Abstract
Background
Gastric carcinoma (GC) is one of the most common malignant tumors. Although increasing studies have indicated that circular RNAs function as ideal biomarkers for multiple cancers, only a few researches elucidated the correlation between circular RNA PTK2 (circPTK2) and human cancers.
Aim
To further explore the expression status, biological function, and regulatory mechanism of circPTK2 in GC.
Methods
Bioinformatics analysis and function or mechanism experiments including RT-qPCR, flow cytometry, Western blot, luciferase reporter assay, and xenografts assays were applied to investigate the function of circPTK2 and miR-139-3p.
Results
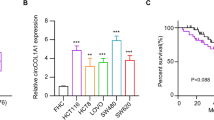
High expression of circPTK2 was presented in GC tissues and cells. The circPTK2 knockdown notably suppressed cell proliferation and promoted cell apoptosis in GC. In mechanism, circPTK2 served as a sponge of miR-139-3p. Inhibition of miR-139-3p could reverse circPTK2 silence-mediated effects on GC cell proliferation and apoptosis. Furthermore, the xenograft tumor model was established to investigate the role of circPTK2 in GC tumor growth. Experimental results delineated that the reduction in tumor growth in response to circPTK2 knockdown was partly recovered by miR-139-3p inhibitor.
Conclusions
CircPTK2 promotes GC development by sponging miR-139-3p, which may function as an effective gene target for managing GC.





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893–2917.
Kamangar F, Dores GM, Anderson WF. Patterns of cancer incidence, mortality, and prevalence across five continents: defining priorities to reduce cancer disparities in different geographic regions of the world. J Clin Oncol. 2006;24:2137–2150.
Van Cutsem E, Sagaert X, Topal B, Haustermans K, Prenen H. Gastric cancer. Lancet (London, England). 2016;388:2654–2664.
de Mestier L, Lardiere-Deguelte S, Volet J, Kianmanesh R, Bouche O. Recent insights in the therapeutic management of patients with gastric cancer. Dig Liver Dis.. 2016;48:984–994.
Digklia A, Wagner AD. Advanced gastric cancer: current treatment landscape and future perspectives. World J Gastroenterol. 2016;22:2403–2414.
Smolle E, Haybaeck J. Non-coding RNAs and lipid metabolism. Int J Mol Sci. 2014;15:13494–13513.
Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–338.
Jin J, Chen A, Qiu W, et al. Dysregulated circRNA_100876 suppresses proliferation of osteosarcoma cancer cells by targeting microRNA-136. J Cell Biochem. 2019;120:15678–15687.
Ma HB, Yao YN, Yu JJ, Chen XX, Li HF. Extensive profiling of circular RNAs and the potential regulatory role of circRNA-000284 in cell proliferation and invasion of cervical cancer via sponging miR-506. Am J Transl Res. 2018;10:592–604.
Zong L, Sun Q, Zhang H, et al. Increased expression of circRNA_102231 in lung cancer and its clinical significance. Biomed Pharmacother. 2018;102:639–644.
Shen F, Liu P, Xu Z, et al. CircRNA_001569 promotes cell proliferation through absorbing miR-145 in gastric cancer. J Biochem. 2019;165:27–36.
Wang S, Zhang X, Li Z, et al. Circular RNA profile identifies circOSBPL10 as an oncogenic factor and prognostic marker in gastric cancer. Oncogene. 2019;38:6985–7001.
Xu ZQ, Yang MG, Liu HJ, Su CQ. Circular RNA hsa_circ_0003221 (circPTK2) promotes the proliferation and migration of bladder cancer cells. J Cell Biochem. 2018;119:3317–3325.
Wang L, Tong X, Zhou Z, et al. Circular RNA hsa_circ_0008305 (circPTK2) inhibits TGF-beta-induced epithelial-mesenchymal transition and metastasis by controlling TIF1gamma in non-small cell lung cancer. Mol Cancer. 2018;17:140.
Verduci L, Strano S, Yarden Y, Blandino G. The circRNA-microRNA code: emerging implications for cancer diagnosis and treatment. Mol Oncol. 2019;13:669–680.
