Abstract
Immunoglobulin light (IGL) chain genes encoding σ and λ from channel catfish, Ictalurus punctatus, and λ from Atlantic cod, Gadus morhua, were identified by mining of expressed sequence tag databases, 5′-RACE and RT-PCR protocols. cDNAs for each of these IGL chains encode typical variable (V), joining (J), and constant (C) regions and Southern blot analyses and genomic sequencing show that genes encoding these isotypes, like other teleost IGL genes, are found in a cluster organization of one or two V gene segments, followed by single J and C gene segments, all in the same transcriptional orientation. However, unlike the teleost κ genes, genes encoding catfish σ and λ are few in number and the two isotypes are each encoded by only two clusters. Similarly, Atlantic cod λ genes are predicted to be encoded by two or three clusters. As expected, sequence and phylogenetic analyses comparisons demonstrate that catfish Vσ and Cσ genes are most similar to Vσ and Cσ genes of other ectothermic vertebrates. Although catfish and Atlantic cod Vλ genes cluster with other vertebrate Vλ genes, their Cλ sequences cluster in a distinct group separate from other vertebrate IGL C sequences. However, support for classifying these sequences as λ, is their V and J recombination signal sequence (RSS) organization. The catfish and Atlantic cod genes have typical λ-like RSS with the Vλ RSS consisting of heptamer-23 bp spacer-nonamer and the Jλ RSS consisting of heptamer-12 bp spacer-nonamer. This is the first report demonstrating the presence of Igλ in teleosts.








Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Arakawa H, Shimizu T, Takeda S (1996) Re-evaluation of the probabilities for productive arrangements on the kappa and lambda loci. Int Immunol 8:91–99. doi:10.1093/intimm/8.1.91
Asenbauer H, Combriato G, Klobeck HG (1999) The immunoglobulin lambda light chain enhancer consists of three modules which synergize in activation of transcription. Eur J Immunol 29:713–724. doi:10.1002/(SICI)1521-4141(199902)29:02<713::AID-IMMU713>;3.0.CO;2-M
Bengtén E, Wilson M, Miller N, Clem LW, Pilström L, Warr GW (2000) Immunoglobulin isotypes: structure, function, and genetics. Curr Top Microbiol Immunol 248:189–219
Chen K, He B, Santini P, Chiu A, Chadburn A, Xu W, Cerutti A, Cattalini M, Plebani A (2008) Cmu to Cdelta class switch recombination and IgD production contribute to mucosal immunity. FASEB J 22:Abstract 854.7
Church GM, Gilbert W (1984) Genomic sequencing. Proc Natl Acad Sci USA 81:1991–1995. doi:10.1073/pnas.81.7.1991
Coleclough C, Perry RP, Karjalainen K, Weigert M (1981) Aberrant rearrangements contribute significantly to the allelic exclusion of immunoglobulin gene expression. Nature 290:372–378. doi:10.1038/290372a0
Corcoran AE (2005) Immunoglobulin locus silencing and allelic exclusion. Semin Immunol 17:141–154. doi:10.1016/j.smim.2005.01.002
Coscia MR, Giacomelli S, De Santi C, Varriale S, Oreste U (2008) Immunoglobulin light chain isotypes in the teleost Trematomus bernacchii. Mol Immunol 45:3096–3106. doi:10.1016/j.molimm.2008.03.006
Criscitiello MF, Flajnik MF (2007) Four primordial immunoglobulin light chain isotypes, including lambda and kappa, identified in the most primitive living jawed vertebrates. Eur J Immunol 37:2683–2694. doi:10.1002/eji.200737263
