Abstract
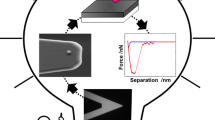
It is generally accepted that the detection limit of bacterial cells using biosensors is one cell. The present chapter describes the specific visualization of bacterial nanofragments using atomic force microscopy (AFM ). The development of new sensitive biosensor methods for the analysis of bacteria is a very important area in biotechnology. AFM is a form of scanning probe microscopy where a cantilever with a very sharp tip is periodically scanned across special surface with immobilized sample. AFM can also be used to measure forces between the tip and sample. The high-resolution imaging of living object is easily possible using AFM. AFM is a powerful, multifunctional imaging platform that allows to visualize and manipulate with biological samples from single molecules to living cells. An artificial nose is unique system for non-contact method for microbial detection. Using this approach, it is possible to detect bacterial “footstep,” even after removing bacteria from the assay medium. These two approaches can be used for detection of less than one bacterial cell.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Abbreviations
- AFM:
-
Atomic force microscopy
- CFU:
-
Colony-forming unit
- LB:
-
Lysogeny broth
- OD:
-
Optical density
- VOCs:
-
Volatile organic components
References
Ahluwalia A, Basta G, Ricci D, Francesconi R, Domenici C, Grattarola M, Palchetti L, Preininger C, De Rossi D (1999) Langmuir-Blodgett films of antibodies as mediators of endothelial cell adhesion on polyurethanes. J Biomater Sci Polym 10(3):295–304
Allison DP, Sullivan CJ, Mortensen NP, Retterer ST, Doktycz M (2011) Bacterial immobilization for imaging by atomic force microscopy. J Vis Exp 10(54):pii: 2880. https://doi.org/10.3791/2880
Andre G, Kulakauskas S, Chapot-Chartier MP, Navet B, Deghorain M, Bernard E, Hols P, Dufrêne YF (2010) Imaging the nanoscale organization of peptidoglycan in living Lactococcus lactis cells. Nat Commun 1:27. https://doi.org/10.1038/ncomms1027
Benn G, Pyne ALB, Ryadnov MG, Hoogenboom BW (2019) Imaging live bacteria at the nanoscale: comparison of immobilisation strategies. Analyst 144(23):6944–6952
Binnig G, Quate CF, Gerber CH (1986) Atomic force microscopy. Phys Rev Lett 56:930–933
Chen P, Xu L, Liu J, Hol FJ, Keymer JE, Taddei F, Han D, Lindner AB (2014) Nanoscale probing the kinetics of oriented bacterial cell growth using atomic force microscopy. Small 10(15):3018–3025
Dubrovin EV, Fedyukina GN, Kraevsky SV, Ignatyuk TE, Yaminsky I, Ignatov SG (2012) AFM specific identification of bacterial cell fragments on biofunctional surfaces. Open Microbiol J 6:22–30
Dufrêne YF, Ando T, Garcia R, Alsteens D, Martinez-Martin D, Engel A, Gerber C, Müller DJ (2017) Imaging modes of atomic force microscopy for application in molecular and cell biology. Nat Nanotechnol 12(4):295–307
Elosua C, Matias IR, Bariain C, Arregui FJ (2006) Volatile Organic Compound Optical Fiber Sensors: A Review. Sensors (Basel) 6(11):1440–1465
Flemming HC, Wingender J, Szewzyk U, Steinberg P, Rice SA, Kjelleberg S (2016) Biofilms: an emergent form of bacterial life. Nat Rev Microbiol 14(9):563–575
Kaundal S, Deep A, Kaur G, Thakur KG (2020) Molecular and Biochemical Characterization of YeeF/YezG, a Polymorphic Toxin-Immunity Protein Pair From Bacillus subtilis. Front Microbiol 11:95. https://doi.org/10.3389/fmicb.2020.00095
Kominami H, Kobayashi K, Ido S, Kimiya H, Yamada H (2018) Immunoactivity of self-assembled antibodies investigated by atomic force microscopy. RSC Adv 8(51):29378–29384
Kommnick C, Lepper A, Hensel M (2019) Correlative light and scanning electron microscopy (CLSEM) for analysis of bacterial infection of polarized epithelial cells. Sci Rep 9:17079
