Abstract
Purpose
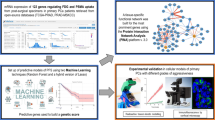
Patients with oligometastatic prostate cancer (PC) may benefit from metastasis-directed therapy (MDT), delaying disease progression and the start of palliative systemic treatment. However, a significant proportion of oligometastatic PC patients progress to polymetastatic PC within a year following MDT, suggesting an underestimation of the metastatic load by current staging modalities. Molecular markers could help to identify true oligometastatic patients eligible for MDT.
Methods
Patients with asymptomatic biochemical recurrence following primary PC treatment were classified as oligo- or polymetastatic based on 18F-choline PET/CT imaging. Oligometastatic patients had up to three metastases at baseline and did not progress to more than three lesions following MDT or surveillance within 1 year of diagnosis of metastases. Polymetastatic patients had > 3 metastases at baseline or developed > 3 metastases within 1 year following imaging. A model aiming to prospectively distinguish oligo- and polymetastatic PC patients was trained using clinicopathological parameters and serum-derived microRNA expression profiles from a discovery cohort of 20 oligometastatic and 20 polymetastatic PC patients. To confirm the models predictive performance, it was applied on biomarker data obtained from an independent validation cohort of 44 patients with oligometastatic and 39 patients with polymetastatic disease.
Results
Oligometastatic PC patients had a more favorable prognosis compared to polymetastatic ones, as defined by a significantly longer median CRPC-free survival (not reached versus 38 months; 95% confidence interval 31–45 months with P < 0.001). Despite the good performance of a predictive model trained on the discovery cohort, with an AUC of 0.833 (0.693–0.973; 95% CI) and a sensitivity of 0.894 (0.714–1.000; 95% CI) for oligometastatic disease, none of the miRNA targets were found to be differentially expressed between oligo- and polymetastatic PC patients in the signature validation cohort. The multivariate model had an AUC of 0.393 (0.534 after cross-validation) and therefore, no predictive ability.
Conclusions
Although PC patients with oligometastatic disease had a more favorable prognosis, no serum-derived biomarkers allowing for prospective discrimination of oligo- and polymetastatic prostate cancer patients could be identified.


Similar content being viewed by others
References
Hellman S, Weichselbaum RR (1995) Oligometastases. J Clin Oncol 13(1):8–10
De Bleser E, Tran PT, Ost P (2017) Radiotherapy as metastasis-directed therapy for oligometastatic prostate cancer. Curr Opin Urol 27(6):587–595
Schweizer MT, Zhou XC, Wang H, Yang T, Shaukat F, Partin AW et al (2013) Metastasis-free survival is associated with overall survival in men with PSA-recurrent prostate cancer treated with deferred androgen deprivation therapy. Ann Oncol 24(11):2881–2886
Ost P, Decaestecker K, Lambert B, Fonteyne V, Delrue L, Lumen N et al (2014) Prognostic factors influencing prostate cancer-specific survival in non-castrate patients with metastatic prostate cancer. Prostate 74(3):297–305
Sridharan S, Steigler A, Spry NA, Joseph D, Lamb DS, Matthews JH et al (2016) Oligometastatic bone disease in prostate cancer patients treated on the TROG 03.04 RADAR trial. Radiother Oncol J Eur Soc Ther Radiol Oncol 121(1):98–102
Sweeney CJ, Chen YH, Carducci M, Liu G, Jarrard DF, Eisenberger M et al (2015) Chemohormonal therapy in metastatic hormone-sensitive prostate cancer. N Engl J Med 373(8):737–746
De Bruycker A, Lambert B, Claeys T, Delrue L, Mbah C, De Meerleer G et al (2017) Prevalence and prognosis of low-volume, oligorecurrent, hormone-sensitive prostate cancer amenable to lesion ablative therapy. BJU Int 120(6):815–821
Hong MK, Macintyre G, Wedge DC, Van Loo P, Patel K, Lunke S et al (2015) Tracking the origins and drivers of subclonal metastatic expansion in prostate cancer. Nat Commun 6:6605
Gundem G, Van Loo P, Kremeyer B, Alexandrov LB, Tubio JMC, Papaemmanuil E et al (2015) The evolutionary history of lethal metastatic prostate cancer. Nature 520(7547):353–357
Weichselbaum RR, Hellman S (2011) Oligometastases revisited. Nat Rev Clin Oncol 8(6):378–382
Ost P, Bossi A, Decaestecker K, De Meerleer G, Giannarini G, Karnes RJ et al (2015) Metastasis-directed therapy of regional and distant recurrences after curative treatment of prostate cancer: a systematic review of the literature. Eur Urol 67(5):852–863
Ost P, Reynders D, Decaestecker K, Fonteyne V, Lumen N, De Bruycker A et al (2018) Surveillance or metastasis-directed therapy for oligometastatic prostate cancer recurrence: a prospective, randomized, multicenter phase II trial. J Clin Oncol 36(5):446–453
Wei J, Zhu H, Liao X (2018) Trigger pSA predicting recurrence from positive choline PET/CT with prostate cancer after initial treatment. Oncotarget 9(18):14630–14641
Perera M, Papa N, Christidis D, Wetherell D, Hofman MS, Murphy DG et al (2016) Sensitivity, specificity, and predictors of positive (68)Ga-prostate-specific membrane antigen positron emission tomography in advanced prostate cancer: a systematic review and meta-analysis. Eur Urol 70(6):926–937
