants. Behav Ecol 19(1):9–15. https://doi.org/10.1093/beheco/arm093
Ruppert KM, Kline RJ, Rahman MS (2019) Past, present, and future perspectives of environmental DNA (eDNA) metabarcoding: a systematic review in methods, monitoring, and applications of global eDNA. Glob Ecol Cons 17:e00547. https://doi.org/10.1016/j.gecco.2019.e00547
Samuels A, Silk JB, Altmann J (1987) Continuity and change in dominance relations among female baboons. Anim Behav 35(3):785–793. https://doi.org/10.1016/S0003-3472(87)80115-X
Schülke O, Bhagavatula J, Vigilant L, Ostner J (2010) Social bonds enhance reproductive success in male macaques. Curr Biol 20(24):2207–2210. https://doi.org/10.1016/j.cub.2010.10.058
Shanley DP, Sear R, Mace R, Kirkwood TBL (2007) Testing evolutionary theories of menopause. Proc R Soc Lond B 274(1628):2943–2949. https://doi.org/10.1098/rspb.2007.1028
Silk JB (2002) Using the ‘F’-word in primatology. Behaviour 139(2–3):421
Silk JB, Beehner JC, Bergman TJ, Crockford C, Engh AL, Moscovice LR, Wittig RM, Seyfarth RM, Cheney DL (2009) The benefits of social capital: close social bonds among female baboons enhance offspring survival. Proc R Soc Lond B 276(1670):3099–3104. https://doi.org/10.1098/rspb.2009.0681
Smith JE (2014) Hamilton’s legacy: kinship, cooperation and social tolerance in mammalian groups. Anim Behav 92:291–304. https://doi.org/10.1016/j.anbehav.2014.02.029
Snyder-Mackler N, Burger JR, Gaydosh L, Belsky DW, Noppert GA, Campos FA, Bartolomucci A, Yang YC, Aiello AE, O’Rand A, Harris KM, Shively CA, Alberts SC, Tung J (2020) Social determinants of health and survival in humans and other animals. Science 368(6493):eaax9553. https://doi.org/10.1126/science.aax9553
Stanton MA, Mann J (2012) Early social networks predict survival in wild bottlenose dolphins. PLoS One 7(10):e47508. https://doi.org/10.1371/journal.pone.0047508
Stern SJ, Friedlaender AS (2018) Migration and movement. In: Würsig B, Thewissen JGM, Kovacs KM (eds) Encyclopedia of marine mammals, 3rd edn. Academic Press, London, pp 602–606
Surbeck M, Mundry R, Hohmann G (2011) Mothers matter! Maternal support, dominance status and mating success in male bonobos (Pan paniscus). Proc R Soc Lond B 278(1705):590–598
Surbeck M, Boesch C, Crockford C, Thompson ME, Furuichi T, Fruth B, Hohmann G, Ishizuka S, Machanda Z, Muller MN, Pusey A, Sakamaki T, Tokuyama N, Walker K, Wrangham R, Wroblewski E, Zuberbühler K, Vigilant L, Langergraber K (2019) Males with a mother living in their group have higher paternity success in bonobos but not chimpanzees. Curr Biol 29(10):R354–R355. https://doi.org/10.1016/j.cub.2019.03.040
Towers JR, Hallé MJ, Symonds HK, Sutton GJ, Morton AB, Spong P, Borrowman JP, Ford JKB (2018) Infanticide in a mammal-eating killer whale population. Sci Rep 8(1):4366. https://doi.org/10.1038/s41598-018-22714-x
Tyack PL (2018) Behavior, overview. In: Würsig B, Thewissen JGM, Kovacs KM (eds) Encyclopedia of marine mammals, 3rd edn. Academic Press, London, pp 86–93
van Noordwijk MA, Arora N, Willems EP, Dunkel LP, Amda RN, Mardianah N, Ackermann C, Krützen M, van Schaik CP (2012) Female philopatry and its social benefits among Bornean orangutans. Behav Ecol Sociobiol 66(6):823–834. https://doi.org/10.1007/s00265-012-1330-7
van Schaik CP, van Hooff JARAM (1994) Male bonds: Afilliative relationships among nonhuman primate males. Behaviour 130(3–4):309–337. https://doi.org/10.1163/156853994X00587
Watts DP (1998) Coalitionary mate guarding by male chimpanzees at Ngogo, Kibale National Park, Uganda. Behav Ecol Sociobiol 44(1):43–55. https://doi.org/10.1007/s002650050513
Weinrich MT, Rosenbaum H, Scott Baker C, Blackmer AL, Whitehead H (2006) The influence of maternal lineages on social affiliations among humpback whales (Megaptera novaeangliae) on their feeding grounds in the southern Gulf of Maine. J Hered 97(3):226–234. https://doi.org/10.1093/jhered/esj018
Weng Z, Yang Y, Wang X, Wu L, Hua S, Zhang H, Meng Z (2021) Parentage analysis in giant grouper ( Epinephelus lanceolatus) using microsatellite and SNP markers from genotyping-by-sequencing data. Genes 12(7):1042
Whitehead H (1996) Babysitting, dive synchrony, and indications of alloparental care in sperm whales. Behav Ecol Sociobiol 38(4):237–244. https://doi.org/10.1007/s002650050238
Whitehead H, Connor R (2005) Alliances I. how large should alliances be? Anim Behav 69(1):117–126. https://doi.org/10.1016/j.anbehav.2004.02.021
Whitehead H, Mann J (2000) Female reproductive strategies of cetaceans. In: Mann J, Connor C, Tyack PL, Whitehead H (eds) Cetacean societies: field studies of dolphins and whales. University of Chicago Press, Chicago, IL, pp 219–246
Whitehead H, Rendell L (2021) The cultural lives of whales and dolphins. University of Chicago Press, Chicago, IL
Whiten A, van de Waal E (2017) Social learning, culture and the ‘socio-cultural brain’ of human and non-human primates. Neurosci Biobehav Rev 82:58–75. https://doi.org/10.1016/j.neubiorev.2016.12.018
Wiszniewski J, Corrigan S, Beheregaray LB, Möller LM (2012) Male reproductive success increases with alliance size in Indo-Pacific bottlenose dolphins (Tursiops aduncus). J Anim Ecol 81(2):423–431. https://doi.org/10.1111/j.1365-2656.2011.01910.x
Würsig B, Rich J, Dara Orbach DN (2023) Sex and behavior. In: Würsig B, Orbach DN (eds) Sex in cetaceans. Springer Nature, Cham
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2023 The Author(s)
About this chapter
Cite this chapter
Gerber, L., Krützen, M. (2023). Genetic Tools to Investigate the Consequences of Sex. In: Würsig, B., Orbach, D.N. (eds) Sex in Cetaceans. Springer, Cham. https://doi.org/10.1007/978-3-031-35651-3_2
Download citation
DOI: https://doi.org/10.1007/978-3-031-35651-3_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-35650-6
Online ISBN: 978-3-031-35651-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)