Abstract
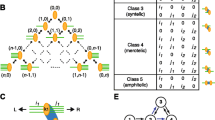
Cell division is a complex process requiring the cell to have many internal checks so that division may proceed and be completed correctly. Failure to divide correctly can have serious consequences, including progression to cancer. During mitosis, chromosomal segregation is one such process that is crucial for successful progression. Accurate segregation of chromosomes during mitosis requires regulation of the interactions between chromosomes and spindle microtubules. If left uncorrected, chromosome attachment errors can cause chromosome segregation defects which have serious effects on cell fates. In early prometaphase, where kinetochores are exposed to multiple microtubules originating from the two poles, there are frequent errors in kinetochore-microtubule attachment. Erroneous attachments are classified into two categories, syntelic and merotelic. In this paper, we consider a stochastic model for a possible function of syntelic and merotelic kinetochores, and we provide theoretical evidence that merotely can contribute to lessening the stochastic noise in the time for completion of the mitotic process in eukaryotic cells.




Similar content being viewed by others
References
Ault JG, Rieder CL (1992a) Chromosome mal-orientation and reorientation during mitosis. Cell Motil Cytoskeleton 22:155–159
Ault JG, Rieder CL (1992b) Chromosome mal-orientation and reorientation during mitosis. Cell Motil Cytoskelet 22(3):155–159
Biggins S, Severin FF, Bhalla N, Sassoon I, Hyman AA, Murray AW (1999) The conserved protein kinase ipl1 regulates microtubule binding to kinetochores in budding yeast. Genes Dev 13(5):532–544
Blake W, Kaern M, Cantor C, Collins J (2003) Noise in eukaryotic gene expression. Nature 422:633–637
Bolton MA, Lan W, Powers SE, McCleland ML, Kuang J, Stukenberg PT (2002) Aurora B kinase exists in a complex with survivin and INCENP and its kinase activity is stimulated by survivin binding and phosphorylation. Mol Biol Cell 13(9):3064–3077
Carmena M, Earnshaw WC (2003) The cellular geography of aurora kinases. Nat Rev Mol Cell Biol 4(11):842–854
Cheeseman IM, Anderson S, Jwa M, Green EM, seog Kang J, Yates JR, Chan CS, Drubin DG, Barnes G (2002) Phospho-regulation of kinetochore-microtubule attachments by the aurora kinase ipl1p. Cell 111(2):163–172
Cheeseman IM, Desai A (2002) Molecular architecture of the kinetochore-microtubule interface. Nat Rev Mol Cell Biol 9(1):1471–0072
Cimini D, Degrassi F (2005) Aneuploidy: a matter of bad connections. Trends Cell Biol 15(8):442–451
Cimini D, Moree B, Canman J, Salmon E (2003) Merotelic kinetochore orientation occurs frequently during early mitosis in mammalian tissue cells, and error correction is achieved by two different mechanisms. J Cell Sci 116:4213–4225
Ditchfield C, Johnson VL, Tighe A, Ellston R, Haworth C, Johnson T, Mortlock A, Keen N, Taylor SS (2003) Aurora b couples chromosome alignment with anaphase by targeting bubr1, mad2, and cenp-e to kinetochores. J Cell Biol 161(2):267280
Eldar A, Elowitz MB (2010) Functional roles for noise in genetic circuits. Nature 467:167–173
Elowitz M, Levine A, Siggia E, Swain P (2002) Stochastic gene expression in a single cell. Science 297:1183–1186
Francisco L, Wang W, Chan CS (1994) Type 1 protein phosphatase acts in opposition to ipl1 protein kinase in regulating yeast chromosome segregation. Mol Cell Biol 14(7):4731–4740
Gadgil C, Lee C, Othmer H (2005) A stochastic analysis of first-order reaction networks. Bull Math Biol 67:901–946
Gardiner C (1985) Handbook of stochastic methods for physics chemistry and the natural sciences, 2nd edn. Springer, New York
Gay G, Courtheoux T, Reyes C, Tournier S, Gachet Y (2012) A stochastic model of kinetochore-microtubule attachment accurately describes fission yeast chromosome segregation. J Cell Biol 196:757–774
Gillespie D (1976) A general method for numerically simulating the stochastic time evolution of coupled chemical reactions. J Comput Phys 22:403–434
Gillespie D (2007) Stochastic simulation of chemical kinetics. Annu Rev Phys Chem 58:25–55
Goshima G, Ndlec F, Vale RD (2005) Mechanisms for focusing mitotic spindle poles by minus enddirected motor proteins. J Cell Biol 171(2):229–240
Gregan J, Polakova S, Zhang L, Toli-Nrrelykke IM, Cimini D (2011) Merotelic kinetochore attachment: causes and effects. Trends Cell Biol 21(6):374–381
Hasty J, Collins JJ (2002) Translating the noise. Nat Genet 31:13–15
