Abstract
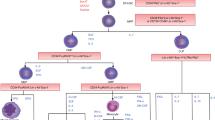
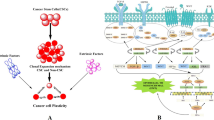
One of the important questions when studying established cancer cell lines is whether such cells contain a subpopulation of primitive cancer stem cells that maintains the expansion of the cell line. To address this issue, we performed studies on the established human embryonal carcinoma cell line NTera2 by evaluating the potential stemness of cells sorted according to their expression of the cell surface stem cell markers CD133 and SSEA4. By performing in vitro and in vivo assays, we observed different properties of cells expressing both, one, or neither of these antigens. While sorted SSEA4+ subpopulations exhibited the greatest propensity for migration toward normal serum and the highest seeding efficiency in the lungs of immunodeficient mice, CD133−SSEA4− cells displayed high seeding efficiency to the bone marrow after injection in vivo. It is worth noting that these properties did not depend on the size of the evaluated cells. To address the question of whether cancer stem cell phenotypes in cell lines are fixed or fluctuating, we sorted single cells according to their expression of CD133 and SSEA4 antigens and observed that cells which did not express these cancer stem cell markers gave rise to cells that express these markers after expansion in vitro. Therefore, our results support the idea that within established cancer cell lines, the phenotype of the cell subpopulation expressing cancer stem cell markers is not fixed but fluctuates during cell line expansion, and cells negative for these markers may acquire their expression.




Similar content being viewed by others
References
Mizrak, D., Brittan, M., & Alison, M. (2008). CD133: Molecule of the moment. The Journal of Pathology, 214(1), 3–9.
Field, M., et al. (2010). Embryonic stem cell markers distinguishing cancer stem cells from normal human neuronal stem cell populations in malignant glioma patients. Clinical Neurosurgery, 57, 151–159.
Grasso, C., et al. (2016). Iterative sorting reveals CD133+ and CD133- melanoma cells as phenotypically distinct populations. BMC Cancer, 16(1), 726.
Kucia, M., et al. (2007). Morphological and molecular characterization of novel population of CXCR4+ SSEA-4+ Oct-4+ very small embryonic-like cells purified from human cord blood: Preliminary report. Leukemia, 21(2), 297–303.
Gang, E. J., et al. (2007). SSEA-4 identifies mesenchymal stem cells from bone marrow. Blood, 109(4), 1743–1751.
Lou, Y. W., et al. (2014). Stage-specific embryonic antigen-4 as a potential therapeutic target in glioblastoma multiforme and other cancers. Proceedings of the National Academy of Sciences of the United States of America, 111(7), 2482–2487.
Gunjal, P., et al., (2015) An emerging question about putative cancer stem cells in established cell lines-are they true stem cells or a fluctuating cell phenotype? Journal of Cancer Stem Cell Research. 3.
Chadalavada, R. S., et al. (2007). Constitutive gene expression predisposes morphogen-mediated cell fate responses of NT2/D1 and 27X-1 human embryonal carcinoma cells. Stem Cells, 25(3), 771–778.
Malecki, M., et al. (2013). TRA-1-60+, SSEA-4+, POU5F1+, SOX2+, NANOG+ clones of pluripotent stem cells in the embryonal carcinomas of the testes. Journal of Stem Cell Research and Therapy, 3(1). doi:10.4172/2157-7633.1000134.
Park, E. K., et al. (2015). Transcriptional repression of cancer stem cell marker CD133 by tumor suppressor p53. Cell Death & Disease, 6, e1964.
Immervoll, H., et al. (2008). Expression of the "stem cell marker" CD133 in pancreas and pancreatic ductal adenocarcinomas. BMC Cancer, 8, 48.
Suzuki, S., et al. (2010). Identification and characterization of cancer stem cells in ovarian yolk sac tumors. Cancer Science, 101(10), 2179–2185.
Nomura, A., et al. (2015). CD133 initiates tumors, induces epithelial-mesenchymal transition and increases metastasis in pancreatic cancer. Oncotarget, 6(10), 8313–8322.
Sansone, P., et al. (2016). Self-renewal of CD133(hi) cells by IL6/Notch3 signalling regulates endocrine resistance in metastatic breast cancer. Nature Communications, 7, 10442.
Wenk, J., et al. (1994). Glycolipids of germ cell tumors: Extended globo-series glycolipids are a hallmark of human embryonal carcinoma cells. International Journal of Cancer, 58(1), 108–115.
