Abstract
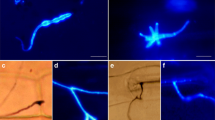
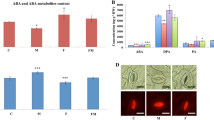
Heterobasidion annosum is widely known as a major root and butt rot pathogen of conifer trees, but little information is available on its interaction with the roots of herbaceous angiosperm plants. We investigated the infection biology of H. annosum during challenge with the angiosperm model Arabidopsis and monitored the host response after exposure to different hormone elicitors, chemicals (chitin, glucan and chitosan) and fungal species that represent diverse basidiomycete life strategies [e.g., pathogen (H. annosum), saprotroph (Stereum sanguinolentum) and mutualist (Lactarius rufus)]. The results revealed that the tree pathogen (H. annosum) and the saprotroph (S. sanguinolentum) could infect the Col-8 (Columbia) ecotype of Arabidopsis in laboratory inoculation experiments. Germinated H. annosum spores had appressorium-like penetration structures attached to the surface of the Arabidopsis roots. Subsequent invasive fungal growth led to the disintegration of the vascular region of the root tissues. Progression of root rot symptoms in Arabidopsis was similar to the infection development that was previously documented in Scots pine seedlings. Scots pine PsDef1 and Arabidopsis DEFLs (AT5G44973.1) and PDF1.2 were induced at the initial stage of the infection. However, differences in the expression patterns of the defensin gene homologs from the two plant groups were observed under various conditions, suggesting functional differences in their regulation. The potential use of the H. annosum–Arabidopsis pathosystem as a model for studying forest tree diseases is discussed.







Similar content being viewed by others
Abbreviations
- ACC:
-
1-Aminocyclopropane-1-carboxylic-acid
- AMP:
-
Antimicrobial protein
- DEFLs :
-
Defensin-like sequences
- MeJA:
-
Methyl jasmonate
- PR:
-
Pathogenesis related
- PsDef1 :
-
Pinus sylvestris defensin 1
- qRT-PCR:
-
Real-time quantitative reverse transcription PCR
- SA:
-
Salicylic acid
References
Adomas A, Asiegbu FO (2006) Analysis of organ-specific responses of Pinus sylvestris to shoot (Gremmeniella abietina) and root (Heterobasidion annosum) pathogens. Physiol Mol Plant Pathol 69:140–152
An C, Mou Z (2012) Non-host defence response in a novel Arabidopsis–Xanthomonas citri subsp. citri pathosystem. PLoS One 7:e31130
Asiegbu FO, Choi W, Li G, Nahalkova J, Dean RA (2003) Isolation of a novel antimicrobial peptide gene (Sp-AMP) homologue from Pinus sylvestris (Scots pine) following infection with the root rot fungus Heterobasidion annosum. FEMS Microbiol Lett 228:27–31
Asiegbu FO, Adomas A, Stenlid J (2005) Conifer root and butt rot caused by Heterobasidion annosum (Fr.) Bref. s.l. Mol Plant Pathol 6:395–409
Azinheira HG, Silva MD, Talhinhas P, Medeira C, Maia I, Petitot AS, Fernandez D (2010) Non-host resistance responses of Arabidopsis thaliana to the coffee leaf rust fungus (Hemileia vastatrix). Botany 88:621–629
Benhamou N, Theriault G (1992) Treatment with chitosan enhances resistance of tomato plants to the crown and root rot pathogen, Fusarium oxysporum f. sp. radicis-lycopersici. Physiol Mol Plant Pathol 41:33–52
Boyd LA, Ridout C, O’Sullivan DM, Leach JE, Leung H (2013) Plant–pathogen interactions: disease resistance in modern agriculture. Trends Genet 29:233–240
Broekaert W, Cammue BP, De Bolle MF, Thevissen K, Samblanx GW, Osborn RW (1997) Antimicrobial peptides from plants. Crit Rev Plant Sci 16:297–323
Broekaert WF, Delaure SL, De Bolle MF, Cammue BP (2006) The role of ethylene in host–pathogen interactions. Annu Rev Phytopathol 44:393–416