Hauf S, Cole RW, LaTerra S, Zimmer C, Schnapp G, Walter R, Heckel A, van Meel J, Rieder CL, Peters JM (2003) The small molecule hesperadin reveals a role for aurora b in correcting kinetochoremicrotubule attachment and in maintaining the spindle assembly checkpoint. J Cell Biol 161:281–294
Higham D (2008) Modeling and simulating chemical reactions. SIAM Rev 50(2):347–368
Kallio MJ, McCleland ML, Stukenberg P, Gorbsky GJ (2002) Inhibition of aurora b kinase blocks chromosome segregation, overrides the spindle checkpoint, and perturbs microtubule dynamics in mitosis. Curr Biol 12(11):900–905
Karsenti E, Vernos I (2001) The mitotic spindle: a self-made machine. Science 294(5542):543–547
Kepler T, Elston T (2001) Stochasticity in transcriptional regulation: origins, consequences, and mathematical representations. Biophys J 81(6):3116–3136
Kirschner M, Mitchison T (1986) Beyond self-assembly: from microtubules to morphogenesis. Cell 45(3):329–342
Lampson MA, Cheeseman IM (2011) Sensing centromere tension: aurora b and the regulation of kinetochore function. Trends Cell Biol 21(3):133–140
Lampson MA, Renduchitala K, Khodjakov A, Kapoor TM (2004) Correcting improper chromosome-spindle attachments during cell division. Nat Cell Biol 6(3):232–237
Laurenzi I (2000) An analytical solution of the stochastic master equation for reversible biomolecular reaction kinetics. J Chem Phys 113(8):3315–3322
Maresca TJ, Salmon ED (2010) Welcome to a new kind of tension: translating kinetochore mechanics into a wait-anaphase signal. J Cell Sci 123(6):825–835
Meunier S, Vernos I (2012) Microtubule assembly during mitosis from distinct origins to distinct functions? J Cell Sci 125(12):2805–2814
Mistry H, MacCallum D, Jackson R, Chaplain M, Davidson F (2008) Modeling the temporal evolution of the spindle assembly checkpoint and role of Aurora B kinase. PNAS 105(51):20215–20220
Mitchison TJ, Salmon ED (2001) Mitosis: a history of division. Nat Cell Biol 3:E17–E21
Morrow CJ, Tighe A, Johnson VL, Scott MI, Ditchfield C, Taylor SS (2005) Bub1 and aurora B cooperate to maintain BubR1-mediated inhibition of APC/CCdc20. J Cell Sci 118(16):3639–3652
Musacchio A, Salmon E (2007) The spindle-assembly checkpoint in space and time. Nat Rev Mol Cell Biol 8(5):379–393
Ozbudak E, Thattai M, Kurtser I, Grossman A, van Oudenaarden A (2002) Regulation of noise in the expression of a single gene. Nat Genet 31(1):69–73
Paul R, Wollman R, Silkworth WT, Nardi IK, Cimini D, Mogilner A (2009) Computer simulations predict that chromosome movements and rotations accelerate mitotic spindle assembly without compromising accuracy. Proc Nat Acad Sci 106(37):15708–15713
Rasner JM, O’Shea EK (2004) Control of stochasticity in eukaryotic gene expression. Science 304:1811–1814
Rieder CL, Maiato H (2004) Stuck in division or passing through: what happens when cells cannot satisfy the spindle assembly checkpoint. Dev Cell 7(5):637–651
Ruchaud S, Carmena M, Earnshaw WC (2007) Chromosomal passengers: conducting cell division. Nat Rev Mol Cell Biol 8(10):798–812
Swain P, Elowitz M, Siggia E (2002) Intrinsic and extrinsic contributions to stochasticity in gene expression. Proc Natl Acad Sci USA 99(20):12795–12800
Tanaka K, Mukae N, Dewar H, van Breugel M, James EK, Prescott AR, Antony C, Tanaka TU (2005) Molecular mechanisms of kinetochore capture by spindle microtubules. Nature 434:987–994
Tanaka T (2008) Bi-orienting chromosomes: acrobatics on the mitotic spindle. Chromosoma 117:521–533
Tanaka TU, Rachidi N, Janke C, Pereira G, Galova M, Schiebel E, Stark MJ, Nasmyth K (2002) Evidence that the ipl1-Sli15 (Aurora Kinase-INCENP) complex promotes chromosome bi-orientation by altering kinetochore-spindle pole connections. Cell 108(3):317–329
Thattai M, van Oudenaarden A (2001) Intrinsic noise in gene regulatory networks. Proc Natl Acad Sci USA 98(15):8614–8619
Tulu US, Fagerstrom C, Ferenz NP, Wadsworth P (2006) Molecular requirements for kinetochore-associated microtubule formation in mammalian cells. Curr Biol 16:536–541
van Kampen N (2007) Stochastic processes in physics and chemistry, 3rd edn. Elsevier, Amsterdam
Acknowledgments
This research has been supported in part by the Mathematical Biosciences Institute and the National Science Foundation under Grant DMS 0931642. The research of Sijia Liu has been supported in part by an Alberta Wolfe Research Fellowship from the Iowa State University Mathematics department. The research of Blerta Shtylla has been supported in part by the National Science Foundation under Grant NSF DMS 1358932. MAJC gratefully acknowledges the support of the ERC Advanced Investigator Grant 227619, “M5CGS - From Mutations to Metastases: Multiscale Mathematical Modelling of Cancer Growth and Spread”.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix 1: Numerical Computations
