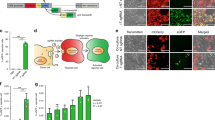
Abstract
Vesicle trafficking entails packaging and transport of membrane-associated proteins to their target membranes, and recycling or degradation of endocytosed proteins. Biochemical and cell biological studies of vesicle trafficking often require the introduction of epitope tags or fluorescent protein markers for protein purification and tracking in cells. Previously, such tagging experiments in mammalian cells mainly used overexpression systems, which could lead to artifacts. Abnormally high expression levels also prevent us from studying individual vesicle trafficking events with precision. With the advent of CRISPR technologies, epitope tags and fluorescent proteins can now be introduced into endogenous proteins in almost any cell type that are proliferating in culture. This chapter describes approaches for inserting tags at the endogenous loci of genes, with the vesicle tethering protein complex, exocyst, as an example.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Rodriguez-Boulan E, Kreitzer G, Müsch A (2005) Organization of vesicular trafficking in epithelia. Nat Rev Mol Cell Biol 6(3):233–247
Orlando K, Guo W (2009) Membrane organization and dynamics in cell polarity. Cold Spring Harb Perspect Biol 1(5):a001321
Booher KR, Kaiser P (2008) A PCR-based strategy to generate yeast strains expressing endogenous levels of amino-terminal epitope-tagged proteins. Biotechnol J 3(4):524–529. https://doi.org/10.1002/biot.200800012
Vavouri T, Semple JI, Garcia-Verdugo R, Lehner B (2009) Intrinsic protein disorder and interaction promiscuity are widely associated with dosage sensitivity. Cell 138(1):198–208. https://doi.org/10.1016/j.cell.2009.04.029
Birchler JA, Veitia RA (2012) Gene balance hypothesis: connecting issues of dosage sensitivity across biological disciplines. Proc Natl Acad Sci U S A 109(37):14746–14753
Doyon JB, Zeitler B, Cheng J et al (2011) Rapid and efficient clathrin-mediated endocytosis revealed in genome-edited mammalian cells. Nat Cell Biol 13(3):331–337. https://doi.org/10.1038/ncb2175
Rivera-Molina F, Toomre D (2013) Live-cell imaging of exocyst links its spatiotemporal dynamics to various stages of vesicle fusion. J Cell Biol 201:673–680. https://doi.org/10.1083/jcb.201212103
Yu J (2016) Single-molecule studies in live cells. Annu Rev Phys Chem 67:565–585. https://doi.org/10.1146/annurev-physchem-040215-112451
Chen B, Zou W, Xu H et al (2018) Efficient labeling and imaging of protein-coding genes in living cells using CRISPR-tag. Nat Commun 9(1):5065. https://doi.org/10.1038/s41467-018-07498-y
Ahmed SM, Nishida-Fukuda H, Li Y et al (2018) Exocyst dynamics during vesicle tethering and fusion. Nat Commun 9:5140. https://doi.org/10.1038/s41467-018-07467-5
Doudna JA, Charpentier E (2014) The new frontier of genome engineering with CRISPR-Cas9. Science 346(6213):1258096
Sander JD, Joung JK (2014) CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 32(4):347–355
Jiang F, Doudna JA (2017) CRISPR-Cas9 structures and mechanisms. Annu Rev Biophys 46:505–529
Jinek M, Chylinski K, Fonfara I et al (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821. https://doi.org/10.1126/science.1225829
Deltcheva E, Chylinski K, Sharma CM et al (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471(7340):602–607. https://doi.org/10.1038/nature09886
Chylinski K, Le Rhun A, Charpentier E (2013) The tracrRNA and Cas9 families of type II CRISPR-Cas immunity systems. RNA Biol 10(5):726–737. https://doi.org/10.4161/rna.24321
Liu M, Rehman S, Tang X et al (2019) Methodologies for improving HDR efficiency. Front Genet 9:691
Rodriguez EA, Campbell RE, Lin JY et al (2017) The growing and glowing toolbox of fluorescent and photoactive proteins. Trends Biochem Sci 42(2):111–129
Erdmann RS, Baguley SW, Richens JH et al (2019) Labeling strategies matter for super-resolution microscopy: a comparison between HaloTags and SNAP-tags. Cell Chem Biol 26(4):584–592.e6. https://doi.org/10.1016/j.chembiol.2019.01.003
England CG, Luo H, Cai W (2015) HaloTag technology: a versatile platform for biomedical applications. Bioconjug Chem 26(6):975–986. https://doi.org/10.1021/acs.bioconjchem.5b00191
Mali P, Yang L, Esvelt KM et al (2013) RNA-guided human genome engineering via Cas9. Science 339(6121):823–826. https://doi.org/10.1126/science.1232033
Jao LE, Wente SR, Chen W (2013) Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc Natl Acad Sci U S A 110(34):13904–13909. https://doi.org/10.1073/pnas.1308335110
Evans PD, Cook SN, Riggs PD, Noren CJ (1995) LITMUS(TM): multipurpose cloning vectors with a novel system for bidirectional in vitro transcription. BioTechniques 19(1):130–135
Roberts B, Haupt A, Tucker A et al (2017) Systematic gene tagging using CRISPR/Cas9 in human stem cells to illuminate cell organization. Mol Biol Cell 28(21):2854–2874. https://doi.org/10.1091/mbc.E17-03-0209
Maruyama T, Dougan SK, Truttmann MC et al (2015) Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat Biotechnol 33(5):538–542. https://doi.org/10.1038/nbt.3190
Chu VT, Weber T, Wefers B et al (2015) Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat Biotechnol 33(5):543–548. https://doi.org/10.1038/nbt.3198
Acknowledgments
I thank Dr. Ian Macara, Dr. Loic Fort, and Christian de Caestecker for their comments and insights on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ahmed, S.M. (2022). Generation of Endogenously Tagged Membrane Trafficking Regulators Using CRISPR Genome Editing. In: Shen, J. (eds) Membrane Trafficking. Methods in Molecular Biology, vol 2473. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2209-4_5
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2209-4_5
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2208-7
Online ISBN: 978-1-0716-2209-4
eBook Packages: Springer Protocols