Daggfeldt A, Bengtén E, Pilström L (1993) A cluster type organization of the loci of the immunoglobulin light chain in Atlantic cod (Gadus morhua L.) and rainbow trout (Oncorhynchus mykiss Walbaum) indicated by nucleotide sequences of cDNA and hybridization analysis. Immunogenetics 38:199–209. doi:10.1007/BF00211520
Danilova N, Bussmann J, Jekosch K, Steiner LA (2005) The immunoglobulin heavy-chain locus in zebrafish: identification and expression of a previously unknown isotype, immunoglobulin Z. Nat Immunol 6:295–302. doi:10.1038/ni1166
Das S, Nikolaidis N, Klein J, Nei M (2008) Evolutionary redefinition of immunoglobulin light chain isotypes in tetrapods using molecular markers. Proc Natl Acad Sci USA 105:16647–16652. doi:10.1073/pnas.0808800105
DeLuca D, Wilson M, Warr GW (1983) Lymphocyte heterogeneity in the trout, Salmo gairdneri, defined with monoclonal antibodies to IgM. Eur J Immunol 13:546–551. doi:10.1002/eji.1830130706
Edelman GM, Gally JA (1962) The nature of Bence-Jones proteins. Chemical similarities to polypetide chains of myeloma globulins and normal gamma-globulins. J Exp Med 116:207–227. doi:10.1084/jem.116.2.207
Fleurant M, Changchien L, Chen CT, Flajnik MF, Hsu E (2004) Shark Ig light chain junctions are as diverse as in heavy chains. J Immunol 173:5574–5582
Gadol N, Waldman RH, Clem LW (1976) Inhibition of macrophage migration by normal guinea pig intestinal secretions. Proc Soc Exp Biol Med 151:654–658
Geier JK, Schlissel MS (2006) Pre-BCR signals and the control of Ig gene rearrangements. Semin Immunol 18:31–39. doi:10.1016/j.smim.2005.11.001
Ghaffari SH, Lobb CJ (1993) Structure and genomic organization of immunoglobulin light chain in the channel catfish. An unusual genomic organizational pattern of segmental genes. J Immunol 151:6900–6912
Ghaffari SH, Lobb CJ (1997) Structure and genomic organization of a second class of immunoglobulin light chain genes in the channel catfish. J Immunol 159:250–258
Giudicelli V, Lefranc MP (1999) Ontology for immunogenetics: the IMGT-ONTOLOGY. Bioinformatics 15:1047–1054
Greenberg AS, Steiner L, Kasahara M, Flajnik MF (1993) Isolation of a shark immnoglobulin light chain cDNA clone encoding a protein resembling mammalian k light chains: implications for the evolution of light chains. Proc Natl Acad Sci USA 90:10603–10607. doi:10.1073/pnas.90.22.10603
Haire RN, Ota T, Rast JP, Litman RT, Chan FY, Zon LI, Litman GW (1996) A third Ig light chain gene isotype in Xenopus laevis consists of six distinct VL families and is related to mammalian lambda genes. J Immunol 157:1544–1550
Haire RN, Rast JP, Litman RT, Litman GW (2000) Characterization of three isotypes of immunoglobulin light chains and T-cell antigen receptor alpha in zebrafish. Immunogenetics 51:915–923. doi:10.1007/s002510000229
Hansen JD, Landis ED, Phillips RB (2005) Discovery of a unique Ig heavy chain isotype in rainbow trout: implications for a novel B-cell developmental pathway in teleost fish. Proc Natl Acad Sci USA 102:6919–6924. doi:10.1073/pnas.0500027102
Heery DM, Gannon F, Powell R (1990) A simple method for subcloning DNA fragments from gel slices. Trends Genet 6:173. doi:10.1016/0168-9525(90)90158-3
Hogan RJ, Waldbieser GC, Goudie CA, Antao A, Godwin UB, Wilson MR, Miller NW, Clem LW, McConnell TJ, Wolters WR, Chinchar VG (1999) Molecular and immunologic characterization of gynogenetic channel catfish (Ictalurus punctatus). Mar Biotechnol 1:317–327. doi:10.1007/PL00011781