Dori M, Bicciato S. Integration of bioinformatic predictions and experimental data to identify circRNA-miRNA associations. Genes. 2019;10:642.
Chen L, Zhang S, Wu J, et al. circRNA_100290 plays a role in oral cancer by functioning as a sponge of the miR-29 family. Oncogene. 2017;36:4551–4561.
Yang CY, Zhang FX, He JN, Wang SQ. CircRNA_100876 promote proliferation and metastasis of breast cancer cells through adsorbing microRNA-361-3p in a sponge form. Eur Rev Med Pharmacol Sci. 2019;23:6962–6970.
Zhang Y, Liu H, Li W, et al. CircRNA_100269 is downregulated in gastric cancer and suppresses tumor cell growth by targeting miR-630. Aging. 2017;9:1585–1594.
Pei FL, Cao MZ, Li YF. Circ_0000218 plays a carcinogenic role in colorectal cancer progression by regulating miR-139-3p/RAB1A axis. J Biochem. 2020;167:55–65.
Xue F, Li QR, Xu YH, Zhou HB. MicroRNA-139-3p inhibits the growth and metastasis of ovarian cancer by inhibiting ELAVL1. OncoTargets Ther. 2019;12:8935–8945.
Huang P, Xi J, Liu S. MiR-139-3p induces cell apoptosis and inhibits metastasis of cervical cancer by targeting NOB1. Biomed Pharmacother. 2016;83:850–856.
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Chen H, Guan R, Lei Y, et al. Lymphangiogenesis in gastric cancer regulated through Akt/mTOR-VEGF-C/VEGF-D axis. BMC Cancer. 2015;15:103.
Msika S, Benhamiche AM, Jouve JL, Rat P, Faivre J. Prognostic factors after curative resection for gastric cancer. A population-based study. Eur J Cancer (Oxford, England: 1990). 2000;36:390–396.
Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12:381–388.
Ng WL, Mohd Mohidin TB, Shukla K. Functional role of circular RNAs in cancer development and progression. RNA Biol. 2018;15:995–1005.
Si-Tu J, Cai Y, Feng T, et al. Upregulated circular RNA circ-102004 that promotes cell proliferation in prostate cancer. Int J Biol Macromol. 2019;122:1235–1243.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–531.
Romero-Cordoba SL, Salido-Guadarrama I, Rodriguez-Dorantes M, Hidalgo-Miranda A. miRNA biogenesis: biological impact in the development of cancer. Cancer Biol Ther. 2014;15:1444–1455.
Tian W, Wu W, Li X, Rui X, Wu Y. MiRNA-139-3p inhibits the proliferation, invasion, and migration of human glioma cells by targeting MDA-9/syntenin. Biochem Biophys Res Commun. 2019;508:295–301.
Sannigrahi MK, Sharma R, Singh V, Panda NK, Rattan V, Khullar M. Role of host miRNA Hsa-miR-139-3p in HPV-16-induced carcinomas. Clin Cancer Res. 2017;23:3884–3895.
Pei FL, Cao MZ, Li YF. Circ_0000218 plays a carcinogenic role in colorectal cancer progression by regulating miR-139-3p/RAB1A axis. J Biochem. 2019;167:55–65.
Wu L, Liu D, Yang Y. Enhanced expression of circular RNA circ-DCAF6 predicts adverse prognosis and promotes cell progression via sponging miR-1231 and miR-1256 in gastric cancer. Exp Mol Pathol. 2019;110:104273.
Chan JJ, Tay Y. Noncoding RNA:RNA regulatory networks in cancer. Int J Mol Sci. 2018;19:1310.
Acknowledgments
We thank all participators for their help.
Author information
Authors and Affiliations
Contributions
The study conception and design, material preparation, data collection, and analysis were performed by DY and CZ; the first draft of the manuscript was written by DY; DY and CZ commented on previous versions of the manuscript. Both of the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yu, D., Zhang, C. Circular RNA PTK2 Accelerates Cell Proliferation and Inhibits Cell Apoptosis in Gastric Carcinoma via miR-139-3p. Dig Dis Sci 66, 1499–1509 (2021). https://doi.org/10.1007/s10620-020-06358-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-020-06358-4