Abstract
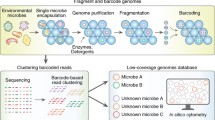
The ability to isolate and cultivate cells in the laboratory has been key to progress in the life sciences. Methods that allow for automated and rapid isolation and cultivation of microorganisms in the laboratory can provide access to organisms that have previously not been propagated in the laboratory, thereby enabling in-depth studies of the physiology of these microbes. Here we describe an automated high-throughput method that combines encapsulation of cells into agarose microcapsules. After isolating individual cells by encapsulation, the population of cells can be incubated as a whole. Cells that divide and form distinct microcolonies within the microcapsule can subsequently be separated from each other by flow cytometry.
The original version of this chapter was revised: Corresponding author name was updated and acknowledgement content was added to the chapter. The erratum to this chapter is available at 10.1007/8623_2016_197
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ishikuri S, Hattori T (1985) Formation of bacterial colonies in successive time intervals. Appl Environ Microbiol 49:870–873
Winding A, Binnerup SJ, Sørensen J (1994) Viability of indigenous soil bacteria assayed by respiratory activity and growth. Appl Environ Microbiol 60:2869–2875
Janssen PH (2008) New cultivation strategies for terrestrial microorganisms. In: Zengler K (ed) Accessing uncultivated microorganisms: from the environment to organisms and genomes and back. ASM Press, Washington, DC, pp 173–192
Watve M, Shejval V, Sonawane C et al (2000) The “K” selected oligotrophic bacteria: a key to uncultured diversity? Curr Sci 78:1535–1542
Button DK, Schut F, Quang P, Martin R, Roberston BR (1993) Viability and isolation of marine bacteria by dilution culture: theory, procedures, and initial results. Appl Environ Microbiol 59:881–891
Schoenborn L, Yates PS, Grinton BE, Hugenholtz P, Janssen PH (2004) Liquid serial dilution is inferior to solid media for isolation of cultures representative of the phylum-level diversity of soil bacteria. Appl Environ Microbiol 70:4363–4366
Connon SA, Giovannoni SJ (2002) High-throughput methods for culturing microorganisms in very-low-nutrient media yield diverse new marine isolates. Appl Environ Microbiol 68:3878–3885
de Man JC (1975) The probability of most probable numbers. Eur J Appl Microbiol 1:67–78
Porter J, Edwards C, Morgan JA, Pickup RW (1993) Rapid, automated separation of specific bacteria from lake water and sewage by flow cytometry and cell sorting. Appl Environ Microbiol 59:3327–3333
Ferrari BC, Oregaard G, Sorensen SJ (2004) Recovery of GFP-labeled bacteria for culturing and molecular analysis after cell sorting using a benchtop flow cytometer. Microb Ecol 48:239–245
Ishii S, Ohno H, Tsuboi M, Otsuka S, Senoo K (2011) Identification and isolation of active N2O reducers in rice paddy soil. ISME J 5:1936–1945
Taylor RJ, Falconnet D, Niemisto A et al (2009) Dynamic analysis of MAPK signaling using a high-throughput microfluidic single-cell imaging platform. Proc Natl Acad Sci U S A 106:3758–3763
Marcy Y, Ouverney C, Bik EM et al (2007) Dissecting biological “dark matter” with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc Natl Acad Sci U S A 104:11889–11894
Pompano RR, Liu W, Du W, Ismagilov RF (2011) Microfluidics using spatially defined arrays of droplets in one, two, and three dimensions. Annu Rev Anal Chem 4:59–81
Wang BL, Ghaderi A, Zhou H et al (2014) Microfluidic high-throughput culturing of single cells for selection based on extracellular metabolite production or consumption. Nat Biotechnol 32:473–478
Fröhlich J, König H (1999) Rapid isolation of single microbial cells from mixed natural and laboratory populations with the aid of a micromanipulator. Syst Appl Microbiol 22:249–257
Ishoy T, Kvist T, Westermann P, Ahring BK (2006) An improved method for single cell isolation of prokaryotes from meso-, thermo- and hyperthermophilic environments using micromanipulation. Appl Microbiol Biotechnol 69:510–514
Zengler K, Walcher M, Clark G et al (2005) High-throughput cultivation of microorganisms using microcapsules. Methods Enzymol 397:124–130
Acknowledgements
I am grateful to Lars Behrendt, Søren Johannes S½rensen, and Jakob Winther (University of Copenhagen, Denmark) for providing results prior to publication that led to further improvements of the method.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this protocol
Cite this protocol
Zengler, K. (2014). Protocols for High-Throughput Isolation and Cultivation. In: McGenity, T., Timmis, K., Nogales , B. (eds) Hydrocarbon and Lipid Microbiology Protocols. Springer Protocols Handbooks. Springer, Berlin, Heidelberg. https://doi.org/10.1007/8623_2014_38
Download citation
DOI: https://doi.org/10.1007/8623_2014_38
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-45178-6
Online ISBN: 978-3-662-45179-3
eBook Packages: Springer Protocols