Hohman VS, Schluter SF, Marchalonis JJ (1992) Complete sequence of a cDNA clone specifying sandbar shark immunoglobulin light chain: gene organisation and implications for the evolution of light chains. Proc Natl Acad Sci USA 89:276–280. doi:10.1073/pnas.89.1.276
Hsu E, Criscitiello MF (2006) Diverse immunoglobulin light chain organizations in fish retain potential to revise B cell receptor specificities. J Immunol 177:2452–2462
Hsu E, Lefkovits I, Flajnik M, Du Pasquier L (1991) Light chain heterogeneity in the amphibian Xenopus. Mol Immunol 28:985–994. doi:10.1016/0161-5890(91)90184-L
Ishikawa J, Imai E, Moritomo T, Nakao M, Yano T, Tomana M (2004) Characterisation of a fourth immunoglobulin light chain isotype in the common carp. Fish Shellfish Immunol 16:369–379. doi:10.1016/j.fsi.2003.06.002
Kubo RT, Rosenblum IY, Benedict AA (1970) The unblocked N-terminal sequence of chicken IgG lambda-like light chains. J Immunol 105:534–536
Lee SS, Greenberg A, Hsu E (2000) Evolution and somatic diversification of immunoglobulin light chains. Curr Top Microbiol Immunol 248:285–300
Lefranc MP, Pommie C, Kaas Q, Duprat E, Bosc N, Guiraudou D, Jean C, Ruiz M, Da Piedade I, Rouard M, Foulquier E, Thouvenin V, Lefranc G (2005) IMGT unique numbering for immunoglobulin and T cell receptor constant domains and Ig superfamily C-like domains. Dev Comp Immunol 29:185–203. doi:10.1016/j.dci.2004.07.003
Liu YJ, de Bouteiller O, Arpin C, Briere F, Galibert L, Ho S, Martinez-Valdez H, Banchereau J, Lebecque S (1996) Normal human IgD+IgM− germinal center B cells can express up to 80 mutations in the variable region of their IgD transcripts. Immunity 4:603–613. doi:10.1016/S1074-7613(00)80486-0
Lobb CJ (1986) Preferential expression of catfish light chain immunoglobulin isotypes in anti-dinitrophenyl antibodies. J Immunogenet 13:19–28. doi:10.1111/j.1744-313X.1986.tb01079.x
Lobb CJ, Clem LW (1982) Fish lymphocytes differ in the expression of surface immunoglobulin. Dev Comp Immunol 6:473–479
Lobb CJ, Olson MO, Clem LW (1984) Immunoglobulin light chain classes in a teleost fish. J Immunol 132:1917–1923
Lundqvist ML, Middleton DL, Radford C, Warr GW, Magor KE (2006) Immunoglobulins of the non-galliform birds: antibody expression and repertoire in the duck. Dev Comp Immunol 30:93–100. doi:10.1016/j.dci.2005.06.019
Magor KE, Higgins DA, Middleton DL, Warr GW (1994) cDNA sequence and organization of the immunoglobulin light chain gene of the duck, Anas platyrhynchos. Dev Comp Immunol 18:523–531. doi:10.1016/S0145-305X(06)80006-6
McCormack WT, Carlson LM, Tjoelker LW, Thompson CB (1989) Evolutionary comparison of the avian IgL locus: combinatorial diversity plays a role in the generation of the antibody repertoire in some avian species. Int Immunol 1:332–341. doi:10.1093/intimm/1.4.332
Miller NW, Bly JE, van Ginkel F, Ellsaesser CF, Clem LW (1987) Phylogeny of lymphocyte heterogeneity: identification and separation of functionally distinct subpopulations of channel catfish lymphocytes with monoclonal antibodies. Dev Comp Immunol 11:739–747. doi:10.1016/0145-305X(87)90061-9
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16:1215. doi:10.1093/nar/16.3.1215
Miller NW, Rycyzyn MA, Wilson MR, Warr GW, Naftel JP, Clem LW (1994) Development and characterization of channel catfish long term B cell lines. J Immunol 152:2180–2189
Mostoslavsky R, Alt FW, Rajewsky K (2004) The lingering enigma of the allelic exclusion mechanism. Cell 118:539–544. doi:10.1016/j.cell.2004.08.023