Extensions of the analysis presented in this paper for more complex models that include explicitly the effect of Aurora B, such as the models discussed by Mistry et al. (2008), are highly non-trivial, and hence numerical approaches become crucial. In this appendix, we employ the Gillespie simulation algorithm (Gillespie 1976, 2007) to compute an ensemble of independent realizations of the master Eq. (2). This allows us, among other things, to numerically approximate the distribution of the hitting time \(T\) of the amphitelic state, i.e., of the state where all kinetochores are amphitelically attached. As we will see, the results of the numerical computations corroborate the analytical derivation of Sect. 3.1.
As explained in Sect. 2, because of the small numbers of chemically interacting species in the system, we treat the evolution of the latter as a Markovian stochastic process, rather than as a deterministic process (Higham 2008), and use the standard master equation formalism (Gardiner 1985; van Kampen 2007). In this context, Monte Carlo methods are commonly used to generate realizations of the underlying stochastic process. For chemical reaction networks this is frequently done using one of the two stochastic simulation algorithms, called the direct method and first reaction method, respectively, developed by Gillespie (1976).
Simulations were conducted by choosing a value in the range discussed for the parameters \(k_1\) and \(k_3\) and performing 10,000 realizations using Gillespie’s direct method on these parameter values. Following the assumptions outlined in Mistry et al. (2008), each realization started with all 46 kinetochore pairs in the unattached state and then proceeded by updating the populations of the kinetochore attachment types in the previous time step according to Gillespie’s direct method, with possible transitions following the network diagram in Fig. 1. A realization was then deemed to have finished when all 46 kinetochore pairs reached the amphitelic state. The amount of time required to reach this state, i.e., the hitting time for the specific realization, was then recorded. This process was repeated for each of the 10,000 realizations. The results for all realizations for a specific choice of parameter values are shown in Fig. 5.
The data in Fig. 5 (and similar data sets for other combinations of parameter values) were used to compute the empirical mean \(\mu \) and empirical standard deviation \(\sigma \) of the all-amphitelic hitting time distribution. Ensembles of 10,000 realizations were generated for all possible combinations of values in the range of the parameters \(k_1\) and \(k_3\) described in Sect. 3, and the respective coefficients of variation were calculated for all possible pairs of parameter values. The results of these simulations are plotted in Fig. 2b. This can be compared with the plot in Fig. 2a which shows the values of the CV over the same range of values for the parameters \(k_1 \text{ and } k_3\) as computed by using the analytically derived probability distribution for the hitting time, discussed in Sect. 3.1. The comparison of Fig. 2a, b clearly shows that both the stochastic simulation approach based on Gillespie’s direct method and the analytical derivation of Sect. 3.1 return similar results.
Appendix 2: On the Coefficient of Variation and the Fano Factor
A significant number of cellular processes involve reactions whose components are present in low concentrations, and this is especially true for the reactions involved in gene expression (Rasner and O’Shea 2004). The low concentrations of the components naturally give rise to stochastic fluctuations, typically called noise, in the number of molecules produced by these reaction (Eldar and Elowitz 2010). Two quantities are commonly used in the literature to measure the noise of the molecular concentrations: the CV, defined as the standard deviation divided by the mean, and the Fano factor \(({\mathcal F})\), defined as the variance divided by the mean (Rasner and O’Shea 2004). While the \(CV\) is largely accepted as the standard measurement of noise, the Fano factor is a valid measure of noise under specific circumstances (Gadgil et al. 2005). Furthermore, inappropriate application of the Fano factor can lead to misinterpret the role of noise biological processes (Swain et al. 2002; Gadgil et al. 2005), and the following literature review is provided to clarify the meaning, utility, and appropriate application of the Fano factor.
