Abstract
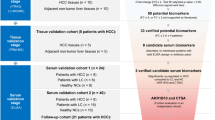
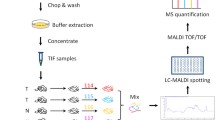
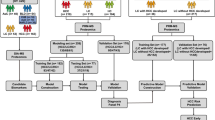
The recent advancements in proteomic technologies have reconstituted our research strategies over different type of liver diseases including hepatocellular carcinoma (HCC). Combined analyses on HCC proteome and clinicopathological data of patients have allowed identification of many promising biomarkers that can be further developed into noninvasive diagnostic assays for cancer surveillance. Capitalizing our established proteomic platform primarily based on two-dimensional polyacrylamide gel electrophoresis (2DE) and MALDI-TOF/TOF mass spectrometry, our groups have identified lamin B1 (LMNB1) and vimentin (VIM) as promising biomarkers for detection of early HCC. Protein levels of both biomarkers were significantly elevated in cancerous tissues when compared to the controls in disease-free and cirrhotic liver subjects. Further investigation of the circulating LMNB1 mRNA level in patients’ blood samples by standard PCR showed 76% sensitivity and 82% specificity for detection of early HCC. In parallel, an ELISA assay for measuring circulating vimentin level in patients’ serum samples could detect small HCC at 40.91% sensitivity and 87.5% specificity. The candidate biomarkers were evaluated with the diagnostic performance of α-fetoprotein (AFP) for HCC. In this article, we address the current protocols for HCC biomarker discovery, ranging from clinical sample preparation, 2DE proteomic profiling and informatics analysis, and assay development and clinical validation study. Focus is emphasized on the methods for sample preservation and low-abundance protein enrichment.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Hao K, Luk JM, Lee NP, Mao M, Zhang C, Ferguson MD, Lamb J, Dai H, Ng IO, Sham PC, Poon RT (2009) Predicting prognosis in hepatocellular carcinoma after curative surgery with common clinicopathologic parameters. BMC Cancer 9:389
Spangenberg HC, Thimme R, Blum HE (2009) Targeted therapy for hepatocellular carcinoma. Nat Rev Gastroenterol Hepatol 6:423–432
Hoshida Y, Villanueva A, Kobayashi M, Peix J, Chiang DY, Camargo A, Gupta S, Moore J, Wrobel MJ, Lerner J, Reich M, Chan JA, Glickman JN, Ikeda K, Hashimoto M, Watanabe G, Daidone MG, Roayaie S, Schwartz M, Thung S, Salvesen HB, Gabriel S, Mazzaferro V, Bruix J, Friedman SL, Kumada H, Llovet JM, Golub TR (2008) Gene expression in fixed tissues and outcome in hepatocellular carcinoma. N Engl J Med 359:1995–2004
Laurent-Puig P, Legoix P, Bluteau O, Belghiti J, Franco D, Binot F, Monges G, Thomas G, Bioulac-Sage P, Zucman-Rossi J (2001) Genetic alterations associated with hepatocellular carcinomas define distinct pathways of hepatocarcinogenesis. Gastroenterology 120:1763–1773
Kaposi-Novak P, Libbrecht L, Woo HG, Lee Y-H, Sears NC, Coulouarn C, Conner EA, Factor VM, Roskams T, Thorgeirsson SS (2009) Central role of c-Myc during malignant conversion in human hepatocarcinogenesis. Cancer Res 69:2775–2782
Wang SM, Ooi LLPJ, Hui KM (2007) Identification and validation of a novel gene signature associated with the recurrence of human hepatocellular carcinoma. Clin Cancer Res 13:6275–6283
Burchard J, Zhang C, Liu AM, Poon RT, Lee NP, Wong KF, Sham PC, Lam BY, Ferguson MD, Tokiwa G, Smith R, Leeson B, Beard R, Lamb JR, Lim L, Mao M, Dai H, Luk JM (2010) microRNA-122 as a regulator of mitochondrial metabolic gene network in hepatocellular carcinoma. Mol Syst Biol 6:402
Sun S, Yi X, Poon RT, Yeung C, Day PJ, Luk JM (2009) A protein-based set of reference markers for liver tissues and hepatocellular carcinoma. BMC Cancer 9:309
