Abstract
A universal feature of the biochemistry of any living system is that all the molecules and catalysts that are required for reactions of the system can be built up from an available food source by repeated application of reactions from within that system. RAF (reflexively autocatalytic and food-generated) theory provides a formal way to study such processes. Beginning with Kauffman’s notion of “collectively autocatalytic sets,” this theory has been further developed over the last decade with the discovery of efficient algorithms and new mathematical analysis. In this paper, we study how the behaviour of a simple binary polymer model can be extended to models where the pattern of catalysis more precisely reflects the ligation and cleavage reactions involved. We find that certain properties of these models are similar to, and can be accurately predicted from, the simple binary polymer model; however, other properties lead to slightly different estimates. We also establish a number of new results concerning the structure of RAFs in these systems.





Similar content being viewed by others
Notes
The first equation holds because in RAND, and for every molecule x, N(x) has a Poisson distribution with mean—and, therefore, variance—equal to m; the second equation is from the identity Var[N(χ)]=E[Var[N|χ]]+Var[E[N|χ]], together with the fact that half the molecules have maximal length.
References
Bonchev, D., & Mekenyan, O. (1994). Graph theoretical approaches to chemical reactivity. Dordrecht: Kluwer.
Crick, F. H. C. (1958). On protein synthesis. Symp. Soc. Exp. Biol., 12, 138–163.
Crick, F. H. C. (1970). Central dogma of molecular biology. Nature, 227, 561–563.
Eigen, M. (1971). Self-organization of matter and the evolution of biological macromolecules. Naturwissenschaften, 58, 465–523.
Eigen, M., & Schuster, P. (1979). The hypercycle. Berlin: Springer.
Garey, M. R., & Johnson, D. S. (1979). Computers and intractability: a guide to the theory of NP-completeness. New York: Freeman.
Hordijk, W., & Steel, M. (2004). Detecting autocatalytic, self-sustaining sets in chemical reaction systems. J. Theor. Biol., 227(4), 451–461.
Hordijk, W., & Steel, M. (2012a). Autocatalytic sets extended: dynamics, inhibition, and a generalization. J. Syst. Chem., 3, 5.
Hordijk, W., & Steel, M. (2012b). Predicting template-based catalysis rates in a simple catalytic reaction model. J. Theor. Biol., 295, 132–138.
Hordijk, W., & Steel, M. (2013). A formal model of autocatalytic sets emerging in an RNA replicator system. J. Syst. Chem., 4, 3.
Hordijk, W., Kauffman, S. A., & Steel, M. (2011). Required levels of catalysis for emergence of autocatalytic sets in models of chemical reaction systems. Int. J. Mol. Sci., 12(5), 3085–3101.
Hordijk, W., Steel, M., & Kauffman, S. (2012). The structure of autocatalytic sets: evolvability, enablement, and emergence. Acta Biotheor., 60(4), 379–392.
Kauffman, S. A. (1971). Cellular homeostasis, epigenesis and replication in randomly aggregated macromolecular systems. J. Cybern., 1(1), 71–96.
Kauffman, S. A. (1986). Autocatalytic sets of proteins. J. Theor. Biol., 119, 1–24.
Kauffman, S. A. (1993). The origins of order. Oxford: Oxford University Press.
Mincheva, M., & Roussel, M. R. (2007). Graph-theoretic methods for the analysis of chemical and biochemical networks, I: multistability and oscillations in ordinary differential equation models. J. Math. Biol., 55, 61–86.
Mossel, E., & Steel, M. (2005). Random biochemical networks: the probability of self-sustaining autocatalysis. J. Theor. Biol., 233(3), 327–336.
Schrödinger, E. (1944). What is life? Cambridge: Cambridge University Press.
Steel, M. (2000). The emergence of a self-catalysing structure in abstract origin-of-life models. Appl. Math. Lett., 3, 91–95.
Steel, M., Hordijk, W., & Smith, J. (2013). Minimal autocatalytic networks. J. Theor. Biol., 332, 96–107.
Vaidya, N., Manapat, M. L., Chen, I. A., Xulvi-Brunet, R., Hayden, E. J., & Lehman, N. (2012). Spontaneous network formation among cooperative RNA replicators. Nature, 491, 72–77.
Vasas, V., Fernando, C., Santos, M., Kauffman, S., & Sathmáry, E. (2012). Evolution before genes. Biol. Direct, 7, 1.
Watson, J. D., & Crick, F. H. C. (1953). Genetical implications of the structure of deoxyribonucleic acid. Nature, 171, 964–967.
Wills, P., & Henderson, L. (2000). Self-organisation and information-carrying capacity of collectively autocatalytic sets of polymers: ligation systems. In Y. Bar-Yam (Ed.), Unifying themes in complex systems: proceedings of the first international conference on complex systems (pp. 613–623). Jackson: Perseus.
