Abstract
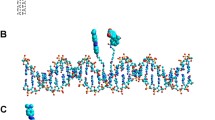
Protein-induced DNA bending plays a key role in many essential biological processes, such as DNA replication, recombination, and transcription, and can be analyzed by a variety of biochemical and biophysical methods, such as electrophoresis mobility shift assay (EMSA), X-ray crystallography, nuclear magnetic resonance (NMR), and DNA ring closure assay. In this chapter, I will provide a detailed protocol for studying protein-induced DNA bending by utilizing the plasmid pBendAT. pBendAT carries a 230 bp DNA segment containing five pairs of restriction–endonuclease recognition sites and can be used to produce a set of five DNA fragments of identical length, with each fragment having a different positioning of a protein-binding site. Binding of a protein to this site will divide a DNA fragment into two DNA segments. If protein binding leads to DNA bending, then the two DNA segments will be at an angle to each other and the distance between the DNA ends will shorten. As a result, the gel mobility of the protein–DNA complex will be affected as the mobility of a rigid DNA fragment is inversely proportional to the end-to-end distance. Therefore, by analyzing how the position of protein binding within a fragment affects the gel mobility of the protein–DNA complexes, we are able to determine the DNA bending angle and the location of the bend. The DNA fragments of identical length can also be conveniently generated by PCR amplification using pBendAT as the DNA template. Since the 230 bp DNA fragment of pBendAT does not contain more than two consecutive AT base pairs, pBendAT is particularly suitable for the assessment of DNA bending induced by proteins recognizing AT-rich DNA sequences cloned in the 230 bp DNA fragment.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
van D V, Verrijzer CP (1993) Bending of DNA by transcription factors. Bioessays 15:25–32
Chen B, Xiao Y, Liu C, Li C, Leng F (2010) DNA linking number change induced by sequence-specific DNA-binding proteins. Nucleic Acids Res 38:3643–3654
Kuznetsov SV, Sugimura S, Vivas P, Crothers DM, Ansari A (2006) Direct observation of DNA bending/unbending kinetics in complex with DNA-bending protein IHF. Proc Natl Acad Sci USA 103:18515–18520
Matthews KS, Nichols JC (1998) Lactose repressor protein: functional properties and structure. Prog Nucleic Acid Res Mol Biol 58:127–164
Lewis M, Chang G, Horton NC, Kercher MA, Pace HC, Schumacher MA, Brennan RG, Lu P (1996) Crystal structure of the lactose operon repressor and its complexes with DNA and inducer. Science 271:1247–1254
Lawson CL, Swigon D, Murakami KS, Darst SA, Berman HM, Ebright RH (2004) Catabolite activator protein: DNA binding and transcription activation. Curr Opin Struct Biol 14:10–20
Harman JG (2001) Allosteric regulation of the cAMP receptor protein. Biochim Biophys Acta 1547:1–17
Schultz SC, Shields GC, Steitz TA (1991) Crystal structure of a CAP–DNA complex: the DNA is bent by 90 degrees. Science 253:1001–1007
Kim J, Zwieb C, Wu C, Adhya S (1989) Bending of DNA by gene-regulatory proteins: construction and use of a DNA bending vector. Gene 85:15–23
Wu HM, Crothers DM (1984) The locus of sequence-directed and protein-induced DNA bending. Nature 308:509–513
Crothers DM, Gartenberg MR, Shrader TE (1991) DNA bending in protein–DNA complexes. Methods Enzymol 208:118–146
Thompson JF, Landy A (1988) Empirical estimation of protein-induced DNA bending angles: applications to lambda site-specific recombination complexes. Nucleic Acids Res 16:9687–9705
Zwieb C, Adhya S (2009) Plasmid vectors for the analysis of protein-induced DNA bending. Methods Mol Biol 543:547–562
Chen B, Young J, Leng F (2010) DNA bending by the mammalian high-mobility group protein AT hook 2. Biochemistry 49:1590–1595
Sambrook J, Maniatis T, Fritsch EF (1989) Molecular cloning a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, New York
About this protocol
Cite this protocol
Leng, F. (2013). DNA Bending by Proteins: Utilizing Plasmid pBendAT as a Tool. In: Makovets, S. (eds) DNA Electrophoresis. Methods in Molecular Biology, vol 1054. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-565-1_18
Download citation
DOI: https://doi.org/10.1007/978-1-62703-565-1_18
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-564-4
Online ISBN: 978-1-62703-565-1
eBook Packages: Springer Protocols