Abstract
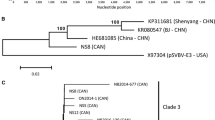
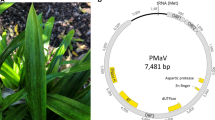
The presence of Gooseberry vein banding associated virus (GVBaV), a badnavirus in the family Caulimoviridae, is strongly correlated with gooseberry vein banding disease in Ribes spp. In this study, full-length genomic sequences of four GVBaV isolates from different hosts and geographic regions were determined to be 7649–7663 nucleotides. These isolates share identities of 96.4–97.3% for the complete genomic sequence, indicating low genetic diversity among them. The GVBaV genome contains three open reading frames (ORFs) on the plus strand that potentially encode proteins of 26, 16, and 216 kDa. The size and organization of GVBaV ORFs 1–3 are similar to those of most other badnaviruses. The putative amino acid sequence of GVBaV ORF 3 contained motifs that are conserved among badnavirus proteins including aspartic protease, reverse transcriptase, and ribonuclease H. The highly conserved putative plant tRNAmet-binding site is also present in the 935-bp intergenic region of GVBaV. The identities of the genomic sequences of GVBaV and other badnaviruses range from 49.1% (Sugarcane bacilliform Mor virus) to 51.7% (Pelargonium vein banding virus, PVBV). Phylogenetic analysis using the amino acid sequence of the ORF 3 putative protein shows that GVBaV groups most closely to Dioscorea bacilliform virus, PVBV, and Taro bacilliform virus. These results confirm that GVBaV is a pararetrovirus of the genus Badnavirus in the family Caulimoviridae.




Similar content being viewed by others
References
A.N. Adams, A.F. Posnette, in Virus Diseases of Small Fruits, ed. by R.H. Converse (USDA Agriculture Handbook No. 631, 1987), pp. 129–130
J.D. Postman, Plant Dis. 77, 953 (1993)
A.N. Adams, J. Hortic. Sci. 54, 23–25 (1979)
T.A. Jones, W.J. McGavin, A.D.W. Geering, B.E.L. Lockhart, Plant Dis. 85, 417–422 (2001)
T.A. Jones, W.J. McGavin, C.A. Watkins, Acta Hortic. 308, 31–35 (1992)
R. Hull, A. Geering, G. Harper, B.E. Lockhart, J.E. Schoelz, in Virus Taxonomy, 8th Report of the International Committee for Taxonomy of Viruses, ed. by C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball (Elsevier Academic Press, San Diego, 2005), pp. 385–396
E. Jacquot, L.S. Hagen, M. Jacquemond, P.P. Yot, Virology 225, 191–195 (1996)
C.-P. Cheng, B.E.L. Lockhart, N.E. Olszewski, Virology 223, 263–271 (1995)
S.L. Medberry, B.E.L. Lockhart, N.E. Olszewski, Nucleic Acids Res. 18, 5505–5513 (1990)
G. Harper, R. Hull, Virus Genes 17, 271–278 (1998)
I. Tzafrir, L. Ayala-Navarrette, B.E.L. Lockhart, N.E. Olszewski N.E, Virology 232, 359–368 (1997)
P. Pfeiffer, T. Hohn, Cell 33, 781–789 (1983)
R.D. Qu, M. Bhattacharyya, G.S. Laco, A. De Kochko, B.L. Rao, M.B. Kaniewska, J.S. Elmer, D.E. Rochester, C.E. Smith, R.N. Beachy, Virology 185, 354–364 (1991)
G. Harper, D. Hart, S. Moult, R. Hull, A. Geering, J. Thomas, Arch. Virol. 150, 2407–2420 (2005)
