Abstract
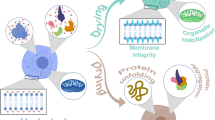
Intrinsically disordered proteins (IDPs) describe a group of proteins that do not have a regular tertiary structure and typically have very little ordered secondary structure. Despite not following the biochemical dogma of “structure determines function” and “function determines structure,” IDPs have been identified as having numerous biological functions. We describe here the steps to express and purify the intrinsically disordered stress response protein, Late embryogenesis abundant protein 3-2 from Arabidopsis thaliana (AtLEA 3-2), with 15N and 13C isotopes in E. coli, although the protocol can be adapted for any IDP with or without isotopic labeling. The atlea 3-2 gene has been cloned into the pET-SUMO vector that in addition to the SUMO portion encodes an N-terminal hexahistidine sequence (His-tag). This vector allows for the SUMO-AtLEA 3-2 fusion protein to be purified using Ni-affinity chromatography and, through the use of ubiquitin-like-specific protease 1 (Ulp1, a SUMO protease), results in an AtLEA 3-2 with a native N-terminus. We also describe the expression and purification of Ulp1 itself.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Tompa P (2002) Intrinsically unstructured proteins. Trends Biochem Sci 27:527–533
Uversky VN (2013) A decade and a half of protein intrinsic disorder: biology still waits for physics. Protein Sci 22:693–724. https://doi.org/10.1002/pro.2261
Liu J, Perumal NB, Oldfield CJ et al (2006) Intrinsic disorder in transcription factors. Biochemistry 45:6873–6888. https://doi.org/10.1021/bi0602718
Iakoucheva LM, Brown CJ, Lawson JD et al (2002) Intrinsic disorder in cell-signaling and cancer-associated proteins. J Mol Biol 323:573–584. https://doi.org/10.1016/S0022-2836(02)00969-5
Sun X, Rikkerink EHA, Jones WT, Uversky VN (2013) Multifarious roles of intrinsic disorder in proteins illustrate its broad impact on plant biology. Plant Cell 25:38–55. https://doi.org/10.1105/tpc.112.106062
Graether SP, Boddington KF (2014) Disorder and function: a review of the dehydrin protein family. Front Plant Sci 5:e576. https://doi.org/10.3389/fpls.2014.00576
Oldfield CJ, Cheng Y, Cortese MS et al (2005) Comparing and combining predictors of mostly disordered proteins. Biochemistry 44:1989–2000. https://doi.org/10.1021/bi047993o
Varadi M, Kosol S, Lebrun P et al (2014) pE-DB: a database of structural ensembles of intrinsically disordered and of unfolded proteins. Nucleic Acids Res 42:D326–D335. https://doi.org/10.1093/nar/gkt960
Gellissen G (2006) Production of recombinant proteins: novel microbial and eukaryotic expression systems. John Wiley & Sons, Hoboken, New Jersey
Yanaka S, Yagi H, Yogo R et al (2018) Stable isotope labeling approaches for NMR characterization of glycoproteins using eukaryotic expression systems. J Biomol NMR 71:193–202. https://doi.org/10.1007/s10858-018-0169-2
Baneyx F (1999) Recombinant protein expression in Escherichia coli. Curr Opin Biotechnol 10:411–421. https://doi.org/10.1016/S0958-1669(99)00003-8
Graether SP (2019) Troubleshooting guide to expressing intrinsically disordered proteins for use in NMR experiments. Front Mol Biosci 5:49. https://doi.org/10.3389/fmolb.2018.00118
Reverter D, Lima CD (2009) Preparation of SUMO proteases and kinetic analysis using endogenous substrates. Methods Mol Biol 497:225–239. https://doi.org/10.1007/978-1-59745-566-4_15
Na J-H, Lee W-K, Yu YG (2018) How do we study the dynamic structure of unstructured proteins: a case study on Nopp140 as an example of a large, intrinsically disordered protein. Int J Mol Sci 19:381. https://doi.org/10.3390/ijms19020381
Breindel L, Burz DS, Shekhtman A (2018) Interaction proteomics by using in-cell NMR spectroscopy. J Proteome 191:202–211. https://doi.org/10.1016/j.jprot.2018.02.006
Marley J, Lu M, Bracken C (2001) A method for efficient isotopic labeling of recombinant proteins. J Biomol NMR 20:71–75. https://doi.org/10.1023/A:1011254402785
Dhamole PB, Mahajan P, Feng H (2010) Sugaring out: a new method for removal of acetonitrile from preparative RP-HPLC eluent for protein purification. Process Biochem 45:1672–1676. https://doi.org/10.1016/j.procbio.2010.06.020
Kalthoff C (2003) A novel strategy for the purification of recombinantly expressed unstructured protein domains. J Chromatogr B 786:247–254. https://doi.org/10.1016/S1570-0232(02)00908-X
Livernois AM, Hnatchuk DJ, Findlater EE, Graether SP (2009) Obtaining highly purified intrinsically disordered protein by boiling lysis and single step ion exchange. Anal Biochem 392:70–76. https://doi.org/10.1016/j.ab.2009.05.023
Gallagher SR (2012), One‐dimensional SDS gel electrophoresis of proteins. Curr Protoc Prot Sci 68:10.1.1–10.1.44 https://doi.org/10.1002/0471140864.ps1001s68
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Singh, K.K., Graether, S.P. (2020). Expression and Purification of an Intrinsically Disordered Protein. In: Kragelund, B.B., Skriver, K. (eds) Intrinsically Disordered Proteins. Methods in Molecular Biology, vol 2141. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0524-0_8
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0524-0_8
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0523-3
Online ISBN: 978-1-0716-0524-0
eBook Packages: Springer Protocols