Sun S, Day PJ, Lee NP, Luk JM (2009) Biomarkers for early detection of liver cancer: focus on clinical evaluation. Protein Pept Lett 16:473–478
Sun S, Lee NP, Poon RT, Fan ST, He QY, Lau GK, Luk JM (2007) Oncoproteomics of hepatocellular carcinoma: from cancer markers’ discovery to functional pathways. Liver Int 27:1021–1038
Luk JM, Lam BY, Lee NP, Ho DW, Sham PC, Chen L, Peng J, Leng X, Day PJ, Fan ST (2007) Artificial neural networks and decision tree model analysis of liver cancer proteomes. Biochem Biophys Res Commun 361:68–73
Luk JM, Su YC, Lam SC, Lee CK, Hu MY, He QY, Lau GK, Wong FW, Fan ST (2005) Proteomic identification of Ku70/Ku80 autoantigen recognized by monoclonal antibody against hepatocellular carcinoma. Proteomics 5:1980–1986
Lee NP, Leung KW, Cheung N, Lam BY, Xu MZ, Sham PC, Lau GK, Poon RT, Fan ST, Luk JM (2008) Comparative proteomic analysis of mouse livers from embryo to adult reveals an association with progression of hepatocellular carcinoma. Proteomics 8:2136–2149
Lee NP, Chen L, Lin MC, Tsang FH, Yeung C, Poon RT, Peng J, Leng X, Beretta L, Sun S, Day PJ, Luk JM (2009) Proteomic expression signature distinguishes cancerous and nonmalignant tissues in hepatocellular carcinoma. J Proteome Res 8:1293–1303
Luk JM, Lam CT, Siu AF, Lam BY, Ng IO, Hu MY, Che CM, Fan ST (2006) Proteomic profiling of hepatocellular carcinoma in Chinese cohort reveals heat-shock proteins (Hsp27, Hsp70, GRP78) up-regulation and their associated prognostic values. Proteomics 6:1049–1057
Yi X, Luk JM, Lee NP, Peng J, Leng X, Guan XY, Lau GK, Beretta L, Fan ST (2008) Association of mortalin (HSPA9) with liver cancer metastasis and prediction for early tumor recurrence. Mol Cell Proteomics 7:315–325
Lu WJ, Lee NP, Fatima S, Luk JM (2009) Heat shock proteins in cancer: signaling pathways, tumor markers and molecular targets in liver malignancy. Protein Pept Lett 16:508–516
Lu WJ, Lee NP, Kaul SC, Lan F, Poon RT, Wadhwa R, Luk JM (2011) Induction of mutant p53-dependent apoptosis in human hepatocellular carcinoma by targeting stress protein mortalin. Int J Cancer 129: 1806–1814.
Lu WJ, Lee NP, Kaul SC, Lan F, Poon RT, Wadhwa R, Luk JM (2011) Mortalin-p53 interaction in cancer cells is stress dependent and constitutes a selective target for cancer therapy. Cell Death Differ 18:1046–1056
Chen L, Ho DW, Lee NP, Sun S, Lam B, Wong KF, Yi X, Lau GK, Ng EW, Poon TC, Lai PB, Cai Z, Peng J, Leng X, Poon RT, Luk JM (2010) Enhanced detection of early hepatocellular carcinoma by serum SELDI-TOF proteomic signature combined with alpha-fetoprotein marker. Ann Surg Oncol 17:2518–2525
Fukuda S, Itamoto T, Nakahara H, Kohashi T, Ohdan H, Hino H, Ochi M, Tashiro H, Asahara T (2005) Clinicopathologic features and prognostic factors of resected solitary small-sized hepatocellular carcinoma. Hepatogastroenterology 52:1163–1167
Kikuchi LO, Paranagua-Vezozzo DC, Chagas AL, Mello ES, Alves VA, Farias AQ, Pietrobon R, Carrilho FJ (2009) Nodules less than 20 mm and vascular invasion are predictors of survival in small hepatocellular carcinoma. J Clin Gastroenterol 43:191–195
Lok AS, Sterling RK, Everhart JE, Wright EC, Hoefs JC, Di Bisceglie AM, Morgan TR, Kim HY, Lee WM, Bonkovsky HL, Dienstag JL (2010) Des-gamma-carboxy prothrombin and alpha-fetoprotein as biomarkers for the early detection of hepatocellular carcinoma. Gastroenterology 138:493–502
Kim MJ (2011) Current limitations and potential breakthroughs for the early diagnosis of hepatocellular carcinoma. Gut Liver 5:15–21