Longo G, Rio LM, Trampuz A, Dietler G, Bizzini A, Kasas S (2013) Antibiotic-induced modifications of the stiffness of bacterial membranes. J Microbiol Methods 93(2):80–84
Meyer RL, Zhou X, Tang L, Arpanaei A, Kingshott P, Besenbacher F (2010) Immobilisation of living bacteria for AFM imaging under physiological conditions. Ultramicroscopy 110(11):1349–1357
Mittelviefhaus M, Müller DB, Zambelli T, Vorholt JA (2019) A modular atomic force microscopy approach reveals a large range of hydrophobic adhesion forces among bacterial members of the leaf microbiota. ISME J (7):1878–1882
Pan F, Altenried S, Liu M, Hegemann D, Bülbül E, Moeller J, Schmahl WW, Maniura-Weber K, Ren Q (2020) A nanolayer coating on polydimethylsiloxane surfaces enables mechanistic study of bacterial adhesion influenced by material surface physicochemistry. Mater Horiz 7:93–103. https://doi.org/10.1039/C9MH01191A
Shi H, Sun J, Han R, Ding C, Hu F, Yu S (2019) The strategy for correcting interference from water in Fourier transform infrared spectrum based bacterial typing. Talanta:120347. https://doi.org/10.1016/j.talanta.2019.120347
Soon RL, Nation RL, Hartley PG, Larson I, Li J (2009) Atomic force microscopy investigation of the morphology and topography of colistin-heteroresistant Acinetobacter baumannii strains as a function of growth phase and in response to colistin treatment. Antimicrob Agents Chemother 53(12):4979–4986
Stitzel SE, Albert KJ, Ignatov SG, David R, Walt DR (2002) Artificial nose employing microsphere sensors for detection of volatile organic compounds. Proc. SPIE 4575, chemical and biological early warning monitoring for water, food, and ground. https://doi.org/10.1117/12.456916.
Tilocca B, Cao A, Migheli Q (2020) Scent of a killer: microbial volatilome and its role in the biological control of plant pathogens. Front Microbiol 11:41. https://doi.org/10.3389/fmicb.2020.00041
Turishchev SY, Marchenko D, Sivakov V, Belikov EA, Chuvenkova OA, Parinova EV, Koyuda D, Chumakov RG, Lebedev AM, Kulikova TV, Berezhnoy AA, Valiakhmedova IV, Praslova NV, Preobrazhenskaya EV, Antipov SS (2019) On the possibility of Photo Emission Electron Microscopy for E. coli advanced studies. Result Phys. https://doi.org/10.1016/j.rinp.2019.102821
Viljoen A, Foster SJ, Fantner GE, Hobbs JK, Dufrêne YF (2020) Scratching the surface: bacterial cell envelopes at the nanoscale. Scratching the surface: bacterial cell envelopes at the nanoscale. MBio 11:1. https://doi.org/10.1128/mBio.03020-19
Voloshin AG, Filippovich SY, Bachurina GP, Besaeva SG, Ignatov SG (2010) Spectrophotometric analysis of volatile compounds in microorganisms. Appl Biochem Microbiol 46(3):303–306. https://doi.org/10.1134/S0003683810030099
Wang H, Koydemir HC, Qiu Y, Bai B, Zhang Y et al (2020) Early-detection and classification of live bacteria using time-lapse coherent imaging and deep learning. arXiv:2001.10695 [physics.ins-det], 2020.
Welch NG, Scoble JA, Muir BW, Pigram PJ (2017) Orientation and characterization of immobilized antibodies for improved immunoassays. Biointerphases 12(2):02D301. https://doi.org/10.1116/1.4978435
Acknowledgments
The work was supported by Rospotrebnadzor. F.S.Y. and G.P.B. were supported by the State Аssignment 0104-2019-0024 to Research Center of Biotechnology RAS. The authors wish to thank E.V. Dubrovin and D.S. Ignatov for the help.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ignatov, S.G., Voloshin, A.G., Bachurina, G.P., Filippovich, S.Y., Dyatlov, I.A. (2021). Is It Possible to Detect Less Than One Bacterial Cell?. In: Rai, M., Reshetilov, A., Plekhanova, Y., Ingle, A.P. (eds) Macro, Micro, and Nano-Biosensors. Springer, Cham. https://doi.org/10.1007/978-3-030-55490-3_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-55490-3_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-55489-7
Online ISBN: 978-3-030-55490-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)