The Fano factor, or the noise strength (e.g., Rasner and O’Shea 2004), was introduced by Thattai and van Oudenaarden (2001) in their analysis of gene expression in prokaryotes. It measures noise as a deviation away from the Poisson distribution, and as opposed to the coefficient of variation, the amount of noise measured by the Fano factor does not diminish for increasing mean values (Ozbudak et al. 2002). Following the derivation of Thattai and van Oudenaarden (2001), let \(r\) and \(p\) denote the number of mRNA and protein molecules, respectively, and define
where \(k_R\) and \(k_P\) can be viewed as the rates for transcription and translation, respectively. Their first-order analysis of single gene expression at steady state determined that the protein numbers are given by a distribution with mean
and variance
Thus the Fano factor is calculated as
where \(\eta ={\upgamma }_P/{\upgamma }_R\) and \(b=k_p/{\upgamma }_R\), typically called the burst size, is the average number of protein molecules created from a strand of mRNA. However, the life span of a mRNA strand is much shorter than the life span of a protein molecule (Thattai and van Oudenaarden 2001). Therefore, \(\eta \) is typically small and (9) implies \({\mathcal F}\cong b+1\), and the authors conclude that intrinsic noise is controlled at the translational level (Thattai and van Oudenaarden 2001). Subsequent work by van Oudenaarden and collaborators, (Ozbudak et al. 2002), varied the transcription and translation rates independently and experimentally verified the conclusion that noise in the protein number is controlled at the translational level (Ozbudak et al. 2002; Hasty and Collins 2002).
Subsequent research was conducted by Kepler and Elston (2001) modeling stochastic fluctuations in protein number arising from operator fluctuations, the addition or removal of proteins bound to DNA which overlap the promoter region of RNA polymerase. The results of those simulation demonstrated that the noise, as measured by the CV, in protein number could arise from the regulation of transcription (Kepler and Elston 2001; Hasty and Collins 2002). Additionally, research conducted by Blake et al. (2003) on the noise, as measured by the Fano factor, in eukaryotic gene expression arise from transcriptional regulation, “providing an important distinction between the sources of noise in prokaryotic and eukaryotic cells.” While it was eventually shown that the sources of noise in eukaryotic and prokaryotic gene expression are the same (see, e.g., Rasner and O’Shea 2004; Eldar and Elowitz 2010), the discussion above clearly demonstrates the lack of consensus concerning the validity of the Fano factor as a measurement of noise.
Research conducted by Swain et al. (2002) clearly demonstrated complications that arise from using the Fano factor as the only measurement of noise. The source of the potential complications stems from the fact that Fano factor removes the dependence on transcription rate and scales with the translation rate (Swain et al. 2002; Rasner and O’Shea 2004). As demonstrated by the in depth theoretical analysis of Gadgil et al. (2005), the Poisson distribution is the limiting distribution of all open conversion networks. Therefore, as stated in Gadgil et al. (2005):
The Fano factor as the measure of noise leads to the prediction that the fluctuations of all components in an open conversion reaction system are identical. Thus a species that has a mean of 10 molecules will exhibit the same amount of noise as a species that has a mean concentration of 1M.
Any usage of the Fano factor in this paper is appropriate. The network presented in this paper is closed. Furthermore, any comparisons made between the Fano factor and the coefficient of variation are done with the same data.
In conclusion, we would like to thank the reviewers for their thoughtful criticism, especially the reviewer at whose request this appendix was written. Additionally, we refer interested readers to the following papers: Gadgil et al. (2005) contains thorough review of theoretical first-order analysis techniques, and Eldar and Elowitz (2010) provides an excellent review for the role of noise in gene expression. Furthermore, Eldar and Elowitz (2010) contains numerous references to review papers covering mathematical representations for the sources of noise in gene expression, the analysis of noise in the context of dynamic circuits, and the advantages of noise on phenotypic variability.
Rights and permissions
About this article
Cite this article
Matzavinos, A., Shtylla, B., Voller, Z. et al. Stochastic Modelling of Chromosomal Segregation: Errors Can Introduce Correction. Bull Math Biol 76, 1590–1606 (2014). https://doi.org/10.1007/s11538-014-9969-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-014-9969-z