Sun S, Poon RT, Lee NP, Yeung C, Chan KL, Ng IO, Day PJ, Luk JM (2010) Proteomics of hepatocellular carcinoma: serum vimentin as a surrogate marker for small tumors (<or =2 cm). J Proteome Res 9:1923–1930
Sun S, Xu MZ, Poon RT, Day PJ, Luk JM (2010) Circulating lamin B1 (LMNB1) biomarker detects early stages of liver cancer in patients. J Proteome Res 9:70–78
Hu MY, Lam CT, Liu KD, Xu Z, Fatima S, Su YC, Tsang F, Chen J, Pang JZ, Qin LX, Luk JM (2009) Proteomic identification of a monoclonal antibody recognizing caveolin-1 in hepatocellular carcinoma with metastatic potential. Protein Pept Lett 16:479–485
Liu LX, Lee NP, Chan VW, Xue W, Zender L, Zhang C, Mao M, Dai H, Wang XL, Xu MZ, Lee TK, Ng IO, Chen Y, Kung HF, Lowe SW, Poon RT, Wang JH, Luk JM (2009) Targeting cadherin-17 inactivates Wnt signaling and inhibits tumor growth in liver carcinoma. Hepatology 50:1453–1463
Luk JM, Wong KF (2006) Monoclonal antibodies as targeting and therapeutic agents: prospects for liver transplantation, hepatitis and hepatocellular carcinoma. Clin Exp Pharmacol Physiol 33:482–488
Tsang FH, Lee NP, Luk JM (2009) The use of small peptides in the diagnosis and treatment of hepatocellular carcinoma. Protein Pept Lett 16:530–538
Speers AE, Wu CC (2007) Proteomics of integral membrane proteins – theory and application. Chem Rev 107:3687–3714
Josic D, Clifton JG (2007) Mammalian plasma membrane proteomics. Proteomics 7:3010–3029
Rabilloud T, Adessi C, Giraudel A, Lunardi J (1997) Improvement of the solubilization of proteins in two-dimensional electrophoresis with immobilized pH gradients. Electrophoresis 18:307–316
James GT (1978) Inactivation of the protease inhibitor phenylmethylsulfonyl fluoride in buffers. Anal Biochem 86:574–579
Xu MZ, Chan SW, Liu AM, Wong KF, Fan ST, Chen J, Poon RT, Zender L, Lowe SW, Hong W, Luk JM (2010) AXL receptor kinase is a mediator of YAP-dependent oncogenic functions in hepatocellular carcinoma. Oncogene 30:1229–1240
Wong KF, Luk JM, Cheng RH, Klickstein LB, Fan ST (2007) Characterization of two novel LPS-binding sites in leukocyte integrin betaA domain. FASEB J 21:3231–3239
Li C, Hong Y, Tan YX, Zhou H, Ai JH, Li SJ, Zhang L, Xia QC, Wu JR, Wang HY, Zeng R (2004) Accurate qualitative and quantitative proteomic analysis of clinical hepatocellular carcinoma using laser capture microdissection coupled with isotope-coded affinity tag and two-dimensional liquid chromatography mass spectrometry. Mol Cell Proteomics 3:399–409
Ai J, Tan Y, Ying W, Hong Y, Liu S, Wu M, Qian X, Wang H (2006) Proteome analysis of hepatocellular carcinoma by laser capture microdissection. Proteomics 6:538–546
Guedj N, Dargere D, Degos F, Janneau JL, Vidaud D, Belghiti J, Bedossa P, Paradis V (2006) Global proteomic analysis of microdissected cirrhotic nodules reveals significant biomarkers associated with clonal expansion. Lab Invest 86:951–958
Chan JK, Thompson JW, Gill TA (1995) Quantitative determination of protamines by coomassie blue G assay. Anal Biochem 226:191–193
Nassiri M, Ramos S, Zohourian H, Vincek V, Morales AR, Nadji M (2008) Preservation of biomolecules in breast cancer tissue by a formalin-free histology system. BMC Clin Pathol 8:1
Espina V, Edmiston KH, Heiby M, Pierobon M, Sciro M, Merritt B, Banks S, Deng J, VanMeter AJ, Geho DH, Pastore L, Sennesh J, Petricoin EF III, Liotta LA (2008) A portrait of tissue phosphoprotein stability in the clinical tissue procurement process. Mol Cell Proteomics 7:1998–2018
Rountree CB, Van Kirk CA, You H, Ding W, Dang H, Vanguilder HD, Freeman WM (2010) Clinical application for the preservation of phospho-proteins through in-situ tissue stabilization. Proteome Sci 8:61