Nadel B, Cazenave PA, Sanchez P (1990) Murine lambda gene rearrangements: the stochastic model prevails over the ordered model. EMBO J 9:435–440
Oberdoerffer P, Novobrantseva TI, Rajewsky K (2003) Expression of a targeted lambda 1 light chain gene is developmentally regulated and independent of Ig kappa rearrangements. J Exp Med 197:1165–1172. doi:10.1084/jem.20030402
Ota T, Sitnikova T, Nei M (2000) Evolution of vertebrate immunoglobulin variable gene segments. Curr Top Microbiol Immunol 248:221–245
Partula S, Schwager J, Timmusk S, Pilström L, Charlemagne J (1996) A second immunoglobulin light chain isotype in the rainbow trout. Immunogenetics 45:44–51. doi:10.1007/s002510050165
Qin T, Ren L, Hu X, Guo Y, Fei J, Zhu Q, Butler JE, Wu C, Li N, Hammarstrom L, Zhao Y (2008) Genomic organization of the immunoglobulin light chain gene loci in Xenopus tropicalis: evolutionary implications. Dev Comp Immunol 32:156–165
Ramsden DA, Wu GE (1991) Mouse kappa light-chain recombination signal sequences mediate recombination more frequently than do those of lambda light chain. Proc Natl Acad Sci USA 88:10721–10725. doi:10.1073/pnas.88.23.10721
Rast JP, Anderson MK, Ota T, Litman RT, Margittai M, Shamblott MJ, Litman GW (1994) Immunoglobulin light chain class multiplicity and alternative organizational forms in early vertebrate phylogeny. Immunogenetics 40:83–99. doi:10.1007/BF00188170
Reynaud C-A, Dahan A, Weill J-C (1983) Complete sequence of a chicken λ light chain immunoglobulin derived from the nucleotide sequence of its mRNA. Proc Natl Acad Sci USA 80:4099–4103. doi:10.1073/pnas.80.13.4099
Reynaud C-A, Anquez V, Grimal H, Weill J-C (1987) A hyperconversion mechanism generates the chicken light chain preimmune repertoire. Cell 48:379–388. doi:10.1016/0092-8674(87)90189-9
Saha NR, Suetake H, Suzuki Y (2004) Characterization and expression of the immunoglobulin light chain in the fugu: evidence of a solitaire type. Immunogenetics 56:47–55. doi:10.1007/s00251-004-0662-5
Schwager J, Burckert N, Schwager M, Wilson M (1991) Evolution of immunoglobulin light chain genes: analysis of Xenopus IgL isotypes and their contribution to antibody diversity. EMBO J 10:505–511
Shen L, Stuge TB, Evenhuis JP, Bengten E, Wilson M, Chinchar VG, Clem LW, Miller NW (2003) Channel catfish NK-like cells are armed with IgM via a putative FcmicroR. Dev Comp Immunol 27:699–714. doi:10.1016/S0145-305X(03)00042-9
Sirac C, Carrion C, Duchez S, Comte I, Cogne M (2006) Light chain inclusion permits terminal B cell differentiation and does not necessarily result in autoreactivity. Proc Natl Acad Sci USA 103:7747–7752. doi:10.1073/pnas.0509121103
Solem ST, Stenvik J (2006) Antibody repertoire development in teleosts—a review with emphasis on salmonids and Gadus morhua L. Dev Comp Immunol 30:57–76. doi:10.1016/j.dci.2005.06.007
Storb U, Wilson R, Selsing E, Walfield A (1981) Rearranged and germline immunoglobulin kappa genes: different states of DNase I sensitivity of constant kappa genes in immunocompetent and nonimmune cells. Biochemistry 20:990–996. doi:10.1021/bi00507a053
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. doi:10.1093/nar/25.24.4876
Timmusk S, Partula S, Pilstrom L (2000) Different genomic organization and expression of immunoglobulin light-chain isotypes in the rainbow trout. Immunogenetics 51:905–914. doi:10.1007/s002510000221
van der Burg M, Tumkaya T, Boerma M, de Bruin-Versteeg S, Langerak AW, van Dongen JJ (2001) Ordered recombination of immunoglobulin light chain genes occurs at the IGK locus but seems less strict at the IGL locus. Blood 97:1001–1008. doi:10.1182/blood.V97.4.1001