Schwartz, C. M., et al. (2005). NTera2: A model system to study dopaminergic differentiation of human embryonic stem cells. Stem Cells and Development, 14(5), 517–534.
Aloia, A., et al. (2015). The sialyl-glycolipid stage-specific embryonic antigen 4 marks a subpopulation of chemotherapy-resistant breast cancer cells with mesenchymal features. Breast Cancer Research, 17(1), 146.
Sivasubramaniyan, K., et al. (2015). Expression of stage-specific embryonic antigen-4 (SSEA-4) defines spontaneous loss of epithelial phenotype in human solid tumor cells. Glycobiology, 25(8), 902–917.
Reuter, V. E. (2005). Origins and molecular biology of testicular germ cell tumors. Modern Pathology, 18(Suppl 2), S51–S60.
Saitou, M., & Yamaji, M. (2012). Primordial germ cells in mice. Cold Spring Harbor Perspectives in Biology, 4, a008375.
Schneider, G., et al. (2014). The paternally imprinted DLK1-GTL2 locus is differentially methylated in embryonal and alveolar rhabdomyosarcomas. International Journal of Oncology, 44(1), 295–300.
Watanabe, K., et al. (2010). Cripto-1 is a cell surface marker for a tumorigenic, undifferentiated subpopulation in human embryonal carcinoma cells. Stem Cells, 28(8), 1303–1314.
Azevedo, A. S., et al. (2015). Metastasis of circulating tumor cells: Favorable soil or suitable biomechanics, or both? Cell Adhesion & Migration, 9(5), 345–356.
Cameron, M. D., et al. (2000). Temporal progression of metastasis in lung: Cell survival, dormancy, and location dependence of metastatic inefficiency. Cancer Research, 60(9), 2541–2546.
Zhan, Q., Wang, C., & Ngai, S. (2013). Ovarian cancer stem cells: A new target for cancer therapy. BioMed Research International, 2013, 916819.
Tesfaigzi, J., & Carlson, D. M. (1996). Cell cycle-specific expression of G(0)SPR1 in Chinese hamster ovary cells. Experimental Cell Research, 228(2), 277–282.
Darzynkiewicz, Z., et al. (1982). Cell heterogeneity during the cell cycle. Journal of Cellular Physiology, 113(3), 465–474.
Galkowski, D. et al. (2017). Of cytometry, stem cells and fountain of Youth. Stem Cell Reviews and Reports. doi:10.1007/s12015-017-9733-5.
Whitfield, M. L., et al. (2000). Stem-loop binding protein, the protein that binds the 3′ end of histone mRNA, is cell cycle regulated by both translational and posttranslational mechanisms. Molecular and Cellular Biology, 20(12), 4188–4198.
Morimoto, H., et al. (2009). Phenotypic plasticity of mouse spermatogonial stem cells. PloS One, 4(11), e7909.
Medema, J. P. (2013). Cancer stem cells: The challenges ahead. Nature Cell Biology, 15(4), 338–344.
Habibian, H. K., et al. (1998). The fluctuating phenotype of the lymphohematopoietic stem cell with cell cycle transit. The Journal of Experimental Medicine, 188(2), 393–398.
Kono, T., et al. (2004). Birth of parthenogenetic mice that can develop to adulthood. Nature, 428(6985), 860–864.
Wu, Q., et al. (2006). Regulated expression of two sets of paternally imprinted genes is necessary for mouse parthenogenetic development to term. Reproduction, 131(3), 481–488.
Qian, P., et al. (2016). The Dlk1-Gtl2 locus preserves LT-HSC function by inhibiting the PI3K-mTOR pathway to restrict mitochondrial metabolism. Cell Stem Cell, 18(2), 214–228.
Venkatraman, A., et al. (2013). Maternal imprinting at the H19-Igf2 locus maintains adult haematopoietic stem cell quiescence. Nature, 500(7462), 345–349.
Acknowledgments
This work was supported by the Stella and Henry Endowment, and the NCN OPUS grant 2016/21/B/NZ4/00201 from the National Science Center in Poland to Magdalena Kucia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosures
The authors indicate no potential conflict of interest.
Rights and permissions
About this article
Cite this article
Sellers, Z.P., Schneider, G., Bujko, K. et al. Do Cancer Cell Lines Have Fixed or Fluctuating Stem Cell Phenotypes? – Studies with the NTera2 Cell Line. Stem Cell Rev and Rep 13, 603–610 (2017). https://doi.org/10.1007/s12015-017-9743-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-017-9743-3