Acknowledgements
We thank the Allan Wilson Centre for Molecular Ecology and Evolution and the Alexander von Humboldt Foundation for helping fund part of this research. We also thank an anonymous reviewer for several helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Appendix: Proof of Theorem 2
Appendix: Proof of Theorem 2
Proof
We will reduce the graph theoretic problem VERTEX COVER to k-cat-RAF (a similar reduction was employed in Steel et al. (2013) for a quite different problem). Recall that for a graph G=(V,E), a vertex cover of G is a subset V′ of V with the property that each edge of G is incident with at least one vertex in V′; VERTEX COVER has as its instance a graph G=(V,E) and an integer K and we ask whether or not G has a vertex cover of size at most K. This is a well-known NP-complete problem (Garey and Johnson 1979) (indeed, it is one of Karp’s original 21 NP-complete problems). Given an instance (G=(V,E),K) of VERTEX COVER we show how to construct an instance \((X_{G}, \mathcal{R}_{G}, C_{G}, F_{G}, k)\), of k-cat-RAF for which the answers to the two decision problems are identical (here, \(\mathcal{R}_{G}\) is an RAF).
First we construct F G and X G . For each v∈V let a v ,b v be two distinct elements of F G and let x v be an element of X G −F G . Order E as e 1,…,e |E| and for each j=1,…,|E| let d j be a distinct element of F and y j an element of X G −F G . Let d 0 be another distinct element of F G . Thus F G consists of the 2|V|+|E|+1 elements:
X G −F G consists of |V|+|E| elements:
For each v∈V, define a reaction
For each 1<j≤|E|, define the reaction:
and for j=1 let:
For any subset U of V, let \(\mathcal{R}_{U} = \{r_{v}: v \in U\}\), let
and set \(\mathcal{R}_{G} = \mathcal{R}_{V} \cup\mathcal{R}_{E}\). Thus we have specified X G ,F G and \(\mathcal{R}_{G}\) and it remains to define the catalysis (C G ) assignment, which is as follows:
-
If e j=(u j,v j) (where u j,v j∈V) then \(r'_{j}\) is catalyzed by both \(x_{u^{j}}\) and \(x_{v^{j}}\) (but by no other molecules).
-
In addition, each reaction r v :v∈V is catalyzed by y |E| and by no other molecule—we call the molecule y |E| the super-catalyst.
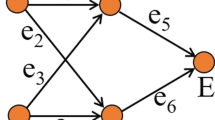
An example of this construction is illustrated in Fig. 6. We have now fully specified the catalysation and thereby the pair \((\mathcal{Q} _{G}, F_{G})\) constructed from G (\(\mathcal{Q}_{G} = (X_{G}, \mathcal{R}_{G}, C_{G})\)).
CLAIM: A finite graph G has a vertex cover of size at most K if and only there is an ordering of \(\mathcal{R}_{G}\) that satisfies (P1) and involves at most K violations of (P2).
To establish this claim, first suppose that V′ is a vertex cover of G of size at most K. Then order \(\mathcal{R}_{V'}\) arbitrarily and place these as the first reactions in a linear ordering, followed by the reactions in \(\mathcal{R}_{E}\) in the order \(r'_{1}, \ldots, r'_{E}\), and finally the remaining reactions in \(\mathcal{R}_{V-V'}\) in arbitrary order as the final segment of the ordering. This ordering just requires |V′|=K violations of (P2) for the initial reactions (i.e., \(\mathcal {R}_{V'}\)), and it also satisfies (P1), and so provides the required ordering of \(\mathcal{R}_{G}\).
Conversely, suppose that there is an ordering of \(\mathcal{R}_{G}\), r 1,…,r |V|+|E| that satisfies (P1) and involves at most K violations of (P2). Let J denote the set of j for which (P2) fails for r j , and let
Each j∈J E corresponds to some edge e of G, so we will let v(j) denote any vertex of G incident with e.
Now, {r j :j∈J V }∪{r v(j):j∈J E } is a subset of \(\mathcal{R}_{V}\) of size at most K, and so corresponds to \(\mathcal {R}_{V'}\) for a subset V′ of V of size at most K. We show that V′ is an edge cover of G, by showing that any given edge e contains at least one vertex from V′.
First, observe that the reaction \(r'_{e} \in\mathcal{R}_{E}\) is one of the reactions r j in the above ordering of \(\mathcal{R}\). We consider two cases: (i) j∈J E and (ii) j∉J E . In Case (i), v(j)∈V′ and so e contains this vertex from V′. In Case (ii), r j is catalyzed by a product of a reaction r i that appears earlier in the ordering. This implies that r i =r v for some vertex v of V; therefore, if v∈V′ then e contains an element of V′. It remains to consider the case where v∉V′ (i.e., i∉J V ). We will show that this case never arises by deriving a contradiction on the assumption that it does. If i∉J V , then r i is catalyzed by a reaction r k that appears earlier than i in the given ordering of \(\mathcal{R}\). However, the only reaction that can catalyze r i is \(r'_{|{E}|}\), which must therefore appear as r k for some k<i in the ordering (since we are assuming that i∉J V ). Summarizing, we have
It is at this point that we invoke (P1). Notice that the reactants for r k do not become available until all the other reactions in \(\mathcal{R}_{E}\)—including r j —have occurred. By (P1), this requires that j<k. Combining this with inequality (5), we obtain the required contradiction required to exclude the last case. This completes the proof. □
Rights and permissions
About this article
Cite this article
Hordijk, W., Wills, P.R. & Steel, M. Autocatalytic Sets and Biological Specificity. Bull Math Biol 76, 201–224 (2014). https://doi.org/10.1007/s11538-013-9916-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-013-9916-4