E. Muller, S. Sackey, Arch. Virol. 150, 53–66 (2005)
L. Kenyon, B.S.M. Lebas, S.E. Seal, Arch. Virol. 153, 877–889 (2008)
R. Li, R. Mock, Q. Huang, J. Abad, J. Hartung, G. Kinard, J. Viral Methods 154, 48–55 (2008)
L. Falquet, M. Pagni, P. Bucher, N. Hulo, C.J. Sigrist, K. Hofmann, A. Bairoch, Nucleic Acids Res. 30, 235–238 (2002)
K. Tamura, J. Dudley, M. Nei, S. Kumar, Mol. Biol. Evol. 24, 1596–1599 (2007)
B.K. Borah, A.M.A. Johnson, D.V.R. Sai Gopal, I. Dasgupta, Virus Genes 39, 137–140 (2009)
M. Bouhida, B.E.L. Lockhart, N.E. Olszewski, J. Gen. Virol. 74, 15–22 (1993)
L.S. Hagen, M. Jacquemond, A. Lepingle, H. Lot, M. Tepfer, Virology 196, 619–628 (1993)
R.W. Briddon, S. Phillips, A. Brunt, R. Hull, Virus Genes 18, 277–283 (1999)
Q. Huang, J.S. Hartung, J. Gen. Virol. 82, 2549–2558 (2001)
I.C. Yang, G.J. Hafner, J.L. Dale, R.M. Harding, Arch. Virol. 148, 937–949 (2003)
L. Su, S. Gao, Y. Huang, C. Ji, D. Wang, Y. Ma, R. Fang, X. Chen, Virus Genes 35, 423–429 (2007)
E.V. Koonin, A.R. Mushegian, E.V. Ryabov, V.V. Dolja, J. Gen. Virol. 72, 2895–2903 (1991)
L.A. Calvert, D. Manuel, M.D. Ospina, R.J. Shepherd, J. Gen. Virol. 76, 1271–1278 (1995)
K.R. Richert-Poggeler, R.J. Shepherd, Virology 236, 137–147 (1997)
S. Seal, E. Muller, Arch. Virol. 152, 819–825 (2007)
H. Kano, M. Koizumi, H. Noda, H. Hibino, K. Ishikawa, T. Omura, P.Q. Cabauatan, H. Koganezawa, Arch. Virol. 124, 157–163 (1992)
L.H. Pearl, W.R. Taylor, A structural model for the retroviral proteases. Nature 329, 351–354 (1987)
Y. Xiong, T.H. Eickbush, EMBO J. 9, 3353–3362 (1990)
H.S. Malik, T.H. Eickbush, Genome Res. 11, 1187–1197 (2001)
M. Bousalem, E.J. Douzery, S.E. Seal, Arch. Virol. 153, 1085–1102 (2008)
A.D.W. Geering, L.A. McMichael, R.G. Dietzgen, J.E. Thomas, Phytopathology 90, 921–927 (2000)
R.J. Geijskes, K.S. Braithwaite, J.L. Dale, R.M. Harding, G.R. Smith, Arch. Virol. 147, 2393–2404 (2002)
A.O. Eni, J.d’.A. Hughes, R. Asiedu, M.E.C. Rey, Arch. Virol. 153, 2263–2272 (2008)
K.N. Gupta, V.K. Baranwal, B.K. Prasanna, J. Singh, Q.M.R. Haq, K. Gopal, Int. J. Virol. 5, 143–153 (2009)
I.M. Roberts, A.T. Jones, Ann. Appl. Biol. 130, 77–89 (1997)
I.P. Adams, R.H. Glover, W.A. Monger, R. Mumford, E. Jackeviciene, M. Navalinskiene, M. Samuitiene, N. Boonham, Mol. Plant Pathol. 10, 537–545 (2009)
J.F. Kreuze, A. Perez, M. Untiveros, D. Quispe, S. Fuentes, I. Barker, R. Simon, Virology 388, 1–7 (2009)
Acknowledgments
We thank Gregory Anderson, Whitney Hymes, and Sam Grinstead for technical assistance and Drs. Joseph Postman, John Hartung, and Bob Martin for critical review of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, D., Mock, R., Kinard, G. et al. Molecular analysis of the complete genomic sequences of four isolates of Gooseberry vein banding associated virus . Virus Genes 43, 130–137 (2011). https://doi.org/10.1007/s11262-011-0614-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-011-0614-8