van Ginkel FW, Miller NW, Lobb CJ, Clem LW (1992) Characterization of anti-hapten antibodies generated in vitro by channel catfish peripheral blood lymphocytes. Dev Comp Immunol 16:139–151. doi:10.1016/0145-305X(92)90014-4
Wu TT, Kabat EA (1970) An analysis of the sequences of the variable regions of Bence Jones proteins and myeloma light chains and their implications for antibody complementarity. J Exp Med 132:211–250. doi:10.1084/jem.132.2.211
Zezza DJ, Stewart SE, Steiner LA (1992) Genes encoding Xenopus laevis Ig L chains. Implications for the evolution of k and l chains. J Immunol 149:3968–3977
Zimmerman AM, Yeo G, Howe K, Maddox BJ, Steiner LA (2008) Immunoglobulin light chain (IgL) genes in zebrafish: genomic configurations and inversional rearrangements between (V(L)-J(L)-C(L)) gene clusters. Dev Comp Immunol 32:421–434. doi:10.1016/j.dci.2007.08.005
Zou YR, Takeda S, Rajewsky K (1993) Gene targeting in the Ig kappa locus: efficient generation of lambda chain-expressing B cells, independent of gene rearrangements in Ig kappa. EMBO J 12:811–820
Acknowledgments
This work was supported by grants from the National Institutes of Health (RO1AI-19530), US Department of Agriculture (2002-35204-12211, 2003-35205-12829, and 2006-35204-16880), UMMC IRSP (59908) awarded to M.W., and UMMC IRSP (55901) awarded to N.W.M. We also thank the Drs Ellen Hsu and Mike Criscitiello for sharing their IGL sequence information pre-publication.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
sTable 1
Channel catfish primers used for PCR, probes and sequencing (PDF 18 kb)
sTable 2
Molecular markers distinguishing κ, λ and σ isotypes (PDF 12 kb)
sFig. 1
Catfish Igσ and Igλ and Atlantic cod Igλ chains. Nucleotide and predicted amino acid sequences of a catfish Igσ, b catfish Igλ and c Atlantic cod Igλ. The predicted leader sequence, VL, JL and C regions are labeled above the sequence. The stop (TGA) codons are marked with (*) and nucleotide and amino acid numbers are at left. GenBank accession numbers are: Catfish Igσ (IpIgσ, EU872021), catfish Igλ (IpIgLλ, EU872022), Atlantic cod Ig λ (GmIgLλ, CAC03754). (PDF 25 kb)
sFig. 2
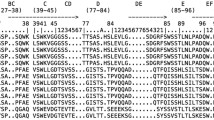
Amino acid alignment of catfish Vσ sequences. IGSV1-1*01, IGSV1-2*01, and IGSV1-3P are genomic sequences obtained from IgLσ gene clusters 3C and 5A, respectively. IGSV1-2*02 was obtained by RT-PCR and IGSV1-1*02 and IGSV1-1*03 by 5'-RACE. The conserved cysteine residues that form the intrachain disulfide bonds are labeled (*). Residues identical to the catfish sequence are shown as (.) and (–) represents gaps in the alignment, an arrow denotes the primer used for 5′ RACE. GenBank accession numbers are: IGSV1-1*01 (EU872024), IGSV1-2*01 and IGSV1-3P (EU872023), IGSV-2*02 (EU925377), IGSV1-1*02 (EU925378), IGSV1-1*03 (EU925379). (PDF 48 kb)
sFig. 3
Amino acid alignment of catfish Cσ sequences. IGSC1*01 is a genomic sequence obtained from the IgLσ gene cluster 3C. IGSC1*02, IGSC1*03 and IGSC1*04 are representative sequences obtained by 5'-RACE. In total, 23 RACE products were amplified from PBL cDNA made from a single catfish: three IGSC1*03, five IGSC1*02, and 15 IGSC1*04 products were sequenced. The conserved cysteine residues that form the intrachain disulfide bonds are labeled (*) and the cysteine that form the interchain disulfide bond is marked by (↓). The number of amino acids is on the right. Residues identical to the catfish sequence are shown as (.). GenBank accession numbers are: IGSC1*01 (EU872024), IGSC1*02 (EU925380), IGSC1*03 (EU925381), IGSC1*04 (EU925382). (PDF 16 kb)