Svensson M, Boren M, Skold K, Falth M, Sjogren B, Andersson M, Svenningsson P, Andren PE (2009) Heat stabilization of the tissue proteome: a new technology for improved proteomics. J Proteome Res 8:974–981
Goodwin RJ, Lang AM, Allingham H, Boren M, Pitt AR (2010) Stopping the clock on proteomic degradation by heat treatment at the point of tissue excision. Proteomics 10:1751–1761
Robinson AA, Westbrook JA, English JA, Boren M, Dunn MJ (2009) Assessing the use of thermal treatment to preserve the intact proteomes of post-mortem heart and brain tissue. Proteomics 9:4433–4444
Scholz B, Skold K, Kultima K, Fernandez C, Waldemarson S, Savitski MM, Svensson M, Boren M, Stella R, Andren PE, Zubarev R, James P (2011) Impact of temperature dependent sampling procedures in proteomics and peptidomics – a characterization of the liver and pancreas post mortem degradome. Mol Cell Proteomics 10:M900229MCP200
Rossbach U, Nilsson A, Falth M, Kultima K, Zhou Q, Hallberg M, Gordh T, Andren PE, Nyberg F (2009) A quantitative peptidomic analysis of peptides related to the endogenous opioid and tachykinin systems in nucleus accumbens of rats following naloxone-precipitated morphine withdrawal. J Proteome Res 8:1091–1098
Qian WJ, Jacobs JM, Liu T, Camp DG II, Smith RD (2006) Advances and challenges in liquid chromatography–mass spectrometry-based proteomics profiling for clinical applications. Mol Cell Proteomics 5:1727–1744
Rifai N, Gillette MA, Carr SA (2006) Protein biomarker discovery and validation: the long and uncertain path to clinical utility. Nat Biotechnol 24:971–983
Anderson NL, Anderson NG (2002) The human plasma proteome: history, character, and diagnostic prospects. Mol Cell Proteomics 1:845–867
Pieper R, Su Q, Gatlin CL, Huang ST, Anderson NL, Steiner S (2003) Multi-component immunoaffinity subtraction chromatography: an innovative step towards a comprehensive survey of the human plasma proteome. Proteomics 3:422–432
Brand J, Haslberger T, Zolg W, Pestlin G, Palme S (2006) Depletion efficiency and recovery of trace markers from a multiparameter immunodepletion column. Proteomics 6:3236–3242
Qian WJ, Kaleta DT, Petritis BO, Jiang H, Liu T, Zhang X, Mottaz HM, Varnum SM, Camp DG II, Huang L, Fang X, Zhang WW, Smith RD (2008) Enhanced detection of low abundance human plasma proteins using a tandem IgY12-SuperMix immunoaffinity separation strategy. Mol Cell Proteomics 7:1963–1973
Zhang H, Li XJ, Martin DB, Aebersold R (2003) Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat Biotechnol 21:660–666
Liu T, Qian WJ, Strittmatter EF, Camp DG II, Anderson GA, Thrall BD, Smith RD (2004) High-throughput comparative proteome analysis using a quantitative cysteinyl-peptide enrichment technology. Anal Chem 76:5345–5353
Baumann S, Ceglarek U, Fiedler GM, Lembcke J, Leichtle A, Thiery J (2005) Standardized approach to proteome profiling of human serum based on magnetic bead separation and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Clin Chem 51:973–980
Zhang L, Xie J, Wang X, Liu X, Tang X, Cao R, Hu W, Nie S, Fan C, Liang S (2005) Proteomic analysis of mouse liver plasma membrane: use of differential extraction to enrich hydrophobic membrane proteins. Proteomics 5:4510–4524
Everberg H, Peterson R, Rak S, Tjerneld F, Emanuelsson C (2006) Aqueous two-phase partitioning for proteomic monitoring of cell surface biomarkers in human peripheral blood mononuclear cells. J Proteome Res 5:1168–1175
Kohnke PL, Mulligan SP, Christopherson RI (2009) Membrane proteomics for leukemia classification and drug target identification. Curr Opin Mol Ther 11:603–610