Cappellaro C, Mrsa V, Tanner W (1998) New potential cell wall glucanases of Saccharomyces cerevisiae and their involvement in mating. J Bacteriol 180:5030–5037
Carvalho AO, Gomes VM (2009) Plant defensins—prospects for the biological functions and biotechnological properties. Peptides 30:1007–1020
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible WR (2005) Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139:5–17
Felten J, Kohler A, Morin E, Bhalerao RP, Palme K, Martin F, Ditengou FA, Legué V (2009) The ectomycorrhizal fungus Laccaria bicolor stimulates lateral root formation in poplar and Arabidopsis through auxin transport and signaling. Plant Physiol 151:1991–2005
Feys BJ, Parker JE (2000) Interplay of signaling pathways in plant disease resistance. Trends Genet 16:449–455
Fossdal CG, Nagy NE, Sharma P, Lonneborg A (2003) The putative gymnosperm plant defensin polypeptide (SPI1) accumulates after seed germination is not readily released and the SPI1 levels are reduced in Pythium dimorphum-infected spruce roots. Plant Mol Biol 52:291–302
Glazebrook J (2005) Contrasting mechanisms of defence against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43:205–227
Gronberg H, Hietala AM, Haahtela K (2009) Analysing scots pine defence-related transcripts and fungal DNA levels in seedlings single- or dual-inoculated with endophytic and pathogenic Rhizoctonia species. For Pathol 39:377–389
Hantula J, Vainio E (2003) Specific primers for the differentiation of Heterobasidion annosum (s.str.) and H. parviporum infected stumps in northern Europe. Silva Fenn 37:181–187
Hashimoto JG, Beadles-Bohling AS, Wiren KM (2004) Comparison of RiboGreen and 18S rRNA quantitation for normalizing real-time RT-PCR expression analysis. Biotechniques 36:54–60
Hudgins JW, Ralph SG, Franceschi VR, Bohlmann J (2006) Ethylene in induced conifer defence: cDNA cloning, protein expression, and cellular and sub cellular localization of 1-aminocyclopropane-1-carboxylate oxidase in resin duct and phenolic parenchyma cells. Planta 224:865–877
Huitema E, Vleeshouwers VGAA, Francis DM, Kamoun S (2003) Active defence responses associated with non-host resistance of Arabidopsis thaliana to the oomycete pathogen Phytophthora infestans. Mol Plant Pathol 4:487–500
Iida K, Kawaguchi S, Kobayashi N, Yoshida Y, Ishii M, Harada E et al (2011) ARTADE2DB: improved statistical inferences for Arabidopsis gene functions and structure predictions by dynamic structure-based dynamic expression (DSDE) analyses. Plant Cell Physiol 52:254–264
Jenssen H, Hamill P, Hancock REW (2006) Peptide antimicrobial agents. Clin Microbiol Rev 19:491–511
Kovaleva V, Kiyamova R, Cramer R, Krynytskyy H, Gout I, Filonenko V, Gout R (2009) Purification and molecular cloning of antimicrobial peptides from Scots pine seedlings. Peptides 30:2136–2143
Kovaleva V, Krynytskyy H, Gout I, Gout R (2011) Recombinant expression, affinity purification and functional characterization of Scots pine defensin 1. Appl Microbiol Biotechnol 89(1093):1101
Lawrence CB, Mitchell TK, Craven KD, Cho Y, Cramer RA, Kim KH (2008) At death’s door: alternaria pathogenicity mechanisms. Plant Pathol J 24:101–111
Li GS, Asiegbu FO (2004) Use of Scots pine seedling roots as an experimental model to investigate gene expression during interaction with the conifer pathogen Heterobasidion annosum (P-type). J Plant Res 117:155–162
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆Ct method. Methods 25:402–408
Loake G, Grant M (2007) Salicylic acid in plant defence—the players and protagonists. Curr Opin Plant Biol 10:466–472