sFig. 4
Comparison of nucleotide sequence identities of teleost Vσ and Vλ sequences. Pairwise alignments were performed using CLUSTAL W and yellow highlights indicate >75% identity. VS and Vl subgroup membership is indicated above the sequence names and Genbank accession numbers are listed in the figure. Accession numbers referring to genomic sequences containing more than 1 V are labeled −1, −2, or −3, depending on the order of the V in the gene fragment. Species abbreviations are: Cc carp, Dr zebrafish, Om rainbow trout, Ip channel catfish, If blue catfish, Gm Atlantic cod. (PDF 7 kb)
sFig. 5
Amino acid alignment of catfish Vλ sequences. IGLV1-1*01 is a genomic sequence obtained from the IgLλ gene cluster 2A. IGLV1-1*02 and IGLV1-1*03 were obtained by RT-PCR and IGLV1-2*01 by 5′-RACE. The conserved cysteine residues that form the intrachain disulfide bonds are labeled (*) and the two cysteines found at the 3' end are labeled (^). Residues identical to the catfish sequence are shown as (.) and (–) represents gaps in the alignment, arrows denotes the primers used for 5′ RACE and genomic PCR. GenBank accession numbers are: IGLV1-1*01 (EU872025), IGLV1-1*02 (EU925383), IGLV1-1*03 (EU925384), IGLV1-2*01 (EU925385), IGLV1-2*02 (FJ716623). (PDF 49 kb)
sFig. 6
Amino acid alignment of representative IGL sequences. Sequences are aligned as recommended by Das et al. 2008 and molecular markers that distinguish κ, λ, and σ isotypes are boxed in blue (or red when not valid for teleosts). The amino acids conforming to the criteria proposed by Das et al. 2008 are highlighted in yellow and additional markers not defined by Das are highlighted in green. CDR regions are removed to optimize the alignment and their respective locations are marked by triangles. Amino acid numbers are according to the standardized rules of the IMGT unique numbering and are shown above the sequence (Lefranc et al. 2005). Accession numbers are catfish G-type (IpG, L25533); Carp L1 (CcL1, V and J (AB073328), C (AB015904)); zebrafish L1 (DrL1, AF246185); rainbow trout L1 (OmL1, X65260); Atlantic cod L1 (GmL1, V and J (AF104898.3), C (X68514)); catfish F-type (IpF, U25705); carp L3 (CcL3, V and J (BAB91006), C (BAB90987)); zebrafish L3 (DrL3, AB246193); X. laevis κ (Xlκ, V (L15570), J and C (M94392)); green anole κ (Acκ, FG748778); human κ (Hsκ, AAH73764); channel catfish λ (Ipλ, EU872022); blue catfish λ (Ifλ, CK403484); rainbow trout λ (Omλ, BX861350); Atlantic cod λ (Gmλ, AJ293807); X. laevis III (XlIII, L76579); green anole λ (Acλ, FG746006); chicken λ (Ggλ, K00678); human λ (Ηsλ, ABU90549); catfish σ (Ιpσ, EU872023); carp L2 (CcL2, V and J (AB091113), C (BAC81202)); zebrafish L2 (DrL2, AAG31698); rainbow trout L2 (OmL2, AAB41310); Atlantic cod σ (Gmσ, V (EX732557), J and C (EC907978)); and X. laevis σ (Xlσ, AAH82413). (PDF 69 kb)
Rights and permissions
About this article
Cite this article
Edholm, ES., Wilson, M., Sahoo, M. et al. Identification of Igσ and Igλ in channel catfish, Ictalurus punctatus, and Igλ in Atlantic cod, Gadus morhua . Immunogenetics 61, 353–370 (2009). https://doi.org/10.1007/s00251-009-0365-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-009-0365-z