van Leeuwen PJ, Stricker P, Hruby G, Kneebone A, Ting F, Thompson B et al (2016) (68) Ga-PSMA has a high detection rate of prostate cancer recurrence outside the prostatic fossa in patients being considered for salvage radiation treatment. BJU Int 117(5):732–739
Murphy DG, Sweeney CJ, Tombal B (2017) “Gotta Catch ‘em All”, or Do We? Pokemet approach to metastatic prostate cancer. Eur Urol 72(1):1–3
Joice GA, Rowe SP, Pienta KJ, Gorin MA (2017) Oligometastatic prostate cancer: shaping the definition with molecular imaging and an improved understanding of tumor biology. Curr Opin Urol 27(6):533–541
Dhondt B, Rousseau Q, De Wever O, Hendrix A (2016) Function of extracellular vesicle-associated miRNAs in metastasis. Cell Tissue Res 365:621–641
Cornford P, Bellmunt J, Bolla M, Briers E, De Santis M, Gross T et al (2017) EAU-ESTRO-SIOG guidelines on prostate cancer. Part II: treatment of relapsing, metastatic, and castration-resistant prostate cancer. Eur Urol. 71(4):630–642
Moore HM, Kelly AB, Jewell SD, McShane LM, Clark DP, Greenspan R et al (2011) Biospecimen reporting for improved study quality (BRISQ). Cancer Cytopathol 119(2):92–102
Lussier YA, Xing HR, Salama JK, Khodarev NN, Huang Y, Zhang Q et al (2011) MicroRNA expression characterizes oligometastasis(es). PLoS One 6(12):e28650
Lussier YA, Khodarev NN, Regan K, Corbin K, Li H, Ganai S et al (2012) Oligo- and polymetastatic progression in lung metastasis(es) patients is associated with specific microRNAs. PLoS One 7(12):e50141
Pitroda SP, Khodarev NN, Huang L, Uppal A, Wightman SC, Ganai S et al (2018) Integrated molecular subtyping defines a curable oligometastatic state in colorectal liver metastasis. Nat Commun 9(1):1793
Banks RE, Stanley AJ, Cairns DA, Barrett JH, Clarke P, Thompson D et al (2005) Influences of blood sample processing on low-molecular-weight proteome identified by surface-enhanced laser desorption/ionization mass spectrometry. Clin Chem 51(9):1637–1649
McLerran D, Grizzle WE, Feng Z, Bigbee WL, Banez LL, Cazares LH et al (2008) Analytical validation of serum proteomic profiling for diagnosis of prostate cancer: sources of sample bias. Clin Chem 54(1):44–52
Wang K, Yuan Y, Cho J-H, McClarty S, Baxter D, Galas DJ (2012) Comparing the MicroRNA spectrum between serum and plasma. PLoS One 7(7):e41561 (Ahuja SK, editor)
Radwan N, Phillips R, Ross A, Rowe SP, Gorin MA, Antonarakis ES et al (2017) A phase II randomized trial of Observation versus stereotactic ablative RadiatIon for OLigometastatic prostate CancEr (ORIOLE). BMC Cancer 17(1):453
Acknowledgements
This work was supported by “Kom op tegen Kanker (Stand up to Cancer), the Flemisch cancer society” (Bert Dhondt: Emmanuel Vander Schueren Research Grant). Piet Ost is a senior clinical investigator of the Research Foundation—Flanders, Belgium.
Author information
Authors and Affiliations
Contributions
BD sample collection, data analysis and reporting, data management, manuscript writing. EDB data management, manuscript writing. TC sample collection, data collection and management. SB sample collection, manuscript editing. NL manuscript editing. JV project development, manuscript editing. AB project development. VF data collection and management, manuscript editing. JP data analysis and reporting. PG data analysis and reporting, manuscript editing. PO project development, data collection and management, manuscript editing. All authors approved the final version of the manuscript.
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
345_2018_2609_MOESM6_ESM.xlsx
Supplementary material 6 Discovery set annotation table with number of reads and number of mature canonical miRNAs per discovery sample (XLSX 14 kb)
345_2018_2609_MOESM14_ESM.xlsx
Supplementary material 14 Discovery set RT-qPCR result tables (normalized relative quantities without error values—no logarithm transformation) (XLSX 35 kb)
345_2018_2609_MOESM15_ESM.xlsx
Supplementary material 15 Discovery set RT-qPCR result tables (normalized relative quantities with error values—logarithm transformation) (XLSX 58 kb)
345_2018_2609_MOESM18_ESM.xlsx
Supplementary material 18 Predictive models that, tuned to optimize sensitivity, obtained by multivariate analysis (XLSX 10 kb)
345_2018_2609_MOESM19_ESM.xlsx
Supplementary material 19 Consensus variable ranking obtained by averaging the 200 rankings of the resampling procedure (XLSX 10 kb)
345_2018_2609_MOESM21_ESM.xlsx
Supplementary material 21 Validation set RT-qPCR result tables (normalized relative quantities without error values—no logarithm transformation) (XLSX 64 kb)
345_2018_2609_MOESM22_ESM.xlsx
Supplementary material 22 Validation set RT-qPCR result tables (normalized relative quantities with error values—logarithm transformation) (XLSX 114 kb)
Rights and permissions
About this article
Cite this article
Dhondt, B., De Bleser, E., Claeys, T. et al. Discovery and validation of a serum microRNA signature to characterize oligo- and polymetastatic prostate cancer: not ready for prime time. World J Urol 37, 2557–2564 (2019). https://doi.org/10.1007/s00345-018-2609-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00345-018-2609-8