Loehrer M, Langenbach C, Goellner K, Conrath U, Schaffrath U (2008) Characterization of nonhost resistance of Arabidopsis to the Asian soybean rust. Mol Plant Microbe Interact 21:1421–1430
Lorenzo O, Solano R (2005) Molecular players regulating the jasmonate signalling network. Curr Opin Plant Biol 8:532–540
Mellersh DG, Heath MC (2003) An investigation into the involvement of defence signaling pathways in components of the nonhost resistance of Arabidopsis thaliana to rust fungi also reveals a model system for studying rust fungal compatibility. Mol Plant Microbe Interact 16:398–404
Nandi A, Kachroo P, Fukushige H, Hildebrand DF, Klessig DF, Shah J (2003) Ethylene and jasmonic acid signaling affect the NPR1-independent expression of defence genes without impacting resistance to Pseudomonas syringae and Peronospora parasitica in the Arabidopsis ssi1 mutant. Mol Plant Microbe Interact 16:588–599
O’Connell R, Herbert C, Sreenivasaprasad S, Khatib M, Esquerre-Tugaye M-T, Dumas B (2004) A novel Arabidopsis–Colletotrichum pathosystem for the molecular dissection of plant–fungal interactions. Mol Plant Microbe Interact 17:272–282
Ospina-Giraldo MD, Mullins E, Kang S (2003) Loss of function of the Fusarium oxysporum SNF1 gene reduces virulence on cabbage and Arabidopsis. Curr Genet 44:49–57
Park JY, Jin J, Lee YL, Kang S, Lee YH (2009) Rice blast fungus (Magnaporthe oryzae) infects Arabidopsis thaliana via a mechanism distinct from that required for the infection of rice. Plant Physiol 149:474–486
Penninckx IA, Eggermont K, Terras FR, Thomma BP, De Samblanx GW, Buchala A, Metraux JP, Manners JM, Broekaert WF (1996) Pathogen-induced systemic activation of a plant defensin gene in Arabidopsis follows a salicylic acid-independent pathway. Plant Cell 8:2309–2323
Penninckx IA, Thomma BPHJ, Buchala A, Metraux JP, Broekaert WF (1998) Concomitant activation of jasmonate and ethylene response pathways is required for induction of a plant defensin gene in Arabidopsis. Plant Cell 10:2103–2113
Pervieux I, Bourassa M, Laurans F, Hamelin RC, Seguin A (2004) A spruce defensin showing strong antifungal activity and increased transcript accumulation after wounding and jasmonate treatments. Physiol Mol Plant Pathol 64:331–341
Rabea EI, Badawy ME, Stevens CV, Smagghe G, Steurbaut W (2003) Chitosan as antimicrobial agent: applications and mode of action. Biomacromolecules 4:1457–1465
Sanchez-Vallet A, Ramos B, Bednarek P, López G, Piślewska-Bednarek M, Schulze-Lefert P, Molina A (2010) Tryptophan-derived secondary metabolites in Arabidopsis thaliana confer non-host resistance to necrotrophic Plectosphaerella cucumerina fungi. Plant J 63:115–127
Santos B, Duran A, Valdivieso MH (1997) CHS5, a gene involved in chitin synthesis and mating in Saccharomyces cerevisiae. Mol Cell Biol 17:2485–2496
Segura A, Moreno M, Molina A, Garcia-Olmedo F (1998) Novel defensin subfamily from spinach (Spinacia oleracea). FEBS Lett 435:159–162
Sharma P, Lonneborg A (1996) Isolation and characterization of a cDNA encoding a plant defensin-like protein from roots of Norway spruce. Plant Mol Biol 31:707–712
Silverstein KAT, Graham MA, Paape TD, VandenBosch KA (2005) Genome organization of more than 300 defensin-like genes in Arabidopsis. Plant Physiol 138:600–610
Sooriyaarachchi S, Jaber E, Covarrubias AS, Ubhayasekera W, Asiegbu FO, Mowbray SL (2011) Expression and β-glucan binding properties of Scots pine (Pinus sylvestris L.) antimicrobial protein (Sp-AMP). Plant Mol Biol 77:33–45
Splivallo R, Fischer U, Gobel C, Feussner I, Karlovsky P (2009) Truffles regulate plant root morphogenesis via the production of auxin and ethylene. Plant Physiol 150:2018–2029