Rahbar AM, Fenselau C (2004) Integration of Jacobson’s pellicle method into proteomic strategies for plasma membrane proteins. J Proteome Res 3:1267–1277
Zhang LJ, Wang XE, Peng X, Wei YJ, Cao R, Liu Z, Xiong JX, Yin XF, Ping C, Liang S (2006) Proteomic analysis of low-abundant integral plasma membrane proteins based on gels. Cell Mol Life Sci 63:1790–1804
Macher BA, Yen TY (2007) Proteins at membrane surfaces-a review of approaches. Mol Biosyst 3:705–713
Stasyk T, Huber LA (2004) Zooming in: fractionation strategies in proteomics. Proteomics 4:3704–3716
Tang X, Yi W, Munske GR, Adhikari DP, Zakharova NL, Bruce JE (2007) Profiling the membrane proteome of Shewanella oneidensis MR-1 with new affinity labeling probes. J Proteome Res 6:724–734
Scheurer SB, Rybak JN, Roesli C, Brunisholz RA, Potthast F, Schlapbach R, Neri D, Elia G (2005) Identification and relative quantification of membrane proteins by surface biotinylation and two-dimensional peptide mapping. Proteomics 5:2718–2728
Rybak JN, Ettorre A, Kaissling B, Giavazzi R, Neri D, Elia G (2005) In vivo protein biotinylation for identification of organ-specific antigens accessible from the vasculature. Nat Methods 2:291–298
Lu B, McClatchy DB, Kim JY, Yates JR III (2008) Strategies for shotgun identification of integral membrane proteins by tandem mass spectrometry. Proteomics 8:3947–3955
Na K, Lee EY, Lee HJ, Kim KY, Lee H, Jeong SK, Jeong AS, Cho SY, Kim SA, Song SY, Kim KS, Cho SW, Kim H, Paik YK (2009) Human plasma carboxylesterase 1, a novel serologic biomarker candidate for hepatocellular carcinoma. Proteomics 9:3989–3999
Chaerkady R, Thuluvath PJ, Kim MS, Nalli A, Vivekanandan P, Simmers J, Torbenson M, Pandey A (2008) O Labeling for a quantitative proteomic analysis of glycoproteins in hepatocellular carcinoma. Clin Proteomics 4:137–155
Dai Z, Fan J, Liu Y, Zhou J, Bai D, Tan C, Guo K, Zhang Y, Zhao Y, Yang P (2007) Identification and analysis of alpha1,6-fucosylated proteins in human normal liver tissues by a target glycoproteomic approach. Electrophoresis 28:4382–4391
Xu Z, Zhou X, Lu H, Wu N, Zhao H, Zhang L, Zhang W, Liang YL, Wang L, Liu Y, Yang P, Zha X (2007) Comparative glycoproteomics based on lectins affinity capture of N-linked glycoproteins from human Chang liver cells and MHCC97-H cells. Proteomics 7:2358–2370
Wollscheid B, Bausch-Fluck D, Henderson C, O’Brien R, Bibel M, Schiess R, Aebersold R, Watts JD (2009) Mass-spectrometric identification and relative quantification of N-linked cell surface glycoproteins. Nat Biotechnol 27:378–386
Cottingham K (2008) Antibodypedia seeks to answer the question: “how good is that antibody?”. J Proteome Res 7:4213
Bjorling E, Uhlen M (2008) Antibodypedia, a portal for sharing antibody and antigen validation data. Mol Cell Proteomics 7:2028–2037
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
Peterson GL (1979) Review of the Folin phenol protein quantitation method of Lowry, Rosebrough, Farr and Randall. Anal Biochem 100:201–220
Stoscheck CM (1990) Quantitation of protein. Methods Enzymol 182:50–68
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer Science+Business Media New York
About this protocol
Cite this protocol
Wong, KF., Luk, J.M. (2012). Discovery of Lamin B1 and Vimentin as Circulating Biomarkers for Early Hepatocellular Carcinoma. In: Josic, D., Hixson, D. (eds) Liver Proteomics. Methods in Molecular Biology, vol 909. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-61779-959-4_19
Download citation
DOI: https://doi.org/10.1007/978-1-61779-959-4_19
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-61779-958-7
Online ISBN: 978-1-61779-959-4
eBook Packages: Springer Protocols