Stenlid J (1985) Population structure of Heterobasidion annosum as determined by somatic incompatibility, sexual incompatibility, and isoenzyme patterns. Can J Bot 63:2268–2273
Terras FR, Penninckx IA, Goderis IJ, Broekaert WF (1998) Evidence that the role of plant defensins in radish defence responses is independent of salicylic acid. Planta 206:117–124
Theissen G, Melzer R (2007) Molecular mechanisms underlying origin and diversification of the Angiosperm flower. Ann Bot 100:603–619
Thevissen K, Osborn RW, Acland DP, Broekaert WF (2000) Specific binding sites for an antifungal plant defensin from Dahlia (Dahlia merckii) on fungal cells are required for antifungal activity. Mol Plant Microbe Interact 13:54–61
Uknes S, Mauch-Mani B, Moyer M, Potter S, Williams S, Dincher S, Chandler D, Slusarenko A, Ward E, Ryals J (1992) Acquired resistance in Arabidopsis. Plant Cell 4:645–656
Van Baarlen P, Woltering EJ, Staats M, van Kan JAL (2007) Histochemical and genetic analysis of host and non-host interactions of Arabidopsis with three Botrytis species: an important role for cell death control. Mol Plant Pathol 8:41–54
Van de Mortel J, Villanueva L, Schat H, Kwekkeboom J, Coughlan S, Moerland P, van Ver Loren Themaat E, Koornneef M, Aarts MGM (2006) Large expression differences in genes for iron and zinc homeostasis, stress response, and lignin biosynthesis distinguish roots of Arabidopsis thaliana and the related metal hyperaccumulator Thlaspi caerulescens. Plant Physiol 142:1127–1147
Van Loon LC, Bakker PA, Pieterse CM (1998) Systemic resistance induced by rhizosphere bacteria. Annu Rev Phytopathol 36:453–483
Van Ooij C (2011) Symbiosis: establishing the roots of a relationship. Nat Rev Micro 9:629
Vasiliauskas R, Stenlid J (1998) Influence of spatial scale on population structure of Stereum sanguinolentum in Northern Europe. Mycol Res 102:93–98
Vasiliauskas R, Menkis A, Finlay RD, Stenlid J (2007) Wood-decay fungi in fine living roots of conifer seedlings. New Phytol 174:441–446
Walter C, Grace L, Donaldson SS, Moody J, Gemmel JE, van der Maos S, Kvaalen H, Lonneborg A (1999) An efficient biolistic transformation protocol for Picea abies embryogenic tissue and regeneration of transgenic plants. Can J For Res 29:1539–1546
Zeneli G, Krokene P, Christiansen E, Krekling T, Gershenzon J (2006) Methyl jasmonate treatment of mature Norway spruce (Picea abies) trees increases the accumulation of terpenoid resin components and protects against infection by Ceratocystis polonica a bark beetle-associated fungus. Tree Physiol 26:977–988
Zhang B, Ramonell K, Somerville S, Stacey G (2002) Characterization of early chitin-induced gene expression in Arabidopsis. Mol Plant Microbe Interact 15:963–970
Zimmerli L, Stein M, Lipka V, Schulze-Lefert P, Somerville S (2004) Host and non-host pathogens elicit different jasmonate/ethylene responses in Arabidopsis. Plant J 40:633–646
Acknowledgments
This work was supported by grants from the Academy of Finland (AKA), Helsinki University Research Fund and the Finnish Doctoral Program in Plant Science (FDPPS). We also thank Dr. Hannu Rita for assistance with statistical analysis.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Electronic supplementary material. Supplementary material S1. Supplementary material S2. Supplementary material S3. Supplementary material S4.
Rights and permissions
About this article
Cite this article
Jaber, E., Xiao, C. & Asiegbu, F.O. Comparative pathobiology of Heterobasidion annosum during challenge on Pinus sylvestris and Arabidopsis roots: an analysis of defensin gene expression in two pathosystems. Planta 239, 717–733 (2014). https://doi.org/10.1007/s00425-013-2012-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-013-2012-z